

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
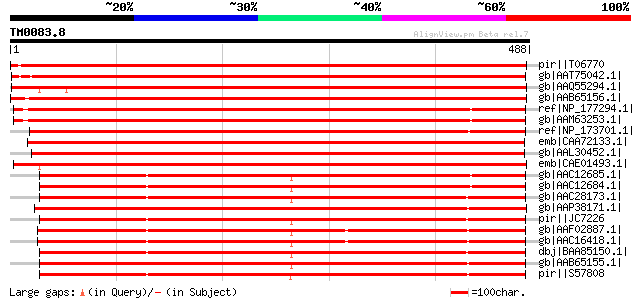
Query= TM0083.8
(488 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
pir||T06770 cellulase (EC 3.2.1.4) precursor - garden pea gi|728... 879 0.0
gb|AAT75042.1| Cel9B [Populus tremula x Populus tremuloides] 820 0.0
gb|AAQ55294.1| endo-1,4-beta-glucanase [Malus x domestica] 800 0.0
gb|AAB65156.1| basic cellulase [Citrus sinensis] gi|7488905|pir|... 793 0.0
ref|NP_177294.1| glycosyl hydrolase family 9 protein [Arabidopsi... 751 0.0
gb|AAM63253.1| putative beta-glucanase [Arabidopsis thaliana] 749 0.0
ref|NP_173701.1| glycosyl hydrolase family 9 protein [Arabidopsi... 746 0.0
emb|CAA72133.1| endo-1,4-beta-D-glucanase [Lycopersicon esculent... 743 0.0
gb|AAL30452.1| endo-beta-1,4-glucanase precursor [Nicotiana taba... 739 0.0
emb|CAE01493.1| P0041A24.5 [Oryza sativa (japonica cultivar-grou... 625 e-177
gb|AAC12685.1| endo-beta-1,4-glucanase [Pinus radiata] gi|113571... 619 e-176
gb|AAC12684.1| endo-beta-1,4-glucanase [Pinus radiata] gi|748447... 605 e-171
gb|AAC28173.1| T2H3.5 [Arabidopsis thaliana] gi|7268989|emb|CAB8... 597 e-169
gb|AAP38171.1| endo-1,4-beta-glucanase [Lilium longiflorum] 590 e-167
pir||JC7226 endo-1,3(4)-beta-glucanase (EC 3.2.1.6) - garden pea 587 e-166
gb|AAF02887.1| endo-1,4-beta glucanase [Arabidopsis thaliana] gi... 587 e-166
gb|AAC16418.1| endo-1,4-beta glucanase; ATCEL2 [Arabidopsis thal... 586 e-166
dbj|BAA85150.1| endo-1,4-beta-glucanase [Pisum sativum] 584 e-165
gb|AAB65155.1| acidic cellulase [Citrus sinensis] gi|7488904|pir... 579 e-164
pir||S57808 cellulase (EC 3.2.1.4) precursor - tomato gi|924622|... 576 e-163
>pir||T06770 cellulase (EC 3.2.1.4) precursor - garden pea
gi|728483|gb|AAA96135.1| endo-1,4-beta-glucanase
Length = 486
Score = 879 bits (2272), Expect = 0.0
Identities = 419/486 (86%), Positives = 453/486 (92%), Gaps = 2/486 (0%)
Query: 1 MMRTSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSD 60
MM+TS F LL +I LL V NV+C PNYREALAKSLLF QGQRSG+LPPDQQ+KWRS+
Sbjct: 1 MMKTSLLF--LLLIITCLLAVENVECKPNYREALAKSLLFFQGQRSGKLPPDQQIKWRSN 58
Query: 61 SGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIR 120
SGL DG NVDLSGGYYDAGDNVKFNFPMAFTTTMLSWS IEYGKRMGPQ+KEARAAIR
Sbjct: 59 SGLSDGLQDNVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSTIEYGKRMGPQMKEARAAIR 118
Query: 121 WGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETA 180
+ DY LKCATSTPGRLYVGVGDPNVDHKCWERPEDMDT RTVY+VS NPGSDVAAETA
Sbjct: 119 YAADYFLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTVRTVYYVSSKNPGSDVAAETA 178
Query: 181 AALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDE 240
AALAAAS+VF+KVDP+YSKLL+RT+QKVYQFALQYQGSYS+SLGSA CPFYCSYSGFKDE
Sbjct: 179 AALAAASIVFRKVDPSYSKLLLRTSQKVYQFALQYQGSYSNSLGSAACPFYCSYSGFKDE 238
Query: 241 LLWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQ 300
LLWGAAWLFRATNAV YY L+K+LGADDQPDIFSWD+KYAGAHVLLS+RALLNGDKNFDQ
Sbjct: 239 LLWGAAWLFRATNAVYYYKLVKSLGADDQPDIFSWDNKYAGAHVLLSKRALLNGDKNFDQ 298
Query: 301 YRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYT 360
YRQEA+NFMCKILPNSPSS+TQYTQGGLMFKLP+SNLQYVT+ITFLLTTY+KYMSA K+T
Sbjct: 299 YRQEADNFMCKILPNSPSSTTQYTQGGLMFKLPESNLQYVTAITFLLTTYSKYMSATKHT 358
Query: 361 FNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHP 420
F+CG+V VTPNTLRS+AKRQVDYILGENPL+MSYMVGYGP FPKRIHHRGSSLPSL+ HP
