

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0400a.3
(419 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
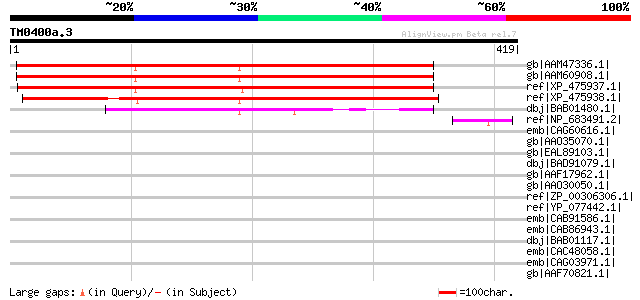
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM47336.1| AT3g27930/K24A2_2 [Arabidopsis thaliana] gi|18087... 449 e-125
gb|AAM60908.1| unknown [Arabidopsis thaliana] 446 e-124
ref|XP_475937.1| unknown protein [Oryza sativa (japonica cultiva... 415 e-114
ref|XP_475938.1| unknown protein [Oryza sativa (japonica cultiva... 367 e-100
dbj|BAB01480.1| unnamed protein product [Arabidopsis thaliana] 232 2e-59
ref|NP_683491.2| endonuclease/exonuclease/phosphatase family pro... 48 6e-04
emb|CAG60616.1| unnamed protein product [Candida glabrata CBS138... 37 1.1
gb|AAO35070.1| hypothetical protein [Clostridium tetani E88] gi|... 37 1.5
gb|EAL89103.1| leucyl-tRNA synthetase [Aspergillus fumigatus Af293] 35 4.3
dbj|BAD91079.1| beta-D-galactosidase [Pyrus pyrifolia] 35 4.3
gb|AAF17962.1| gp028L [Rabbit fibroma virus] gi|9633889|ref|NP_0... 35 4.3
gb|AAO30050.1| unknown protein [Arabidopsis thaliana] gi|2026044... 35 5.7
ref|ZP_00306306.1| COG1571: Predicted DNA-binding protein contai... 34 7.4
ref|YP_077442.1| Polysaccharide deacetylase,Carbohydrate Esteras... 34 7.4
emb|CAB91586.1| putative protein [Arabidopsis thaliana] gi|15231... 34 7.4
emb|CAB86943.1| putative protein [Arabidopsis thaliana] gi|11357... 34 7.4
dbj|BAB01117.1| unnamed protein product [Arabidopsis thaliana] 34 7.4
emb|CAC48058.1| putative lyase [Rhizobium leguminosarum bv. viciae] 34 9.6
emb|CAG03971.1| unnamed protein product [Tetraodon nigroviridis] 34 9.6
gb|AAF70821.1| beta-galactosidase [Lycopersicon esculentum] 34 9.6
>gb|AAM47336.1| AT3g27930/K24A2_2 [Arabidopsis thaliana] gi|18087557|gb|AAL58910.1|
AT3g27930/K24A2_2 [Arabidopsis thaliana]
gi|18405582|ref|NP_566828.1| expressed protein
[Arabidopsis thaliana]
Length = 425
Score = 449 bits (1155), Expect = e-125
Identities = 222/367 (60%), Positives = 271/367 (73%), Gaps = 22/367 (5%)
Query: 6 VKPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTTRIM 65
VK PPP VVLVPPLFD+PPL+AR RMLESSY+++FGKLALRCLF DYF+ FT + +
Sbjct: 6 VKKEPPPPVVLVPPLFDYPPLSARTRMLESSYNLLFGKLALRCLFEDYFEEANRFTGKFL 65
Query: 66 FKPIDEPHVDLIATVSGPLDHKPEESITGNALFRWQR------------------ILRMR 107
KP D+PHVDL+A+VSG +D + E GNA FRWQ +L MR
Sbjct: 66 LKPTDDPHVDLVASVSGAVDGRVEGDFVGNAEFRWQSDVDDPHTFVDLSVSTSNPVLLMR 125
Query: 108 SCAYYPRYGFGAFGVFPLLLR-KREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKS 166
