

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0069.5
(595 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
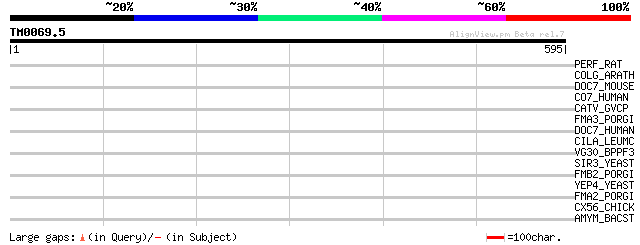
PERF_RAT (P35763) Perforin 1 precursor (P1) (Lymphocyte pore for... 41 0.010
COLG_ARATH (Q8RWD0) Zinc finger protein CONSTANS-LIKE 16 35 0.73
DOC7_MOUSE (Q8R1A4) Dedicator of cytokinesis protein 7 (Fragment) 33 2.8
CO7_HUMAN (P10643) Complement component C7 precursor 33 2.8
CATV_GVCP (O91466) Viral cathepsin (EC 3.4.22.50) (V-cath) (Cyst... 33 2.8
FMA3_PORGI (Q51826) Major fimbrial subunit protein, type III pre... 32 3.6
DOC7_HUMAN (Q96N67) Dedicator of cytokinesis protein 7 (Fragment) 32 4.7
CILA_LEUMC (O53079) Citrate lyase alpha chain (EC 4.1.3.6) (Citr... 32 4.7
VG30_BPPF3 (P03626) 32.7 kDa protein (ORF 301) 32 6.1
SIR3_YEAST (P06701) Regulatory protein SIR3 (Silent information ... 32 6.1
FMB2_PORGI (Q51825) Major fimbrial subunit protein, type II prec... 32 6.1
YEP4_YEAST (P40041) Hypothetical 56.6 kDa protein in THO1-ICL1 i... 31 8.0
FMA2_PORGI (Q51822) Major fimbrial subunit protein, type II prec... 31 8.0
CX56_CHICK (P29415) Gap junction Cx56 protein (Connexin 56) 31 8.0
AMYM_BACST (P19531) Maltogenic alpha-amylase precursor (EC 3.2.1... 31 8.0
>PERF_RAT (P35763) Perforin 1 precursor (P1) (Lymphocyte pore
forming protein) (Cytolysin)
Length = 554
Score = 40.8 bits (94), Expect = 0.010
Identities = 40/164 (24%), Positives = 67/164 (40%), Gaps = 17/164 (10%)
Query: 157 ALARFIEKFGTHILVGLSIGGKDLVLVK----QDVSSNLEPSELKKNLD-----ELGDQL 207
A R I +GTH + + +GG+ VL Q L E+ L +G Q
Sbjct: 210 AYRRLISSYGTHFITAVDLGGRVSVLTALRTCQLTLDGLTADEVGDCLSVEAQVSIGAQA 269
Query: 208 FTGTCNFLPKTKDQKHKVPPAF-DVFGPQIVAFNNSTCVCAKDGITVICAKRGGDTQVSN 266
+ + K ++HK+ +F + + V + D + G +
Sbjct: 270 SVSSEYKACEEKKKQHKIATSFHQTYRERHVEVLGGPLDSSNDLLF------GNQATPEH 323
Query: 267 HSEWLLTVPNKPDAVDFSFIPFTSLLK-AAPGRGFLSHAINLYL 309
S W+ ++P +PD VD+S P LL+ + P R L AI+ Y+
Sbjct: 324 FSTWIASLPTRPDVVDYSLEPLHILLEDSDPKREALRQAISHYV 367
>COLG_ARATH (Q8RWD0) Zinc finger protein CONSTANS-LIKE 16
Length = 417
Score = 34.7 bits (78), Expect = 0.73
Identities = 29/110 (26%), Positives = 42/110 (37%), Gaps = 16/110 (14%)
Query: 429 GKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVK 488
G K + C + VK W D AF+ V + H R+ S A+V
Sbjct: 10 GAKTARACDSCVKRRARWYCAADDAFLCQSCDSLVHSANPLARRHERVRLKTASPAVVKH 69
Query: 489 SN------------WTQGSTQRS----GSGSGSGSSVFSAISTSIQGKDQ 522
SN W G T+++ GSG + SS+F + I +DQ
Sbjct: 70 SNHSSASPPHEVATWHHGFTRKARTPRGSGKKNNSSIFHDLVPDISIEDQ 119
>DOC7_MOUSE (Q8R1A4) Dedicator of cytokinesis protein 7 (Fragment)
Length = 727
Score = 32.7 bits (73), Expect = 2.8
Identities = 16/53 (30%), Positives = 29/53 (54%), Gaps = 2/53 (3%)
Query: 360 KLYVNTAKVTVGKRPVTG--MRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIED 410
++ V T VT K + G +++ L+ M CN+ A++LQ+ T L +K +
Sbjct: 75 EIIVQTVSVTESKESILGGVLKVLLQSMACNQSAVYLQHCFATQRALVSKFPE 127
>CO7_HUMAN (P10643) Complement component C7 precursor
Length = 843
Score = 32.7 bits (73), Expect = 2.8
