

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0060.4
(1482 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
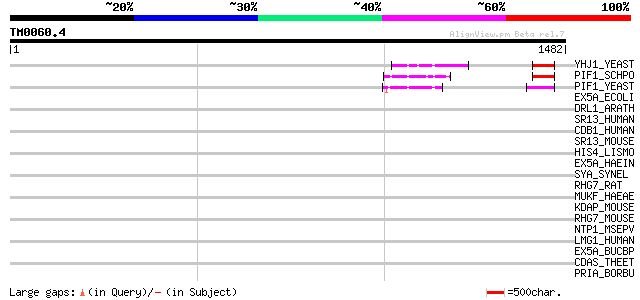
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 57 3e-07
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 54 4e-06
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 51 3e-05
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 42 0.016
DRL1_ARATH (P60838) Probable disease resistance protein At1g12280 36 0.69
SR13_HUMAN (Q9Y3M8) StAR-related lipid transfer protein 13 (StAR... 35 1.5
CDB1_HUMAN (Q9Y5F3) Protocadherin beta 1 precursor (PCDH-beta1) 35 1.5
SR13_MOUSE (Q923Q2) StAR-related lipid transfer protein 13 (StAR... 35 2.0
HIS4_LISMO (Q8Y9G4) 1-(5-phosphoribosyl)-5-[(5-phosphoribosylami... 35 2.0
EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 35 2.0
SYA_SYNEL (Q8DH56) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 34 2.6
RHG7_RAT (Q63744) Rho-GTPase-activating protein 7 (Rho-type GTPa... 34 3.4
MUKF_HAEAE (O30867) Chromosome partition protein mukF 34 3.4
KDAP_MOUSE (O09043) Kidney-derived aspartic protease-like protei... 34 3.4
RHG7_MOUSE (Q9R0Z9) Rho-GTPase-activating protein 7 (Rho-type GT... 33 4.5
NTP1_MSEPV (Q9YW39) Nucleoside triphosphatase I (EC 3.6.1.15) (N... 33 4.5
LMG1_HUMAN (P11047) Laminin gamma-1 chain precursor (Laminin B2 ... 33 4.5
EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 33 5.8
CDAS_THEET (P29964) Cyclomaltodextrinase (EC 3.2.1.54) (CDase) (... 33 5.8
PRIA_BORBU (Q45032) Primosomal protein N' (Replication factor Y) 33 7.6
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 57.4 bits (137), Expect = 3e-07
Identities = 64/217 (29%), Positives = 100/217 (45%), Gaps = 25/217 (11%)
Query: 1021 LFVYGYGGTGKTFLWTTLTYKLRS--EKKIILNVASSGIASLLLHGGRTAHSLFCIPL-N 1077
+F G GTGK+ + T+ +L S K+ I AS+G+A++ + GG T H I + N
Sbjct: 250 VFYTGSAGTGKSVILQTIIRQLSSLYGKESIAITASTGLAAVTI-GGSTLHKWSGIGIGN 308
Query: 1078 ADEDSCCGIVQGSPKAELL---KLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECF 1134
D +Q + +LL + + ++I DE MV + L++ R I +
Sbjct: 309 KTIDQLVKKIQS--QKDLLAAWRYTKVLIIDEISMVDGNLLDKLEQIARRIRKN------ 360
Query: 1135 NKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFC--KVLTLTENMRLFSN 1192
+ PFGG +VL GDF Q+ PV K V+ S +W+ C K + LT+ R N
Sbjct: 361 DDPFGGIQLVLTGDFFQLPPVAKKDEHN--VVKFCFESEMWKRCIQKTILLTKVFRQQDN 418
Query: 1193 SETSDVEKIKVFAEWVLDIGDGVLG-----DYNDGDA 1224
+ I+ + E +DI + DY DG A
Sbjct: 419 KLIDILNAIR-YGELTVDIAKTIRNLNRDIDYADGIA 454
Score = 46.2 bits (108), Expect = 7e-04