S AYYP+YG GAF V+PL+ + + SE+Y +MGLRYGS NLS G T+ PF+ +ELPK
Sbjct: 126 SSAYYPKYGIGAFAVYPLISKITGKSSEEYRIMGLRYGSTNLSVGATVTPFSANNELPKH 185
Query: 167 AWLVSKMGRLTAGVQYEPQHGN---AKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVK 223
AWLVSKMG LT GVQYEP HG+ AK ++ NWSCA YGVGS SPL+PSFN +EL +
Sbjct: 186 AWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQSPLTPSFNIGIELAR 245
Query: 224 SSQFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRI 283
SSQF+ASFYQH+VVQRRV+NP EEN V+GITNYIDFGFELQ+ VDD+ N DS ++
Sbjct: 246 SSQFIASFYQHVVVQRRVQNPFEENQVVGITNYIDFGFELQSRVDDSKTPPNAPDSLLQV 305
Query: 284 GASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSES 343
ASWQANKNFLLK KVG S+++LAFKSWWKPSF F+ISAT + G +Q GFG++ ++
Sbjct: 306 AASWQANKNFLLKGKVGAHSSTLSLAFKSWWKPSFAFNISATTNHRTGNVQCGFGLRVDN 365
Query: 344 LREASFQ 350
LREAS+Q
Sbjct: 366 LREASYQ 372
>gb|AAM60908.1| unknown [Arabidopsis thaliana]
Length = 425
Score = 446 bits (1147), Expect = e-124
Identities = 220/367 (59%), Positives = 270/367 (72%), Gaps = 22/367 (5%)
Query: 6 VKPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTTRIM 65
VK PPP VVLVPPLFD+PPL+AR RMLESSY+++FGKLALRCLF DYF+ FT + +
Sbjct: 6 VKKEPPPPVVLVPPLFDYPPLSARTRMLESSYNLLFGKLALRCLFEDYFEEANRFTAKFL 65
Query: 66 FKPIDEPHVDLIATVSGPLDHKPEESITGNALFRWQR------------------ILRMR 107
KP D+PHVDL+A+VSG +D + E GNA FRWQ +L MR
Sbjct: 66 LKPTDDPHVDLVASVSGAVDGRVEGDFVGNAEFRWQSDVDDPHTFVDLSVSTSNPVLLMR 125
Query: 108 SCAYYPRYGFGAFGVFPLLLR-KREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKS 166
S AYYP+YG GAF V+PL+ + + SE+Y +MGLRYGS LS G T+ PF+ +ELPK
Sbjct: 126 SSAYYPKYGIGAFAVYPLISKITGKSSEEYRIMGLRYGSTXLSVGATVTPFSANNELPKH 185
Query: 167 AWLVSKMGRLTAGVQYEPQHGN---AKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVK 223
AWLVSKMG LT GVQYEP HG+ AK ++ NWSCA YGVGS SPL+PSFN +EL +
Sbjct: 186 AWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQSPLTPSFNMGIELAR 245
Query: 224 SSQFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRI 283
SSQF+ASFYQH+VVQR+V+NP EEN V+GITNYIDFGFELQ+ VDD+ N DS ++
Sbjct: 246 SSQFIASFYQHVVVQRKVQNPFEENQVVGITNYIDFGFELQSRVDDSKTPPNAPDSLLQV 305
Query: 284 GASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSES 343
ASWQANKNFLLK KVG S+++LAFKSWWKPSF F+ISAT + G +Q GFG++ ++
Sbjct: 306 AASWQANKNFLLKGKVGAHSSTLSLAFKSWWKPSFAFNISATTNHRTGNVQCGFGLRVDN 365
Query: 344 LREASFQ 350
LREAS+Q
Sbjct: 366 LREASYQ 372
>ref|XP_475937.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900285|gb|AAT39153.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 424