Sbjct: 359 FSCGSVFVTPNTLRSIAKRQVDYILGENPLRMSYMVGYGPYFPKRIHHRGSSLPSLSVHP 418
Query: 421 QTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGP 480
QTIGCDGGF PFFHS +PNPNILVGAIVGGPNQ+DGFPDDR DYSHSEP TYINGA+VGP
Sbjct: 419 QTIGCDGGFNPFFHSMSPNPNILVGAIVGGPNQNDGFPDDRGDYSHSEPATYINGAIVGP 478
Query: 481 LAYFAG 486
LAYF+G
Sbjct: 479 LAYFSG 484
>gb|AAT75042.1| Cel9B [Populus tremula x Populus tremuloides]
Length = 486
Score = 820 bits (2118), Expect = 0.0
Identities = 388/484 (80%), Positives = 436/484 (89%), Gaps = 2/484 (0%)
Query: 2 MRTSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDS 61
MR +SF LLF + ++L +G VQ PNY EALAKS+LF QGQRSGRLP QQL WRSDS
Sbjct: 1 MRRGASFC-LLFSLSLVL-LGFVQAKPNYNEALAKSILFFQGQRSGRLPGSQQLAWRSDS 58
Query: 62 GLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRW 121
GL DG A+VDL+GGYYDAGDNVKFNFPMAFTTTMLSWS +EYGKRMGP+L ARAAIRW
Sbjct: 59 GLSDGLFAHVDLTGGYYDAGDNVKFNFPMAFTTTMLSWSTLEYGKRMGPELPNARAAIRW 118
Query: 122 GTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAA 181
TDYLLKCAT+TPG+LYVGVGDPNVDHKCWERPEDMDT RTV+ VS +PGSDVA ETAA
Sbjct: 119 ATDYLLKCATATPGKLYVGVGDPNVDHKCWERPEDMDTVRTVFSVSARSPGSDVAGETAA 178
Query: 182 ALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDEL 241
ALAAAS+VF+KVD YS LL+RTA+KV+QFA+QYQG+YSDSLGSAVCPFYCSYSG+KDEL
Sbjct: 179 ALAAASMVFRKVDRKYSALLLRTARKVFQFAMQYQGAYSDSLGSAVCPFYCSYSGYKDEL 238
Query: 242 LWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQY 301
LWGAAWLFRATN + YYN+ K+LGADDQPD+FSWD+KYAG HVLLSRRALLN DKNF+Q+
Sbjct: 239 LWGAAWLFRATNEMSYYNIFKSLGADDQPDLFSWDNKYAGVHVLLSRRALLNNDKNFEQF 298
Query: 302 RQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTF 361
EAE+FMC+ILPNSP +TQYTQGGLM+KLP+SNLQYVTSITFLLTTYAKYM A ++TF
Sbjct: 299 EGEAESFMCRILPNSPYKTTQYTQGGLMYKLPESNLQYVTSITFLLTTYAKYMKATRHTF 358
Query: 362 NCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQ 421
NCGN+LVTPN+L VAKRQVDYILGENP++MSYMVG+GPNFPKRIHHRGSSLPSLA+HPQ
Sbjct: 359 NCGNLLVTPNSLLYVAKRQVDYILGENPIRMSYMVGFGPNFPKRIHHRGSSLPSLASHPQ 418
Query: 422 TIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPL 481
IGCD GF+PFFHS+NPNPNIL GAIVGGPNQ+DG+PD+RSDYSHSEP TYIN A+VGPL
Sbjct: 419 AIGCDSGFEPFFHSANPNPNILTGAIVGGPNQNDGYPDERSDYSHSEPATYINAAMVGPL 478
Query: 482 AYFA 485
AYFA
Sbjct: 479 AYFA 482
>gb|AAQ55294.1| endo-1,4-beta-glucanase [Malus x domestica]
Length = 497
Score = 800 bits (2065), Expect = 0.0
Identities = 385/491 (78%), Positives = 430/491 (87%), Gaps = 6/491 (1%)
Query: 2 MRTSSSFMSLLFLIPMLLFVGNVQC----NPNYREALAKSLLFLQGQRSGRLPPD--QQL 55
MR S S LF+I +VQ NPNYREALAKS+LF QGQRSGRLP QQ+
Sbjct: 3 MRLSLSIFISLFVILGSSISSSVQVLAAGNPNYREALAKSVLFFQGQRSGRLPAGAAQQI 62
Query: 56 KWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEA 115
WRS+SGL DGR A VDL+GGYYDAGDNVKFNFPMAFTTTMLSW A+EYGKRMGPQL E
Sbjct: 63 TWRSNSGLSDGRQAYVDLTGGYYDAGDNVKFNFPMAFTTTMLSWGALEYGKRMGPQLPET 122
Query: 116 RAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDV 175
RAAIRW TDYL+KCA TPGRLYVGVGDPNVDHKCWERPEDMDT RTVY VSP+NPGSDV
Sbjct: 123 RAAIRWATDYLIKCARQTPGRLYVGVGDPNVDHKCWERPEDMDTTRTVYSVSPSNPGSDV 182
Query: 176 AAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYS 235
A ETAAALAAASLVF++VDP YSKLL+ TA+ V QFA+QY+G+YSDSLGSAVCPFYCSYS
Sbjct: 183 AGETAAALAAASLVFRRVDPKYSKLLLNTARNVMQFAIQYRGAYSDSLGSAVCPFYCSYS 242
Query: 236 GFKDELLWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGD 295