Identities = 23/59 (38%), Positives = 40/59 (66%), Gaps = 1/59 (1%)
Query: 1397 IKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKIL 1455
+ +R Q PL+L +A++I+K+QG+T+ + + L +F GQ+YVA+SR T L++L
Sbjct: 642 VGLERTQIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMDTLQVL 699
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 53.5 bits (127), Expect = 4e-06
Identities = 27/59 (45%), Positives = 42/59 (70%), Gaps = 1/59 (1%)
Query: 1397 IKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKIL 1455
++ R Q PL+L++A++I+K+QG+TL V + L VF GQ YVA+SR T+ GL++L
Sbjct: 704 VQASRSQIPLILAYAISIHKAQGQTLDRVKVDL-GRVFEKGQAYVALSRATTQEGLQVL 761
Score = 49.7 bits (117), Expect = 6e-05
Identities = 54/186 (29%), Positives = 87/186 (46%), Gaps = 24/186 (12%)
Query: 998 LNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWT----TLTYKLRSEKKIILNVA 1053
L+ Q +I + ++ +S +F G GTGK+ L L K R + + A
Sbjct: 310 LSDEQKRILDMVVEQQHS-----IFFTGSAGTGKSVLLRKIIEVLKSKYRKQSDRVAVTA 364
Query: 1054 SSGIASLLLHGGRTAHSLFCIPLNADE-DSCCGIVQGSPKA--ELLKLSSLIIWDEAPMV 1110
S+G+A+ + GG T HS + L + D ++ + K L+ LII DE MV
Sbjct: 365 STGLAACNI-GGVTLHSFAGVGLARESVDLLVSKIKKNKKCVNRWLRTRVLII-DEVSMV 422
Query: 1111 SRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATIN 1170
E +D+ L ++ R+ + +KPFGG +VL GDF Q+ PV G ++
Sbjct: 423 DA---ELMDK-LEEVARVIRKD--SKPFGGIQLVLTGDFFQLPPVPENGKESKFCF---- 472
Query: 1171 SSRLWR 1176
S+ W+
Sbjct: 473 ESQTWK 478
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 50.8 bits (120), Expect = 3e-05
Identities = 31/77 (40%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Query: 1379 RQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQ 1438
R V + D D + R Q PL+L+++++I+KSQG+TL V + L VF GQ
Sbjct: 669 RMVLVEPEDWAIEDENEKPLVSRVQLPLMLAWSLSIHKSQGQTLPKVKVDL-RRVFEKGQ 727
Query: 1439 LYVAISRVKTRAGLKIL 1455
YVA+SR +R GL++L
Sbjct: 728 AYVALSRAVSREGLQVL 744
Score = 47.0 bits (110), Expect = 4e-04
Identities = 49/172 (28%), Positives = 77/172 (44%), Gaps = 20/172 (11%)
Query: 995 LNKLNGGQAKI-------YEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRS--E 1045
LN NG + KI E II ++ G +F G GTGK+ L + L+
Sbjct: 223 LNPHNGVKVKIPICLSKEQESIIKL--AENGHNIFYTGSAGTGKSILLREMIKVLKGIYG 280
Query: 1046 KKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSL--II 1103
++ + AS+G+A+ + GG T HS I D D V + L + ++ ++
Sbjct: 281 RENVAVTASTGLAACNI-GGITIHSFAGILGKGDADKLYKKVGRRSRKHLRRWENIGALV 339
Query: 1104 WDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPV 1155
DE M+ + LD R I + ++PFGG ++ GDF Q+ PV
Sbjct: 340 VDEISMLDAELLDKLDFIARKIRKN------HQPFGGIQLIFCGDFFQLPPV 385