Identities = 19/58 (32%), Positives = 34/58 (57%), Gaps = 7/58 (12%)
Query: 149 VPSSWDPTALARFIEKFGTHILVGLSIGGKDLVL-------VKQDVSSNLEPSELKKN 199
+PS +D +A R I+++GTH L S+GG+ VL +KQ+ +++E + K +
Sbjct: 283 LPSLYDYSAYRRLIDQYGTHYLQSGSLGGEYRVLFYVDSEKLKQNDFNSVEEKKCKSS 340
>CATV_GVCP (O91466) Viral cathepsin (EC 3.4.22.50) (V-cath)
(Cysteine proteinase) (CP)
Length = 333
Score = 32.7 bits (73), Expect = 2.8
Identities = 31/93 (33%), Positives = 43/93 (45%), Gaps = 10/93 (10%)
Query: 3 EGIVEKALNSLGKGFD---LTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVD 59
EG V A N GFD S F L G R V+ NE + REL V GP++ V++D
Sbjct: 201 EGGVVSAENEPYYGFDGVCKKSPFELSI-SGSRRYVLQNENKLRELLVVN-GPIS-VAID 257
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPG 92
+ DL Y++ I + +E N + G
Sbjct: 258 V----SDLINYKAGIADICENNEGLNHAVLLVG 286
>FMA3_PORGI (Q51826) Major fimbrial subunit protein, type III
precursor (Fimbrillin) (Fimbrilin)
Length = 353
Score = 32.3 bits (72), Expect = 3.6
Identities = 15/51 (29%), Positives = 26/51 (50%)
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSW 110
I C KG LT++ LS +M+ FN + P+ Y+ + FE+ ++
Sbjct: 243 ILCVKGKLTKHDGTALSSEEMTAAFNAGWIVANNDPTTYYPVLVNFESNNY 293
>DOC7_HUMAN (Q96N67) Dedicator of cytokinesis protein 7 (Fragment)
Length = 1302
Score = 32.0 bits (71), Expect = 4.7
Identities = 16/53 (30%), Positives = 28/53 (52%), Gaps = 2/53 (3%)
Query: 360 KLYVNTAKVTVGKRPVTG--MRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIED 410
++ V T VT K + G +++ L M CN+ A++LQ+ T L +K +
Sbjct: 650 EIVVQTVSVTESKESILGGVLKVLLHSMACNQSAVYLQHCFATQRALVSKFPE 702
>CILA_LEUMC (O53079) Citrate lyase alpha chain (EC 4.1.3.6) (Citrase
alpha chain) (Citrate (pro-3S)-lyase alpha chain)
(Citrate CoA-transferase subunit) (EC 2.8.3.10)
Length = 512
Score = 32.0 bits (71), Expect = 4.7
Identities = 27/91 (29%), Positives = 41/91 (44%), Gaps = 2/91 (2%)
Query: 32 ERLVILNETERRELTVPGFGPVTDVSVDIKCDK-GDLTRYQSDILSFTQ-MSEMFNQKSS 89
++LV++ +T P T V +K DK GD + S FT+ E+ K+
Sbjct: 201 DKLVLITDTIMPYPNTPASIKQTQVDYVVKVDKVGDPDKIGSGATRFTKDPKELKIAKTV 260
Query: 90 IPGKIPSGYFNTVFGFEAGSWASDAANTKFL 120
+ S YF F F+ GS + A T+FL
Sbjct: 261 NDVIVNSKYFKNDFSFQTGSGGAALAVTRFL 291
>VG30_BPPF3 (P03626) 32.7 kDa protein (ORF 301)
Length = 301
Score = 31.6 bits (70), Expect = 6.1
Identities = 28/113 (24%), Positives = 46/113 (39%), Gaps = 10/113 (8%)
Query: 374 PVTGMRLFLEGMKCNRLAIH-LQYLLNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKKF 432
PVT RL M+ +R H L ++ PT +++ I + R L+A S ++
Sbjct: 84 PVTDERL--TAMETHRHTGHDLVFITQAPTFVHHHIRKLVGLHIHLYRSRGLQAASKYEW 141
Query: 433 SHVCTAPVKYNPS-------WSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLF 478
SHVC +P W K+ T +H K + + + L+F
Sbjct: 142 SHVCDSPNDRKEQQRADFVLWKFPKEHFAFYTSAVMHTHKFKMPKKIGVLLVF 194
>SIR3_YEAST (P06701) Regulatory protein SIR3 (Silent information
regulator 3)
Length = 978
Score = 31.6 bits (70), Expect = 6.1
Identities = 17/80 (21%), Positives = 41/80 (51%)
Query: 112 SDAANTKFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILV 171