G+ DELLWGAAWLFRATN + YYN IK+LGA D DIFSWD+K+AGA+VLLSRRALLN D
Sbjct: 243 GYNDELLWGAAWLFRATNELYYYNFIKSLGASDSTDIFSWDNKFAGAYVLLSRRALLNND 302
Query: 296 KNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMS 355
KNF+ Y+QEAE FMC+ILPNSPSSSTQYTQGGL++KLP SNLQYVTSITFLLTTY+KYM+
Sbjct: 303 KNFEPYKQEAEQFMCRILPNSPSSSTQYTQGGLIYKLPGSNLQYVTSITFLLTTYSKYMA 362
Query: 356 AKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPS 415
A+K TF+CGN++VTP LR++AK+QVDYILG NPL+MSYMVGYGP +PKRIHHRGSSLPS
Sbjct: 363 ARKLTFDCGNLVVTPMALRNLAKQQVDYILGVNPLKMSYMVGYGPYYPKRIHHRGSSLPS 422
Query: 416 LAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYING 475
L +H Q+IGCDGGFQPFF+S NPNPNILVGA+VGGPNQ+DGFPDDR DYSHSEP TYING
Sbjct: 423 LTSHRQSIGCDGGFQPFFYSLNPNPNILVGAVVGGPNQNDGFPDDRGDYSHSEPATYING 482
Query: 476 AVVGPLAYFAG 486
A+VGPLA+FAG
Sbjct: 483 AIVGPLAFFAG 493
>gb|AAB65156.1| basic cellulase [Citrus sinensis] gi|7488905|pir||T07885 cellulase
(EC 3.2.1.4) - sweet orange
Length = 488
Score = 793 bits (2049), Expect = 0.0
Identities = 377/486 (77%), Positives = 427/486 (87%), Gaps = 2/486 (0%)
Query: 1 MMRTSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSD 60
M R S LFL LL + N NPNYREALAKSLLF QGQRSGRLP DQQ+ WRS+
Sbjct: 1 MRRGVSLRCCCLFLF--LLLIENAHGNPNYREALAKSLLFFQGQRSGRLPKDQQITWRSN 58
Query: 61 SGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIR 120
SGL DG A+VDL+GGYYDAGDNVKFNFPMAFTTTMLSWS +EYGK+MGP+L+ ARAAIR
Sbjct: 59 SGLSDGLFAHVDLTGGYYDAGDNVKFNFPMAFTTTMLSWSTLEYGKKMGPELQNARAAIR 118
Query: 121 WGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETA 180
W TDYLLKCAT+TPG+LYVGVGDPN DHKCWERPEDMDT R+VY VS +NPGSDVA ETA
Sbjct: 119 WATDYLLKCATATPGKLYVGVGDPNADHKCWERPEDMDTVRSVYSVSASNPGSDVAGETA 178
Query: 181 AALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDE 240
AALAAASLVF+K DP Y+ LL+RTA+ V QFA+QY+G+YSDSLGSAVCPFYCSYSG+KDE
Sbjct: 179 AALAAASLVFRKGDPRYASLLLRTAKNVLQFAMQYRGAYSDSLGSAVCPFYCSYSGYKDE 238
Query: 241 LLWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQ 300
LLWGAAWLFRATN V YYN IK++G D D+FSWD+K+AGAHVLL+R ALLN DKNF+
Sbjct: 239 LLWGAAWLFRATNDVTYYNFIKSVGDDGGTDVFSWDNKFAGAHVLLARGALLNRDKNFEP 298
Query: 301 YRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYT 360
YRQEAE+F+C+ILPNSP ++TQYTQGGLM+K+P+SNLQYVTSI+FLLTTYAKYM A K+
Sbjct: 299 YRQEAEDFICRILPNSPFTTTQYTQGGLMYKMPESNLQYVTSISFLLTTYAKYMRATKHY 358
Query: 361 FNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHP 420
F CGN++V P L ++AKRQVDYILG NP++MSYMVG+GPNFP+RIHHRGSSLPSLA HP
Sbjct: 359 FTCGNMVVNPGLLTNLAKRQVDYILGVNPIKMSYMVGFGPNFPRRIHHRGSSLPSLANHP 418
Query: 421 QTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGP 480
Q+I CDGGF+PFFHSSNPNPNILVGAIVGGPNQ+DGFPDDRSDYSHSEP TYIN A+VGP
Sbjct: 419 QSIRCDGGFEPFFHSSNPNPNILVGAIVGGPNQNDGFPDDRSDYSHSEPATYINAAMVGP 478
Query: 481 LAYFAG 486
LAYFAG
Sbjct: 479 LAYFAG 484
>ref|NP_177294.1| glycosyl hydrolase family 9 protein [Arabidopsis thaliana]
gi|12323721|gb|AAG51817.1| putative beta-glucanase;
74324-76084 [Arabidopsis thaliana]
Length = 484
Score = 751 bits (1939), Expect = 0.0
Identities = 360/482 (74%), Positives = 410/482 (84%), Gaps = 4/482 (0%)
Query: 4 TSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGL 63
TS F LLF L + N NPNY+EAL+KSLLF QGQRSG LP QQ+ WR+ SGL
Sbjct: 2 TSLFFFVLLF---SSLLISNGDANPNYKEALSKSLLFFQGQRSGPLPRGQQISWRASSGL 58
Query: 64 FDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGT 123