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.11.5)
(Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 41.6 bits (96), Expect = 0.016
Identities = 39/121 (32%), Positives = 57/121 (46%), Gaps = 22/121 (18%)
Query: 1335 EGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGS 1394
EG PVM+ RN D + GL NG +G + G G +V+ MP DG+
Sbjct: 478 EGRPVMIARN-DSALGLFNGD-----------IGIALDRGQ--GTRVWFA----MP-DGN 518
Query: 1395 MPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPN---PVFSHGQLYVAISRVKTRAG 1451
+ R ++AMT++KSQG H L LP+ PV + +Y A++R + R
Sbjct: 519 IKSVQPSRLPEHETTWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRRLS 578
Query: 1452 L 1452
L
Sbjct: 579 L 579
>DRL1_ARATH (P60838) Probable disease resistance protein At1g12280
Length = 894
Score = 36.2 bits (82), Expect = 0.69
Identities = 24/75 (32%), Positives = 39/75 (52%), Gaps = 5/75 (6%)
Query: 976 LFNELNFDTVEMS----KLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGK 1031
L ++ +FDTV ++ ++ + GQ + E + + + DG E + +YG GG GK
Sbjct: 130 LSSQGDFDTVTLATPIARIEEMPIQPTIVGQETMLERVWTRLTEDGDEIVGLYGMGGVGK 189
Query: 1032 TFLWTTLTYKLRSEK 1046
T L T + K SEK
Sbjct: 190 TTLLTRINNKF-SEK 203
>SR13_HUMAN (Q9Y3M8) StAR-related lipid transfer protein 13
(StARD13) (START domain-containing protein 13) (46H23.2)
Length = 995
Score = 35.0 bits (79), Expect = 1.5
Identities = 34/127 (26%), Positives = 58/127 (44%), Gaps = 12/127 (9%)
Query: 433 LPSSFTGGRRYMFNNCQDAMGICREYGYPD-LFLTMTCNPKWPEIERHVSARCLSAYDRP 491
LP S RY+ +NC D +G+ R+ G + N +PE +V+ SAYD
Sbjct: 558 LPQSIQQALRYLRSNCLDQVGLFRKSGVKSRIHALRQMNENFPE---NVNYEDQSAYDVA 614
Query: 492 DLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIE 551
D+ + FR + L N F + Y + Q+ A ILL L+ +++ E
Sbjct: 615 DMVKQFFRDLPEPLFTNKLSETFLH---IYQYVSKEQRLQAVQAAILL-LADENR----E 666
Query: 552 LIDSVIC 558
++ +++C
Sbjct: 667 VLQTLLC 673
>CDB1_HUMAN (Q9Y5F3) Protocadherin beta 1 precursor (PCDH-beta1)
Length = 818
Score = 35.0 bits (79), Expect = 1.5
Identities = 23/87 (26%), Positives = 38/87 (43%), Gaps = 2/87 (2%)
Query: 974 ILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTF 1033
IL E N V + K+H E L+ G A+I ++ N D F ++ G GK +
Sbjct: 458 ILTVRENNSPAVFIGKVHAEDLDL--GENAQITYSLLPPKNGDLSVFAYISINSGNGKLY 515
Query: 1034 LWTTLTYKLRSEKKIILNVASSGIASL 1060
T+ Y+ + + ++ G SL
Sbjct: 516 ALRTMDYEAIQDFQFVVKATDGGFLSL 542
>SR13_MOUSE (Q923Q2) StAR-related lipid transfer protein 13
(StARD13) (START domain-containing protein 13)
Length = 1113
Score = 34.7 bits (78), Expect = 2.0
Identities = 34/127 (26%), Positives = 58/127 (44%), Gaps = 12/127 (9%)