Score = 415 bits (1067), Expect = e-114
Identities = 203/365 (55%), Positives = 261/365 (70%), Gaps = 21/365 (5%)
Query: 7 KPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTTRIMF 66
K PPP +VLVPPLFD+PP+AAR RM +Y+++FGKL+L+ LF DYF + T+R+M
Sbjct: 7 KAEPPPPMVLVPPLFDYPPIAARTRMSVPAYELMFGKLSLQNLFEDYFDHAGNMTSRVML 66
Query: 67 KPIDEPHVDLIATVSGPLDHKPEESITGNALFRWQR------------------ILRMRS 108
KP+++PHVDLIATVS D + G+ALFRWQ+ +L++RS
Sbjct: 67 KPLEDPHVDLIATVSAAADKIDGTQVKGDALFRWQKDLDDPHTFVDLLVSTSNPMLQVRS 126
Query: 109 CAYYPRYGFGAFGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKSAW 168
CAY+P+Y GAFG FPLL+ R SEDYG+MG+RYGS NLSFG + +PF ELP AW
Sbjct: 127 CAYHPKYRVGAFGTFPLLMGNRVRSEDYGVMGVRYGSENLSFGSSFVPFPGSAELPSGAW 186
Query: 169 LVSKMGRLTAGVQYEPQHGNAKL---SNLMNWSCAMAYGVGSGSPLSPSFNFSLELVKSS 225
LV + G L+AGVQY+P GN L ++ NW+CA++YGVG SPLSPSF FSLEL +S+
Sbjct: 187 LVGRKGSLSAGVQYKPLSGNKHLMPYTDWKNWNCAISYGVGLTSPLSPSFIFSLELARST 246
Query: 226 QFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRIGA 285
+F+ASFYQHMVVQRRVKNP E++ ++GITNYIDFG EL T +D + + ++S F+ A
Sbjct: 247 EFIASFYQHMVVQRRVKNPFEDDQIVGITNYIDFGLELATRIDKDKPSESANNSLFQFAA 306
Query: 286 SWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSESLR 345
SWQANKNFL K K+GP SS+ALAFKSWW+PSFTFS++A D G YGFGI+ E LR
Sbjct: 307 SWQANKNFLFKGKLGPSKSSVALAFKSWWRPSFTFSVTAVNDHLKGTRSYGFGIRVEDLR 366
Query: 346 EASFQ 350
+ S+Q
Sbjct: 367 QPSYQ 371
>ref|XP_475938.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900286|gb|AAT39154.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 414
Score = 367 bits (941), Expect = e-100
Identities = 191/367 (52%), Positives = 244/367 (66%), Gaps = 32/367 (8%)
Query: 11 PPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTT-RIMFKPI 69
PP +VLVPP F FP AAR RM +Y+V+FGKL R LF+DYF T+ IM +P+
Sbjct: 8 PPAMVLVPPPFTFPAAAARTRMAVPAYEVMFGKLQRRSLFDDYFDQVGSITSGMIMLRPL 67
Query: 70 DEPHVDLIATVSGPLDHKPEESITGNALFRWQRIL------------------RMRSCAY 111
+ HVDL A ++ + G ALFRWQR L ++RSCAY
Sbjct: 68 VDSHVDLTAKMT---------TTGGEALFRWQRYLDDPNTFMDLHLSTPKPMVQLRSCAY 118
Query: 112 YPRYGFGAFGVFPLLLRKREFSE-DYGLMGLRYGSGNLSFGVTLMPFAKKDELPKSAWLV 170
YP+Y GAFG FPLL R+ SE DYG+MGLRYGS NLS G + +PFA ++P AWLV
Sbjct: 119 YPKYRIGAFGTFPLLKANRDCSEGDYGIMGLRYGSENLSIGASFLPFALSGQVPYGAWLV 178
Query: 171 SKMGRLTAGVQYEPQHGN---AKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVKSSQF 227
+ G ++AG+QY+P + L++L NW+CA++YG+GS SPLSPSFNFSLELV+++Q
Sbjct: 179 GRKGNISAGIQYKPLCESMHPVPLTDLKNWNCAISYGMGSTSPLSPSFNFSLELVRNTQL 238