DG A+VDL+GGYYDAGDNVKFN PMAFTTTMLSWSA+EYGKRMGP+L+ AR IRW T
Sbjct: 59 SDGSAAHVDLTGGYYDAGDNVKFNLPMAFTTTMLSWSALEYGKRMGPELENARVNIRWAT 118
Query: 124 DYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAAL 183
DYLLKCA +TPG+LYVGVGDPNVDHKCWERPEDMDT RTVY VS +NPGSDVAAETAAAL
Sbjct: 119 DYLLKCARATPGKLYVGVGDPNVDHKCWERPEDMDTPRTVYSVSASNPGSDVAAETAAAL 178
Query: 184 AAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLW 243
AAAS+VF+KVD YS+LL+ TA+ V QFA+QYQG+YSDSL S+VCPFYCSYSG+KDEL+W
Sbjct: 179 AAASMVFRKVDSKYSRLLLATAKDVMQFAIQYQGAYSDSLSSSVCPFYCSYSGYKDELMW 238
Query: 244 GAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQ 303
GA+WL RATN Y N IK+LG DQPDIFSWD+KYAGA+VLLSRRALLN D NF+QY+Q
Sbjct: 239 GASWLLRATNNPYYANFIKSLGGGDQPDIFSWDNKYAGAYVLLSRRALLNKDSNFEQYKQ 298
Query: 304 EAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNC 363
AENF+CKILP+SPSSSTQYTQGGLM+KLP SNLQYVTSITFLLTTYAKYM A K+TFNC
Sbjct: 299 AAENFICKILPDSPSSSTQYTQGGLMYKLPQSNLQYVTSITFLLTTYAKYMKATKHTFNC 358
Query: 364 GNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTI 423
G+ ++ PN L S++KRQVDYILG+NP++MSYMVG+ NFPKRIHHR SSLPS A Q++
Sbjct: 359 GSSVIVPNALISLSKRQVDYILGDNPIKMSYMVGFSSNFPKRIHHRASSLPSHALRSQSL 418
Query: 424 GCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAY 483
GC+GGFQ F+ + NPNPNIL GAIVGGPNQ+DG+PD R DYSH+EP TYIN A VGPLAY
Sbjct: 419 GCNGGFQSFY-TQNPNPNILTGAIVGGPNQNDGYPDQRDDYSHAEPATYINAAFVGPLAY 477
Query: 484 FA 485
FA
Sbjct: 478 FA 479
>gb|AAM63253.1| putative beta-glucanase [Arabidopsis thaliana]
Length = 484
Score = 749 bits (1934), Expect = 0.0
Identities = 359/482 (74%), Positives = 410/482 (84%), Gaps = 4/482 (0%)
Query: 4 TSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGL 63
TS F LLF L + N NPNY+EAL+KSLLF QGQRSG LP QQ+ WR+ SGL
Sbjct: 2 TSLFFFVLLF---SSLLISNGDANPNYKEALSKSLLFFQGQRSGPLPRGQQISWRASSGL 58
Query: 64 FDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGT 123
DG A+VDL+GGYYDAGDNVKFN PMAFTTTMLSWSA+EYGKRMGP+L+ AR IRW T
Sbjct: 59 SDGSAAHVDLTGGYYDAGDNVKFNLPMAFTTTMLSWSALEYGKRMGPELENARVNIRWAT 118
Query: 124 DYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAAL 183
DYLLKCA +TPG+LYVGVGDPNVDHKCWERPEDMDT RTVY VS +NPGSDVAAETAAAL
Sbjct: 119 DYLLKCARATPGKLYVGVGDPNVDHKCWERPEDMDTPRTVYSVSASNPGSDVAAETAAAL 178
Query: 184 AAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLW 243
AAAS+VF+KV+ YS+LL+ TA+ V QFA+QYQG+YSDSL S+VCPFYCSYSG+KDEL+W
Sbjct: 179 AAASMVFRKVNSKYSRLLLATAKDVMQFAIQYQGAYSDSLSSSVCPFYCSYSGYKDELMW 238
Query: 244 GAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQ 303
GA+WL RATN Y N IK+LG DQPDIFSWD+KYAGA+VLLSRRALLN D NF+QY+Q
Sbjct: 239 GASWLLRATNNPYYANFIKSLGGGDQPDIFSWDNKYAGAYVLLSRRALLNKDSNFEQYKQ 298
Query: 304 EAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNC 363
AENF+CKILP+SPSSSTQYTQGGLM+KLP SNLQYVTSITFLLTTYAKYM A K+TFNC
Sbjct: 299 AAENFICKILPDSPSSSTQYTQGGLMYKLPQSNLQYVTSITFLLTTYAKYMKATKHTFNC 358
Query: 364 GNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTI 423
G+ ++ PN L S++KRQVDYILG+NP++MSYMVG+ NFPKRIHHR SSLPS A Q++
Sbjct: 359 GSSVIVPNALISLSKRQVDYILGDNPIKMSYMVGFSSNFPKRIHHRASSLPSHALRSQSL 418
Query: 424 GCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAY 483
GC+GGFQ F+ + NPNPNIL GAIVGGPNQ+DG+PD R DYSH+EP TYIN A VGPLAY
Sbjct: 419 GCNGGFQSFY-TQNPNPNILTGAIVGGPNQNDGYPDQRDDYSHAEPATYINAAFVGPLAY 477
Query: 484 FA 485
FA
Sbjct: 478 FA 479
>ref|NP_173701.1| glycosyl hydrolase family 9 protein [Arabidopsis thaliana]