Query: 433 LPSSFTGGRRYMFNNCQDAMGICREYGYPD-LFLTMTCNPKWPEIERHVSARCLSAYDRP 491
LP S RY+ +NC D +G+ R+ G + N +P+ +VS SAYD
Sbjct: 676 LPQSIQQALRYLRSNCLDQVGLFRKSGVKSRIHALRQMNENFPD---NVSYEDQSAYDVA 732
Query: 492 DLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIE 551
D+ + FR + L N F + Y + Q+ A ILL L+ +++ E
Sbjct: 733 DMVKQFFRDLPEPLFTNKLSETFLH---IYQYVPKEQRLQAVQAAILL-LADENR----E 784
Query: 552 LIDSVIC 558
++ +++C
Sbjct: 785 VLQTLLC 791
>HIS4_LISMO (Q8Y9G4)
1-(5-phosphoribosyl)-5-[(5-
phosphoribosylamino)methylideneamino]
imidazole-4-carboxamide isomerase (EC 5.3.1.16)
(Phosphoribosylformimino-5-aminoimidazole carboxamide
ribotide isomerase)
Length = 240
Score = 34.7 bits (78), Expect = 2.0
Identities = 26/92 (28%), Positives = 43/92 (46%), Gaps = 3/92 (3%)
Query: 593 MKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFV-VPYNPKLL-- 649
+KNG+C + F +F +KT + D + T +T VDLD P N +++
Sbjct: 9 LKNGQCVRLFQGDFSKKTVVNEDPIAQAKAFATDGATYLHIVDLDGALEGRPINLEVIQK 68
Query: 650 MKYQAHINIEYCNKSNCIKYLFKYINKGVDRV 681
MK A I ++ + + Y+ G+DRV
Sbjct: 69 MKITAKIPVQVGGGIRSMAQVDYYLESGIDRV 100
>EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.11.5)
Length = 640
Score = 34.7 bits (78), Expect = 2.0
Identities = 17/42 (40%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Query: 1409 SFAMTINKSQGKTLSHVGLYLP---NPVFSHGQLYVAISRVK 1447
+F MTI+KSQG H + LP NPV S ++ ++R K
Sbjct: 565 AFMMTIHKSQGSEFKHTVMVLPTEVNPVLSRELVFTGVTRAK 606
>SYA_SYNEL (Q8DH56) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 882
Score = 34.3 bits (77), Expect = 2.6
Identities = 35/149 (23%), Positives = 65/149 (43%), Gaps = 20/149 (13%)
Query: 474 PEIERHVSA-RCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFF-GQAISWMYTIEFQKRG 531
P++ R +A +CL D ++ +++ N G +F G+AI+W + + G
Sbjct: 57 PKVPRATTAQKCLRTNDIENVGRTARHHTFFEMLGNFSFGDYFKGEAIAWAWELMTTVYG 116
Query: 532 LPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSP 591
LP +L+ + D + ++ + LP ++ + + SN+ GP G P
Sbjct: 117 LPPERLLVSVFENDD-EAYDIWHRQV--GLPKERIQRMGEE--SNFWTAGPTG------P 165
Query: 592 CMKNGRCSK----FFPKNFVEKTSFDSDG 616
C G CS+ F+P+ + D DG
Sbjct: 166 C---GPCSEIYYDFYPEKGLANVDLDDDG 191
>RHG7_RAT (Q63744) Rho-GTPase-activating protein 7 (Rho-type
GTPase-activating protein 7) (p122-RhoGAP) (Deleted in
liver cancer 1 protein homolog) (Dlc-1) (StAR-related
lipid transfer protein 12) (StARD12) (START
domain-containing protein 12)
Length = 1091
Score = 33.9 bits (76), Expect = 3.4
Identities = 38/150 (25%), Positives = 68/150 (45%), Gaps = 7/150 (4%)
Query: 433 LPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPD 492
LP S RY+ N+C D +G+ R+ G + E +V+ SAYD D
