

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0070.9
(437 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
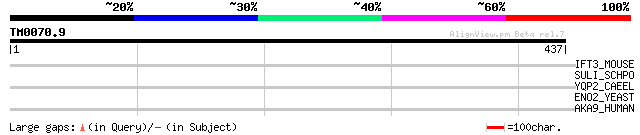
Score E
Sequences producing significant alignments: (bits) Value
IFT3_MOUSE (Q64345) Interferon-induced protein with tetratricope... 35 0.50
SULI_SCHPO (Q9URY8) Probable sulfate permease C869.05c 31 7.2
YQP2_CAEEL (Q09303) Hypothetical protein F07F6.2 in chromosome III 30 9.4
ENO2_YEAST (P00925) Enolase 2 (EC 4.2.1.11) (2-phosphoglycerate ... 30 9.4
AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A ... 30 9.4
>IFT3_MOUSE (Q64345) Interferon-induced protein with
tetratricopeptide repeats 3 (IFIT-3)
(Glucocorticoid-attenuated response gene 49 protein)
(GARG-49) (IRG2)
Length = 403
Score = 34.7 bits (78), Expect = 0.50
Identities = 26/86 (30%), Positives = 41/86 (47%), Gaps = 4/86 (4%)
Query: 17 VEDFMNDTFVEDMMQQEMEFYQQQQHANTVRPKKTRRVIKRDREAGNERLMKDYFSKNPV 76
+E+ F D ++Q ME Q Q+ + K +++ EA ERL+KD K P
Sbjct: 183 LEEKPEKQFSVDALKQAMELNPQNQYLKVLLALK---LLRMGEEAEGERLIKDALGKAPN 239
Query: 77 YTEELFR-RRFRMRKHVFLRIVEALG 101
T+ L + +F +K R +E LG
Sbjct: 240 QTDVLQKAAQFYKKKGNLDRAIELLG 265
>SULI_SCHPO (Q9URY8) Probable sulfate permease C869.05c
Length = 840
Score = 30.8 bits (68), Expect = 7.2
Identities = 29/105 (27%), Positives = 45/105 (42%), Gaps = 8/105 (7%)
Query: 8 DWNTFYNECVEDFMNDTFVEDMMQQEMEFYQQQQ--HANTVRPKKTRRVIKRDREAGNER 65
D+ +NE ++ ND D Q +F Q T+ K I D E
Sbjct: 25 DYPDRFNE-FDNSQNDH--NDYTQNNAQFQNAQTTTFGRTISRVKAYYEIPEDDELDELA 81
Query: 66 LMKDYFSKNPVYTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMS 110
+ +F KN T +F+ K +F I+E L +YNPY+L++
Sbjct: 82 SIPQWFKKN--VTSNIFKNFLHYLKSLF-PIIEWLPNYNPYWLIN 123
>YQP2_CAEEL (Q09303) Hypothetical protein F07F6.2 in chromosome III
Length = 261
Score = 30.4 bits (67), Expect = 9.4
Identities = 16/53 (30%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Query: 8 DWNTFYNECVEDFMNDTFVEDMMQQEMEFYQQQQHANTVRPKKTRRVIKRDRE 60
DW+ E ED N E + +++MEF +Q H + K+ +V+K+ E
Sbjct: 185 DWHV--KEANEDVKNKIEQEGLKRKDMEFEEQLYHLRIEKVKRREQVLKQKLE 235
>ENO2_YEAST (P00925) Enolase 2 (EC 4.2.1.11) (2-phosphoglycerate
dehydratase) (2-phospho-D-glycerate hydro-lyase)
Length = 436
Score = 30.4 bits (67), Expect = 9.4
Identities = 22/82 (26%), Positives = 36/82 (43%), Gaps = 6/82 (7%)
Query: 225 PTIMLEAVASQDLW-IWHAFFGIAGSN---NDITVLNQSPVFNEVLRGAAPMVKFRVNE- 279
P + +E ++D W W FF AG +D+TV N + + + + AA + +VN+
Sbjct: 290 PIVSIEDPFAEDDWEAWSHFFKTAGIQIVADDLTVTNPARIATAIEKKAADALLLKVNQI 349
Query: 280 -TMYHIGYYLADGIYPEWGTFV 300
T+ D WG V
Sbjct: 350 GTLSESIKAAQDSFAANWGVMV 371
>AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A
anchoring protein 9) (PRKA9) (A-kinase anchor protein 450
kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP
350) (hgAKAP 350) (AKAP 120 like protein) (Hyperion
protein) (Yotiao protein
Length = 3911
Score = 30.4 bits (67), Expect = 9.4
Identities = 24/86 (27%), Positives = 40/86 (45%), Gaps = 10/86 (11%)
Query: 21 MNDTFVEDMMQQEMEFYQQQQHANTVRPKKTRRVIKRDREAGNE------RLMKDYFSKN 74
+N+ +EDM Q+ + YQ+ Q A + + R ++R RE + RL + ++
Sbjct: 1594 LNEEQLEDMRQELVRQYQEHQQATELLRQAHMRQMERQREDQEQLQEEIKRLNRQLAQRS 1653
Query: 75 PVYTEELFRRRFRMRKHVFLRIVEAL 100
+ E L R R V L +EAL
Sbjct: 1654 SIDNENLVSERER----VLLEELEAL 1675
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,992,230
Number of Sequences: 164201
Number of extensions: 2331332
Number of successful extensions: 5097
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 5096
Number of HSP's gapped (non-prelim): 6
length of query: 437
length of database: 59,974,054
effective HSP length: 113
effective length of query: 324
effective length of database: 41,419,341
effective search space: 13419866484
effective search space used: 13419866484
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0070.9