gi|2462836|gb|AAB72171.1| beta-glucanase [Arabidopsis
thaliana] gi|25350162|pir||G86362 beta-glucanase
[imported] - Arabidopsis thaliana
Length = 484
Score = 746 bits (1925), Expect = 0.0
Identities = 353/467 (75%), Positives = 404/467 (85%), Gaps = 1/467 (0%)
Query: 19 LFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYY 78
L + N +PNYREAL+KSLLF QGQRSGRLP DQQL WRS SGL DG A+VDL+GGYY
Sbjct: 14 LSLENTYASPNYREALSKSLLFFQGQRSGRLPSDQQLSWRSSSGLSDGSSAHVDLTGGYY 73
Query: 79 DAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLY 138
DAGDNVKFNFPMAFTTTMLSWS++EYGK+MGP+L+ +R AIRW TDYLLKCA +TPG+LY
Sbjct: 74 DAGDNVKFNFPMAFTTTMLSWSSLEYGKKMGPELQNSRVAIRWATDYLLKCARATPGKLY 133
Query: 139 VGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYS 198
VGVGDPN DHKCWERPEDMDT RTVY VSP+NPGSDVAAETAAALAA+S+VF+KVDP YS
Sbjct: 134 VGVGDPNGDHKCWERPEDMDTPRTVYSVSPSNPGSDVAAETAAALAASSMVFRKVDPKYS 193
Query: 199 KLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYY 258
+LL+ TA+KV QFA+QY+G+YS+SL S+VCPFYCSYSG+KDELLWGAAWL RATN Y
Sbjct: 194 RLLLATAKKVMQFAIQYRGAYSNSLSSSVCPFYCSYSGYKDELLWGAAWLHRATNDPYYT 253
Query: 259 NLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPS 318
N IK+LG DQPDIFSWD+KYAGA+VLLSRRA+LN D NF+ Y+Q AENFMCKILPNSPS
Sbjct: 254 NFIKSLGGGDQPDIFSWDNKYAGAYVLLSRRAVLNKDNNFELYKQAAENFMCKILPNSPS 313
Query: 319 SSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAK 378
SST+YT+GGLM+KLP SNLQYVTSITFLLTTYAKYM + K TFNCGN L+ PN L +++K
Sbjct: 314 SSTKYTKGGLMYKLPQSNLQYVTSITFLLTTYAKYMKSTKQTFNCGNSLIVPNALINLSK 373
Query: 379 RQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNP 438
RQVDY+LG NP++MSYMVG+ NFPKRIHHRGSSLPS A ++GC+GGFQ F + NP
Sbjct: 374 RQVDYVLGVNPMKMSYMVGFSSNFPKRIHHRGSSLPSRAVRSNSLGCNGGFQS-FRTQNP 432
Query: 439 NPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
NPNIL GAIVGGPNQ+D +PD R DY+ SEP TYIN A VGPLAYFA
Sbjct: 433 NPNILTGAIVGGPNQNDEYPDQRDDYTRSEPATYINAAFVGPLAYFA 479
>emb|CAA72133.1| endo-1,4-beta-D-glucanase [Lycopersicon esculentum]
gi|7488976|pir||T07025 cellulase (EC 3.2.1.4) - tomato
Length = 479
Score = 743 bits (1919), Expect = 0.0
Identities = 339/468 (72%), Positives = 406/468 (86%)
Query: 17 MLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGG 76
++LF +YR+AL KS+LF +GQRSG+LPP+Q+L WR +SGL DG LA VDLSGG
Sbjct: 7 LILFFSLGTAQFDYRDALEKSILFFEGQRSGKLPPNQRLSWRGNSGLQDGSLAKVDLSGG 66
Query: 77 YYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGR 136
YYDAGDNVKFNFPMA+TTTMLSW+ +EYGKRMG QL+ ARAAIRW TDY LKCA + P +
Sbjct: 67 YYDAGDNVKFNFPMAYTTTMLSWNTLEYGKRMGAQLQHARAAIRWATDYFLKCANAAPNK 126
Query: 137 LYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPT 196
L+VGVGDPN DHKCWERPEDMDT R+VY+VSP+NPGSDVA E AAALAAASLVF+ VDP
Sbjct: 127 LFVGVGDPNADHKCWERPEDMDTIRSVYYVSPSNPGSDVAGEMAAALAAASLVFRSVDPV 186
Query: 197 YSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVK 256
YSK L+ A KV++FA+QY+GSYSDSLGSA CPFYCSYSG+KDEL WGAAWL RATN +
Sbjct: 187 YSKKLLGNAVKVFRFAVQYRGSYSDSLGSAACPFYCSYSGYKDELYWGAAWLLRATNDIS 246
Query: 257 YYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNS 316
Y NLI TLGA+D PD+FSWD+KYAGAHVL+SRR+++ D FD ++Q AE+F+CK+LPNS
Sbjct: 247 YLNLINTLGANDVPDLFSWDNKYAGAHVLMSRRSVVGNDNRFDSFKQRAEDFVCKVLPNS 306
Query: 317 PSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSV 376
P +STQYT+GGL++KLP+ NLQYVTSIT LLTTYAKYM+ KK+TFNCG+++VT T+R+