Sbjct: 654 LPQSIQQAMRYLRNHCLDQVGLFRKSGVKSRIQAL--RQMNESAEDYVNYEGQSAYDVAD 711
Query: 493 LACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQK-RGLPHAHILLWLSTKDKLHSIE 551
+ + FR + LM N K + F Q + Y + Q+ + + A +LL ++ L ++
Sbjct: 712 MLKQYFRDLPEPLMTN-KLSETFLQI--YQYVPKDQRLQAIKAAIMLLPDENREVLQTLL 768
Query: 552 LIDSVICAELPDPKVYPL-LYQCVSNYMVH 580
S + A + + ++ P L C++ + H
Sbjct: 769 YFLSHVTAAVKENQMTPTNLAVCLAPSLFH 798
>MUKF_HAEAE (O30867) Chromosome partition protein mukF
Length = 443
Score = 33.9 bits (76), Expect = 3.4
Identities = 21/98 (21%), Positives = 42/98 (42%), Gaps = 19/98 (19%)
Query: 233 EFNVLAKSFRRVRDHVQEANSNQVALRLFRHRVNDPKTYNLPTV---------------- 276
+ +++A +R D +E N + +R V P Y++ +
Sbjct: 127 QLSIVADEIQRASDSAEEGVENNESEHFWRRNVFAPLKYSVAEIFDSIDLSQRIMDENQQ 186
Query: 277 ---DEVAALIVEDFDTSDCGRDIILRTSSGNLQRIYDT 311
DE+A L+ +D+ + + +L +SGNL+ + DT
Sbjct: 187 SIKDEIAELLTKDWQAAISSCERLLDETSGNLRELQDT 224
>KDAP_MOUSE (O09043) Kidney-derived aspartic protease-like protein
precursor (EC 3.4.23.-) (KDAP-1) (KAP) (Napsin)
Length = 419
Score = 33.9 bits (76), Expect = 3.4
Identities = 33/122 (27%), Positives = 50/122 (40%), Gaps = 16/122 (13%)
Query: 8 LNMMVEWETSSEGSRSNTVHSNPDFCSGQELLPQQRNVSDCTNNFNTNFAFVEAESQMYF 67
LN + WE +E SR++T NP F + + Q + NF V F
Sbjct: 38 LNPLNGWEQLAELSRTSTSGGNPSFVPLSKFMNTQYFGTIGLGTPPQNFTVV-------F 90
Query: 68 DLGEMNM-----ACQYCGAILWYHERAQ-KAKNAISPD---FSICCMKGKISLPYLQDAP 118
D G N+ C + W+H R KA ++ P+ F+I G++S QD
Sbjct: 91 DTGSSNLWVPSTRCHFFSLACWFHHRFNPKASSSFRPNGTKFAIQYGTGRLSGILSQDNL 150
Query: 119 TL 120
T+
Sbjct: 151 TI 152
>RHG7_MOUSE (Q9R0Z9) Rho-GTPase-activating protein 7 (Rho-type
GTPase-activating protein 7) (Deleted in liver cancer 1
protein homolog) (Dlc-1) (StAR-related lipid transfer
protein 12) (StARD12) (START domain-containing protein
12)
Length = 1092
Score = 33.5 bits (75), Expect = 4.5
Identities = 38/150 (25%), Positives = 68/150 (45%), Gaps = 7/150 (4%)
Query: 433 LPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPD 492
LP S RY+ N+C D +G+ R+ G + E +V+ SAYD D
Sbjct: 655 LPQSIQQAMRYLRNHCLDQVGLFRKSGVKSRIQAL--RQMNESAEDNVNYEGQSAYDVAD 712
Query: 493 LACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQK-RGLPHAHILLWLSTKDKLHSIE 551
+ + FR + LM N K + F Q + Y + Q+ + + A +LL ++ L ++
Sbjct: 713 MLKQYFRDLPEPLMTN-KLSETFLQI--YQYVPKDQRLQAIKAAIMLLPDENREVLQTLL 769
Query: 552 LIDSVICAELPDPKVYPL-LYQCVSNYMVH 580
S + A + + ++ P L C++ + H
Sbjct: 770 YFLSDVTAAVKENQMTPTNLAVCLAPSLFH 799
>NTP1_MSEPV (Q9YW39) Nucleoside triphosphatase I (EC 3.6.1.15)
(Nucleoside triphosphate phosphohydrolase I) (NPH I)