Query: 228 VASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRIGASW 287
VASFYQH VVQRRV NP EE +IG TN++DFG EL TS+D A N S+ +F++ ASW
Sbjct: 239 VASFYQHFVVQRRVMNPREEEHIIGTTNFVDFGLELATSLDKDKAKENASNPSFQVAASW 298
Query: 288 QANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSESLREA 347
QA+KNFL+K K+GP SSMALA KSWW+P FTFS +A D G YGFGI E L+E
Sbjct: 299 QASKNFLVKGKLGPSKSSMALAMKSWWRPFFTFSFTAMYDHLKGTGSYGFGISIEDLKEP 358
Query: 348 SFQSIFT 354
S+Q ++
Sbjct: 359 SYQMAYS 365
>dbj|BAB01480.1| unnamed protein product [Arabidopsis thaliana]
Length = 351
Score = 232 bits (591), Expect = 2e-59
Identities = 132/278 (47%), Positives = 166/278 (59%), Gaps = 46/278 (16%)
Query: 80 VSGPLDHKPEESITGNALFRWQRILRMRSCAYYPRYGFGAFGVFPLLLR-KREFSEDYGL 138
VSG +D + E GNA FRWQR+L MRS AYYP+YG GAF V+PL+ + + SE+Y +
Sbjct: 60 VSGAVDGRVEGDFVGNAEFRWQRVLLMRSSAYYPKYGIGAFAVYPLISKITGKSSEEYRI 119
Query: 139 MGLRYGSGNLSFGVTLMPFAKKDELPKSAWLVSKMGRLTAGVQYEPQHGN---AKLSNLM 195
MGLRYGS NLS G T+ PF+ +ELPK AWLVSKMG LT GVQYEP HG+ AK ++
Sbjct: 120 MGLRYGSTNLSVGATVTPFSANNELPKHAWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPR 179
Query: 196 NWSCAMAYGVGSGSPLSPSFNFSLELVKSSQFVASFYQH---MVVQRRVKNPLEENGVIG 252
NWSCA YGVGS SPL+PSFN +EL +SSQF S Q+ ++ +V+NP EEN V+G
Sbjct: 180 NWSCAAGYGVGSQSPLTPSFNIGIELARSSQFSTSLAQYEIDLMDLEQVQNPFEENQVVG 239
Query: 253 ITNYIDFGFELQTSVDDAIATNNISDSTFRIGASWQANKNFLLKAKVGPRISSMALAFKS 312
ITNYIDFGFELQ+ F G A + L
Sbjct: 240 ITNYIDFGFELQSRF-------------FIAGGCVLAGQQELF----------------- 269
Query: 313 WWKPSFTFSISATRDRADGQIQYGFGIQSESLREASFQ 350
AT + G +Q GFG++ ++LREAS+Q
Sbjct: 270 ---------AEATTNHRTGNVQCGFGLRVDNLREASYQ 298
>ref|NP_683491.2| endonuclease/exonuclease/phosphatase family protein [Arabidopsis
thaliana]
Length = 454
Score = 47.8 bits (112), Expect = 6e-04
Identities = 29/86 (33%), Positives = 36/86 (41%), Gaps = 37/86 (43%)
Query: 367 KTVVVSYNILGVDSASKHQDLYSNIRHS-------------------------------- 394
K V+VSYN+LGVD+AS H DLY N+
Sbjct: 99 KLVLVSYNLLGVDNASNHMDLYYNVPRKHLEWSRRKHLICKEISRYNASILCLQEVDRFD 158
Query: 395 -----FQNNGFKGVGKDYTGDARDGC 415
+N GF+GV K TG+A DGC
Sbjct: 159 DLDVLLKNRGFRGVHKSRTGEASDGC 184
>emb|CAG60616.1| unnamed protein product [Candida glabrata CBS138]
gi|50290495|ref|XP_447679.1| unnamed protein product
[Candida glabrata]
Length = 501
Score = 37.0 bits (84), Expect = 1.1
Identities = 25/87 (28%), Positives = 40/87 (45%), Gaps = 5/87 (5%)
Query: 223 KSSQFVASFYQHMVVQRRVKNPLEE----NGVIGITNYIDFGFELQTSVDDAIATNN-IS 277
K S V F + ++ ++ P E N ++G N D G E+Q S+++ + N I