Sbjct: 307 PYTSTQYTKGGLLYKLPEENLQYVTSITSLLTTYAKYMATKKHTFNCGSLVVTEKTIRNF 366
Query: 377 AKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSS 436
AKRQVDYILG NP++MSYMVGYG N+P+RIHHRGSSLPSLA HPQ+ GC+ GFQPF++++
Sbjct: 367 AKRQVDYILGNNPMKMSYMVGYGSNYPRRIHHRGSSLPSLAMHPQSFGCEAGFQPFYYTA 426
Query: 437 NPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
NPNPNILVGAI+GGPNQ+D FPD+R+DYSHSEP TYIN A+VGPLAYF
Sbjct: 427 NPNPNILVGAIIGGPNQNDFFPDERTDYSHSEPATYINAAIVGPLAYF 474
>gb|AAL30452.1| endo-beta-1,4-glucanase precursor [Nicotiana tabacum]
Length = 489
Score = 739 bits (1908), Expect = 0.0
Identities = 338/464 (72%), Positives = 406/464 (86%)
Query: 21 VGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDA 80
+G Q NYREAL KS+LF +GQRSG+LP +Q++ WR SGL DG LA VDL+GGYYDA
Sbjct: 21 LGTAQGVFNYREALEKSILFFEGQRSGKLPHNQRVSWRGSSGLSDGSLAKVDLTGGYYDA 80
Query: 81 GDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVG 140
GDNVKFNFPMA+TTT+LSW+ +EYGKRMGPQL+ ARAAIRW TDYLLKCA + P +L+VG
Sbjct: 81 GDNVKFNFPMAYTTTLLSWNTLEYGKRMGPQLQNARAAIRWATDYLLKCANAAPNKLFVG 140
Query: 141 VGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKL 200
VGDPN DHKCWERPEDMDT R+VY+VSP++PGSDVA E AAALAAASLVF+ VDP YSK
Sbjct: 141 VGDPNSDHKCWERPEDMDTVRSVYYVSPSSPGSDVAGEMAAALAAASLVFRTVDPVYSKK 200
Query: 201 LVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNL 260
L+ A KV++FA+QY+GSYSDSLGSA CPFYCSYSG+KDEL WGAAWL RATN + Y N
Sbjct: 201 LLGNAVKVFRFAVQYRGSYSDSLGSAACPFYCSYSGYKDELYWGAAWLLRATNDISYLNF 260
Query: 261 IKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSS 320
I TLGA+D PD+FSWD+KYAGAHVL++RR+++ DK FD +RQ+AE+F+CKILPNSP +S
Sbjct: 261 INTLGANDVPDLFSWDNKYAGAHVLMARRSVVGNDKRFDPFRQQAEDFVCKILPNSPYTS 320
Query: 321 TQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQ 380
TQYT+GGL++KL + NLQYVTSIT LLTTYAKYM++KK+TFNCG++LVT T+R +AKRQ
Sbjct: 321 TQYTKGGLIYKLTEENLQYVTSITSLLTTYAKYMASKKHTFNCGSLLVTEKTIRILAKRQ 380
Query: 381 VDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNP 440
VDYILG NP++MSYMVGYG N+P+R+HHRGSSLPS+A HPQ+ GCDGGFQP+++++N NP
Sbjct: 381 VDYILGNNPMKMSYMVGYGTNYPRRVHHRGSSLPSMAMHPQSFGCDGGFQPYYYTANANP 440
Query: 441 NILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
NILVGAIVGGPNQ+D FPD+R+DYSHSEP TYIN A+VGPLAYF
Sbjct: 441 NILVGAIVGGPNQNDFFPDERTDYSHSEPATYINAAIVGPLAYF 484
>emb|CAE01493.1| P0041A24.5 [Oryza sativa (japonica cultivar-group)]
gi|50924542|ref|XP_472631.1| P0041A24.5 [Oryza sativa
(japonica cultivar-group)]
Length = 500
Score = 625 bits (1611), Expect = e-177
Identities = 303/491 (61%), Positives = 380/491 (76%), Gaps = 8/491 (1%)
Query: 4 TSSSFMSLLFLIPMLL-FVGNVQC------NPNYREALAKSLLFLQGQRSGRLPPDQQLK 56
+SSS+ +L+ + +L F G+V +P+Y +ALAKS+LF QGQRSGRLPPDQ +K
Sbjct: 6 SSSSWRALVLVAAAVLSFSGHVVVAAAAAGHPDYADALAKSILFFQGQRSGRLPPDQAVK 65
Query: 57 WRSDSGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRM-GPQLKEA 115
WRS+SGL DG ANVDL+GGYYD GDNVKF FPMAFTTTMLSW +EYG RM G L++A
Sbjct: 66 WRSNSGLSDGSAANVDLTGGYYDGGDNVKFGFPMAFTTTMLSWGVVEYGGRMRGRVLRDA 125
Query: 116 RAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDV 175
R A+RW DYLL+ AT+TPG LYVGVGDP+ DH+CWERPEDMDT R VY VS ++PGSDV
Sbjct: 126 RDAVRWAADYLLRAATATPGVLYVGVGDPDADHRCWERPEDMDTPRAVYSVSASSPGSDV 185
Query: 176 AAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYS 235
AAETAAALAAASL + DP YS+ L+ A+ V FA+++QG YSD +G V +Y SYS
Sbjct: 186 AAETAAALAAASLALRAADPGYSRRLLAAARDVMAFAVRHQGKYSDHVGGDVGAYYASYS 245