++ N L L+ +IK+ + D SK++ + + +W+P +++F +V
Sbjct: 873 TEGTNRHTLALETVLIKMVKMLRDNPGYKASKEIKKVICGAWEPAITIEKLKQFSWISVV 932
Query: 172 GLSIGGKDLVLVKQDVSSNL 191
+G K +V+V ++ S+++
Sbjct: 933 NDLVGEKLVVVVLEEPSASI 952
>FMB2_PORGI (Q51825) Major fimbrial subunit protein, type II
precursor (Fimbrillin) (Fimbrilin)
Length = 350
Score = 31.6 bits (70), Expect = 6.1
Identities = 15/51 (29%), Positives = 26/51 (50%)
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSW 110
I C KG LT++ LS +M+ FN + P+ Y+ + FE+ ++
Sbjct: 240 ILCIKGKLTKHDGTPLSSEEMTAAFNAGWIVANNDPTTYYPVLVNFESNNY 290
>YEP4_YEAST (P40041) Hypothetical 56.6 kDa protein in THO1-ICL1
intergenic region
Length = 505
Score = 31.2 bits (69), Expect = 8.0
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 7/103 (6%)
Query: 455 IVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSG-SSVFSAI 513
+VT V+++ RS L K++ V K + S+ GS S S SS
Sbjct: 94 LVTAAVQSVRRNRKRSKKKLLDSKKKIARGKVQKIPLSPPSSSNMGSCSASNASSSDEEA 153
Query: 514 STSIQGKDQKKPALVVVDSSV---FPTG---PPVPVQAQKLLK 550
S + + P+L + S +P G PPVP Q + LLK
Sbjct: 154 SVKEEPAEHALPSLNTITSQKLLPYPNGRTLPPVPTQVRSLLK 196
>FMA2_PORGI (Q51822) Major fimbrial subunit protein, type II
precursor (Fimbrillin) (Fimbrilin)
Length = 348
Score = 31.2 bits (69), Expect = 8.0
Identities = 16/58 (27%), Positives = 27/58 (45%)
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANT 117
I C KG LT++ LS +M+ FN + P+ Y+ + F + ++ D T
Sbjct: 238 ILCVKGKLTKHDGTPLSSEEMTAAFNAGWIVADNNPTTYYPVLVNFNSNNYTYDNGYT 295
>CX56_CHICK (P29415) Gap junction Cx56 protein (Connexin 56)
Length = 510
Score = 31.2 bits (69), Expect = 8.0
Identities = 17/66 (25%), Positives = 33/66 (49%), Gaps = 7/66 (10%)
Query: 478 FSKVSNAIVVKSNWTQGSTQRSGS--GSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVF 535
F+ SNA+ + NW + ++ G S +GSS S++ + +++ L+ +
Sbjct: 378 FNSKSNALAAEQNWVNMAAEQQGKAPSSSAGSSTPSSVRHPLPEQEEPLEQLLPL----- 432
Query: 536 PTGPPV 541
P GPP+
Sbjct: 433 PAGPPI 438
>AMYM_BACST (P19531) Maltogenic alpha-amylase precursor (EC
3.2.1.133) (Glucan 1,4-alpha-maltohydrolase)
Length = 719
Score = 31.2 bits (69), Expect = 8.0
Identities = 25/77 (32%), Positives = 38/77 (48%), Gaps = 3/77 (3%)
Query: 40 TERRELTVPGFGP-VTDVSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGY 98
+ R E+ VP +TDV V +L Y +ILS TQ S +F KS+ P +
Sbjct: 576 SNRIEVYVPNMAAGLTDVKVTAGGVSSNL--YSYNILSGTQTSVVFTVKSAPPTNLGDKI 633
Query: 99 FNTVFGFEAGSWASDAA 115
+ T E G+W++D +
Sbjct: 634 YLTGNIPELGNWSTDTS 650
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,039,314
Number of Sequences: 164201
Number of extensions: 3287090
Number of successful extensions: 7868
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 7860
Number of HSP's gapped (non-prelim): 17
length of query: 595
length of database: 59,974,054
effective HSP length: 116
effective length of query: 479
effective length of database: 40,926,738
effective search space: 19603907502
effective search space used: 19603907502
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0069.5