Sbjct: 374 KPSILVLPFGTYGILTGSIRKPQEHMIMRNDILGYKNIDDEGNEIQKSIEEVVLNNKYIH 433
Query: 278 DSTFRIGASWQANKNFLLKAKVGPRIS 304
S F +G W + LK +V P S
Sbjct: 434 FSDFPLGKPWAYSSFDQLKCRVDPASS 460
>gb|AAO35070.1| hypothetical protein [Clostridium tetani E88]
gi|28210189|ref|NP_781133.1| hypothetical protein
CTC00436 [Clostridium tetani E88]
Length = 260
Score = 36.6 bits (83), Expect = 1.5
Identities = 46/161 (28%), Positives = 72/161 (44%), Gaps = 14/161 (8%)
Query: 215 FNFSLELVKSS-QFVASFYQHMVVQRRV-KNPLEENGVIGITNYIDFGFELQTSVDDAIA 272
+NF + K++ QF+ + + VV KNPL+EN +I +T +L+ DA+
Sbjct: 86 WNFIFNVPKANKQFIITIHDGQVVNYNTSKNPLDENDLIDLT-------QLKVDSTDAV- 137
Query: 273 TNNISDSTFRIGASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRD--RAD 330
I + + G SW NF+L AK+ +I + S S T D A
Sbjct: 138 KKAILEYNLKPGLSWARGYNFVL-AKIDGKIILTVVGNDKDGNFSRININSTTGDIVSAI 196
Query: 331 GQIQYGFGIQSESLREASFQSIFTILNLSPPIADKSKTVVV 371
+I G G+ + + FQS TIL +DK+ T+ V
Sbjct: 197 HKISTGGGLFRNNSQLPIFQSQTTILT-GASYSDKNGTIGV 236
>gb|EAL89103.1| leucyl-tRNA synthetase [Aspergillus fumigatus Af293]
Length = 1127
Score = 35.0 bits (79), Expect = 4.3
Identities = 28/99 (28%), Positives = 41/99 (41%), Gaps = 4/99 (4%)
Query: 183 EPQHGNAK-LSNLMNWSCAMAYGVGSGSPLSPSF---NFSLELVKSSQFVASFYQHMVVQ 238
E +HG K L+ L W+CA YG+GS P P F + S V + + + Y H
Sbjct: 602 ETKHGFEKNLTWLNRWACARTYGLGSKLPWDPKFLVESLSDSTVYMAYYTVAHYLHGDRY 661
Query: 239 RRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNIS 277
PL D+ F + D+ I+ + IS
Sbjct: 662 GNTPGPLNIKPEQMTDEVWDYIFTRRELSDELISKSGIS 700
>dbj|BAD91079.1| beta-D-galactosidase [Pyrus pyrifolia]
Length = 903
Score = 35.0 bits (79), Expect = 4.3
Identities = 28/98 (28%), Positives = 44/98 (44%), Gaps = 15/98 (15%)
Query: 120 FGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKD-ELPKSAWLVSKMGRLTA 178
FG FP+ LR + G+ + + N F + F KK +L + L+S G
Sbjct: 134 FGGFPVWLRD--------IPGIEFRTNNALFKEEMQRFVKKMVDLMQEEELLSWQGGPII 185
Query: 179 GVQYEPQHGNA------KLSNLMNWSCAMAYGVGSGSP 210
+Q E ++GN K + W+ MA G+G+G P
Sbjct: 186 MMQIENEYGNIEGQFGQKGKEYIKWAAEMALGLGAGVP 223
>gb|AAF17962.1| gp028L [Rabbit fibroma virus] gi|9633889|ref|NP_051917.1| gp028L
[Rabbit fibroma virus]
Length = 721
Score = 35.0 bits (79), Expect = 4.3
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 6/65 (9%)
Query: 35 SSYDVVFGKLALRCLFNDYFQSPKHFTTRIMFKPIDEPHVDLIATVSGPLDHKPEESITG 94
+SYD+ +L C DY +FT+ I+ + E H+DL+A++ LDH P E +T
Sbjct: 281 TSYDLTKTELRTIC---DYIHKYDYFTSYIIDCLMKEGHIDLLASI---LDHVPAELLTE 334
Query: 95 NALFR 99
R
Sbjct: 335 EICIR 339