Length = 647
Score = 33.5 bits (75), Expect = 4.5
Identities = 17/58 (29%), Positives = 31/58 (53%), Gaps = 1/58 (1%)
Query: 659 EYCNKSNCIKYLFKYINKGVDRVTMSMSVTQSSNEETTVVDEIKQYHDCRYLSPCEAV 716
E N+ I FK+ N+ +D + S + + + E + D+IKQY C+Y+ C+ +
Sbjct: 328 EKLNEFKSIIGNFKFSNEFIDIFRNNDSFSNAKSSEIEIFDKIKQY-SCKYIEACKII 384
>LMG1_HUMAN (P11047) Laminin gamma-1 chain precursor (Laminin B2
chain)
Length = 1609
Score = 33.5 bits (75), Expect = 4.5
Identities = 17/56 (30%), Positives = 27/56 (47%)
Query: 540 WLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKN 595
W++ D ++ +++ DPKV Y +S++ V G C N S CMKN
Sbjct: 243 WVTATDIRVTLNRLNTFGDEVFNDPKVLKSYYYAISDFAVGGRCKCNGHASECMKN 298
>EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC 3.1.11.5)
Length = 618
Score = 33.1 bits (74), Expect = 5.8
Identities = 20/66 (30%), Positives = 36/66 (54%), Gaps = 6/66 (9%)
Query: 1409 SFAMTINKSQGKTLSHVGLYLP---NPVFSHGQLYVAISRVKTRAGLKILICNEDISQRD 1465
++ MT++KSQG S V L LP + + +Y A++R K + + +E+I +
Sbjct: 543 NWTMTVHKSQGSEFSEVVLILPTIMTSILTKELIYTAVTRSKKKL---TIYSDENIFIKS 599
Query: 1466 VTKNIV 1471
+ KNI+
Sbjct: 600 LKKNII 605
>CDAS_THEET (P29964) Cyclomaltodextrinase (EC 3.2.1.54) (CDase)
(Cyclomaltodextrin hydrolase, decycling)
Length = 574
Score = 33.1 bits (74), Expect = 5.8
Identities = 21/79 (26%), Positives = 41/79 (51%), Gaps = 2/79 (2%)
Query: 702 KQYHDCRYLSPCE--AVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERN 759
++ + +Y S C A+ R F+F K ++ + K++VI NE +D+L+
Sbjct: 495 RENEELKYGSFCTLYAIGRVFAFKREYKGKSIIVVLNNSSKQEVIFLNEVEGKEDILKMK 554
Query: 760 KVKKSMFLAWMEANCKYPL 778
++K+S L +++ N Y L
Sbjct: 555 ELKRSGNLLYLQPNSAYIL 573
>PRIA_BORBU (Q45032) Primosomal protein N' (Replication factor Y)
Length = 660
Score = 32.7 bits (73), Expect = 7.6
Identities = 21/60 (35%), Positives = 35/60 (58%), Gaps = 5/60 (8%)
Query: 991 HVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTL-TYKLRSEKKII 1049
H +CL +LN Q IY+EII S+ +++G G+GKT ++ L Y L E++++
Sbjct: 128 HKKCL-ELNNEQQNIYKEII---GSEKTNVFYLFGIPGSGKTEIFIKLCEYYLALEQQVL 183
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 180,606,537
Number of Sequences: 164201
Number of extensions: 8062383
Number of successful extensions: 16976
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 16959
Number of HSP's gapped (non-prelim): 32
length of query: 1482
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1359
effective length of database: 39,777,331
effective search space: 54057392829
effective search space used: 54057392829
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0060.4