Query: 236 GFKDELLWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGD 295
G++DELLWG+AWL AT Y + + +LGA+D D+FSWD+K AGA VLLSRRAL+NGD
Sbjct: 246 GYQDELLWGSAWLLWATRNASYLDYLASLGANDGVDMFSWDNKLAGARVLLSRRALVNGD 305
Query: 296 KNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMS 355
+ D +R+ AE+F+C+ILP SPSS+TQYT GG+M+K +NLQYVTS +FLLTT+AKYM+
Sbjct: 306 RRLDAFRRLAEDFICRILPGSPSSTTQYTPGGMMYKSGHANLQYVTSASFLLTTFAKYMA 365
Query: 356 AKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPS 415
+TF+C ++ VT TLR++A++QVDYILG NP MSYMVGYG FP+RIHHRG+S+PS
Sbjct: 366 VSNHTFSCQSLPVTAKTLRALARKQVDYILGANPQGMSYMVGYGARFPQRIHHRGASMPS 425
Query: 416 LAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYING 475
+AA+P IGC GF +F++ NPN+ GA+VGGP+Q D FPD+R DY SEPTTY N
Sbjct: 426 VAAYPAHIGCQEGFSGYFNAGGANPNVHTGAVVGGPDQHDAFPDERGDYDRSEPTTYTNA 485
Query: 476 AVVGPLAYFAG 486
A+VG LAYFAG
Sbjct: 486 ALVGCLAYFAG 496
>gb|AAC12685.1| endo-beta-1,4-glucanase [Pinus radiata] gi|11357128|pir||T46610
cellulase (EC 3.2.1.4) 2 precursor - Monterey pine
Length = 515
Score = 619 bits (1596), Expect = e-176
Identities = 293/461 (63%), Positives = 366/461 (78%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
+YR+AL KS+LF +GQRSG+LP +Q+++WR DS L DG +VDL+GGYYDAGDNVKF F
Sbjct: 53 DYRDALTKSILFYEGQRSGKLPLNQRMRWRGDSALSDGAAGHVDLTGGYYDAGDNVKFGF 112
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS +E+G RMG L A+AAIRW TDYLLK AT+ PG +YV VGDPN+DH
Sbjct: 113 PMAFTTTMLSWSVLEFGGRMGDNLDHAKAAIRWATDYLLK-ATARPGVIYVQVGDPNLDH 171
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
KCWERPEDMDT+R VY + ++PGSDVAAETAAALAAAS+VF+ DP YS+ L++TA +V
Sbjct: 172 KCWERPEDMDTSRNVYKIDADHPGSDVAAETAAALAAASMVFRTTDPIYSRHLLQTAMQV 231
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+ FA +++G+YSDSL S VCPFYCSYSG+ DELLWGAAWL RA+ V Y + I++ L
Sbjct: 232 FDFADKHRGAYSDSLQSEVCPFYCSYSGYNDELLWGAAWLQRASRNVSYISYIQSHGQNL 291
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
G +D +IFSWDDK+AGA VLL++ LL K+ ++YR A+NF+C +LP +P+S QYT
Sbjct: 292 GGEDNVNIFSWDDKHAGARVLLAKEVLLRNSKSLEEYRGHADNFVCSLLPGTPNSQAQYT 351
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL++K+ D NLQYVTS TFLL TY KY+ K CGN++VTP +R++AKRQVDYI
Sbjct: 352 PGGLLYKMSDCNLQYVTSTTFLLFTYPKYLRVSKQVVRCGNMVVTPTRIRTLAKRQVDYI 411
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYG FP+RIHHRGSSLPS+ HPQ I C+ GFQ + S+ PNPN LV
Sbjct: 412 LGDNPLRMSYMVGYGAKFPERIHHRGSSLPSIYQHPQIIPCNDGFQSLY-SNAPNPNRLV 470
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP+ +D F D+R+DY+HSEPTTYIN +VG LAY A
Sbjct: 471 GAVVGGPDNNDHFSDERNDYAHSEPTTYINAPLVGSLAYLA 511
>gb|AAC12684.1| endo-beta-1,4-glucanase [Pinus radiata] gi|7484476|pir||T10734
cellulase (EC 3.2.1.4) 1 precursor - Monterey pine
Length = 510
Score = 605 bits (1560), Expect = e-171
Identities = 292/461 (63%), Positives = 357/461 (77%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NYR+AL+K++LF +GQRSG+LP Q++ WR DS L DG +VDL GGYYDAGDNVKF F
Sbjct: 47 NYRDALSKAILFYEGQRSGKLPGSQRMTWRRDSALSDGFSQHVDLVGGYYDAGDNVKFGF 106
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS +E+G MG +L ++ AIRW TDYLLK AT+ PG +YV VGDPN DH
Sbjct: 107 PMAFTTTMLSWSVLEFGGFMGGELANSKDAIRWATDYLLK-ATAHPGTIYVQVGDPNTDH 165
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
KCWERPEDMDTARTVY + +PGSDVAAETAAALAAASLVF+K DP YSK L+ TA +V
Sbjct: 166 KCWERPEDMDTARTVYKIDSQHPGSDVAAETAAALAAASLVFRKSDPQYSKRLIHTAMRV 225