>gb|AAO30050.1| unknown protein [Arabidopsis thaliana] gi|20260446|gb|AAM13121.1|
unknown protein [Arabidopsis thaliana]
gi|22331158|ref|NP_188479.2| nocturnin-related
[Arabidopsis thaliana]
Length = 262
Score = 34.7 bits (78), Expect = 5.7
Identities = 15/37 (40%), Positives = 25/37 (67%)
Query: 365 KSKTVVVSYNILGVDSASKHQDLYSNIRHSFQNNGFK 401
+ + VVSYNILG ++S H++LYSN+ + G++
Sbjct: 104 QERFTVVSYNILGDGNSSYHRELYSNVSVPYLKWGYR 140
>ref|ZP_00306306.1| COG1571: Predicted DNA-binding protein containing a Zn-ribbon
domain [Ferroplasma acidarmanus]
Length = 438
Score = 34.3 bits (77), Expect = 7.4
Identities = 39/112 (34%), Positives = 52/112 (45%), Gaps = 13/112 (11%)
Query: 296 KAKVGPRI--SSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSESLREASFQSIF 353
K K G I SS +LA W S+T+ + R G I YG I S+ E SF S F
Sbjct: 151 KIKTGHGIIGSSASLA---WPGKSYTYELLDYRYPNYGDITYGEKIDISSIPE-SFYSTF 206
Query: 354 T---ILNLSPPIADKSKTVVVSYNILGVDSASKHQDLYSNI--RHSFQNNGF 400
I N P I K KT V+ + + GVD + +Y I RH+ + G+
Sbjct: 207 NNIDIKNRYPAIFPKPKTPVI-FGVRGVDK-NDLITIYDRIKERHNIDDQGY 256
>ref|YP_077442.1| Polysaccharide deacetylase,Carbohydrate Esterase Family 4 [Bacillus
licheniformis ATCC 14580] gi|52001862|gb|AAU21804.1|
Polysaccharide deacetylase,Carbohydrate Esterase Family
4 [Bacillus licheniformis ATCC 14580]
gi|52784013|ref|YP_089842.1| YbaN [Bacillus
licheniformis ATCC 14580] gi|52346515|gb|AAU39149.1|
YbaN [Bacillus licheniformis DSM 13]
Length = 254
Score = 34.3 bits (77), Expect = 7.4
Identities = 19/65 (29%), Positives = 34/65 (52%), Gaps = 2/65 (3%)
Query: 251 IGITNYIDFGFELQTSVDDAIATNNISDSTFRIGASWQANKNFLLK--AKVGPRISSMAL 308
+ +T I +G + + D + N I D+TF + ASW +++ K G +I SM
Sbjct: 59 VALTFNISWGDQKAMPILDTLKANGIKDATFFLSASWAERHPDVVERIRKDGHQIGSMGY 118
Query: 309 AFKSW 313
A+K++
Sbjct: 119 AYKNY 123
>emb|CAB91586.1| putative protein [Arabidopsis thaliana]
gi|15231642|ref|NP_191475.1| F-box family protein
[Arabidopsis thaliana] gi|11357644|pir||T48984
hypothetical protein F25L23.20 - Arabidopsis thaliana
Length = 464
Score = 34.3 bits (77), Expect = 7.4
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYID 258
S S LS + + L V + +F S Y H +RVKNPL E G++G +D
Sbjct: 37 STSLLSRRWRYLLAFVPNLEFDDSAYLHR--DKRVKNPLHEKGLVGFVLTVD 86
>emb|CAB86943.1| putative protein [Arabidopsis thaliana] gi|11357520|pir||T47797
hypothetical protein F17J16.200 - Arabidopsis thaliana
(fragment)
Length = 827
Score = 34.3 bits (77), Expect = 7.4
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYID 258
S S LS + + L V + +F S Y H +RVKNPL E G++G +D
Sbjct: 519 STSLLSRRWRYLLAFVPNLEFDDSAYLHR--DKRVKNPLHEKGLVGFVLTVD 568
>dbj|BAB01117.1| unnamed protein product [Arabidopsis thaliana]
Length = 445
Score = 34.3 bits (77), Expect = 7.4
Identities = 15/32 (46%), Positives = 23/32 (71%)
Query: 370 VVSYNILGVDSASKHQDLYSNIRHSFQNNGFK 401
VVSYNILG ++S H++LYSN+ + G++
Sbjct: 111 VVSYNILGDGNSSYHRELYSNVSVPYLKWGYR 142
>emb|CAC48058.1| putative lyase [Rhizobium leguminosarum bv. viciae]
Length = 397
Score = 33.9 bits (76), Expect = 9.6
Identities = 23/74 (31%), Positives = 36/74 (48%), Gaps = 5/74 (6%)
Query: 240 RVKNPL--EENGVIGITNYID-FGFELQTSVDDAIATNNISDSTFRIGASWQANKNFLLK 296
RV++P E G +T Y F F+L +D A N++ D R+G SW + ++
Sbjct: 291 RVRHPAFQEHPGRATLTGYAGLFAFDLYPDIDVARFVNSLRD--IRLGVSWGGTETLVVP 348
Query: 297 AKVGPRISSMALAF 310
AKV +I +F
Sbjct: 349 AKVALQIPDRMTSF 362
>emb|CAG03971.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1200
Score = 33.9 bits (76), Expect = 9.6
Identities = 29/110 (26%), Positives = 50/110 (45%), Gaps = 14/110 (12%)
Query: 198 SCAMAYGVGSGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYI 257
SC + + SG+ + F++ +K +Q F +Q R KN E N +YI
Sbjct: 872 SCDLGNPMKSGTSVWAGLRFTVPWLKDTQETVQFE----LQIRSKN--ENNSHSEAVSYI 925
Query: 258 DFGFELQTSVDDAIATNNIS--DSTFRIGASWQANKNFLLKAKVGPRISS 305
L+ A+ +S D F + ASW++ KN+ ++ VGP + +
Sbjct: 926 -----LEVEAQAAVILQGVSLPDKVF-LPASWKSTKNYRVEQDVGPGVDT 969
>gb|AAF70821.1| beta-galactosidase [Lycopersicon esculentum]
Length = 892
Score = 33.9 bits (76), Expect = 9.6
Identities = 29/98 (29%), Positives = 44/98 (44%), Gaps = 15/98 (15%)
Query: 120 FGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKD-ELPKSAWLVSKMGRLTA 178
FG FP+ LR + G+ + + N F + + KK +L S L S G
Sbjct: 135 FGGFPIWLRD--------IPGIEFRTDNAPFKEEMERYVKKIVDLMISESLFSWQGGPII 186
Query: 179 GVQYEPQHGNAKLSN------LMNWSCAMAYGVGSGSP 210
+Q E ++GN + S M W+ MA G+G+G P
Sbjct: 187 LLQIENEYGNVESSFGPKGKLYMKWAAEMAVGLGAGVP 224
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 747,843,961
Number of Sequences: 2540612
Number of extensions: 31278833
Number of successful extensions: 72071
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 72037
Number of HSP's gapped (non-prelim): 27
length of query: 419
length of database: 863,360,394
effective HSP length: 131
effective length of query: 288
effective length of database: 530,540,222
effective search space: 152795583936
effective search space used: 152795583936
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0400a.3