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+ FA +++G+YSDSL S VCPFYCSYSG++DELLWGAAWL +AT Y N I++ L
Sbjct: 226 FDFADKHRGAYSDSLRSCVCPFYCSYSGYEDELLWGAAWLHKATRNPTYLNYIQSKGQSL 285
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GAD+ +IF WD+K+AGA VLLS+ LL K+ Y+ A+N++C +LP S +YT
Sbjct: 286 GADESDNIFGWDNKHAGARVLLSKEFLLRNVKSLHDYKGHADNYICSLLPGVSYSQAKYT 345
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL++ L DSNLQYVTS +FLL TYAKY+S+ K+ CG++ +P LR++AKRQVDYI
Sbjct: 346 PGGLLYTLSDSNLQYVTSSSFLLFTYAKYLSSSKHVVTCGSMTFSPKRLRTIAKRQVDYI 405
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYG ++P+RIHHRGSSLPS HPQ I C+ GFQ + S+ PNPNILV
Sbjct: 406 LGDNPLKMSYMVGYGSHYPERIHHRGSSLPSKGNHPQAIPCNAGFQSLY-SNAPNPNILV 464
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP+ D FPDDR+DY SEPTTYIN VG LAY A
Sbjct: 465 GAVVGGPDSMDRFPDDRNDYEQSEPTTYINAPFVGSLAYLA 505
>gb|AAC28173.1| T2H3.5 [Arabidopsis thaliana] gi|7268989|emb|CAB80722.1| putative
endo-1, 4-beta glucanase [Arabidopsis thaliana]
gi|20855941|gb|AAM26639.1| AT4g02290/T2H3_5 [Arabidopsis
thaliana] gi|19310536|gb|AAL85001.1| AT4g02290/T2H3_5
[Arabidopsis thaliana] gi|15235301|ref|NP_192138.1|
glycosyl hydrolase family 9 protein [Arabidopsis
thaliana] gi|7484857|pir||T01419 cellulase (EC 3.2.1.4)
T2H3.5 precursor - Arabidopsis thaliana
Length = 516
Score = 597 bits (1539), Expect = e-169
Identities = 285/461 (61%), Positives = 355/461 (76%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY++AL KS+LF +GQRSG+LP +Q++ WR DSGL DG +VDL GGYYDAGDN+KF F
Sbjct: 52 NYKDALTKSILFFEGQRSGKLPSNQRMSWRRDSGLSDGSALHVDLVGGYYDAGDNIKFGF 111
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS IE+G M +L+ A+ AIRW TDYLLK ATS P +YV VGD N DH
Sbjct: 112 PMAFTTTMLSWSVIEFGGLMKSELQNAKIAIRWATDYLLK-ATSQPDTIYVQVGDANKDH 170
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
CWERPEDMDT R+V+ V N PGSDVAAETAAALAAA++VF+K DP+YSK+L++ A V
Sbjct: 171 SCWERPEDMDTVRSVFKVDKNIPGSDVAAETAAALAAAAIVFRKSDPSYSKVLLKRAISV 230
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+ FA +Y+G+YS L VCPFYCSYSG++DELLWGAAWL +AT +KY N IK L
Sbjct: 231 FAFADKYRGTYSAGLKPDVCPFYCSYSGYQDELLWGAAWLQKATKNIKYLNYIKINGQIL 290
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GA + + F WD+K+AGA +LL++ L+ K +Y+ A+NF+C ++P +P SSTQYT
Sbjct: 291 GAAEYDNTFGWDNKHAGARILLTKAFLVQNVKTLHEYKGHADNFICSVIPGAPFSSTQYT 350
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL+FK+ D+N+QYVTS +FLL TYAKY+++ K +CG + TP LRS+AKRQVDY+
Sbjct: 351 PGGLLFKMADANMQYVTSTSFLLLTYAKYLTSAKTVVHCGGSVYTPGRLRSIAKRQVDYL 410
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYGP FP+RIHHRGSSLP +A+HP I C GF +S +PNPN LV
Sbjct: 411 LGDNPLRMSYMVGYGPKFPRRIHHRGSSLPCVASHPAKIQCHQGF-AIMNSQSPNPNFLV 469
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP+Q D FPD+RSDY SEP TYIN +VG LAYFA
Sbjct: 470 GAVVGGPDQHDRFPDERSDYEQSEPATYINSPLVGALAYFA 510
>gb|AAP38171.1| endo-1,4-beta-glucanase [Lilium longiflorum]
Length = 490
Score = 590 bits (1520), Expect = e-167
Identities = 271/463 (58%), Positives = 355/463 (76%), Gaps = 1/463 (0%)
Query: 24 VQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDN 83
+ N NY++AL+KS++F +GQRSG+LPP+Q L WR++S L DG NVDL+GGYYDAGDN
Sbjct: 21 IPANINYKDALSKSIIFFEGQRSGKLPPNQTLTWRANSALSDGAADNVDLTGGYYDAGDN 80
Query: 84 VKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGD 143
VKF FPMAF+ TMLSWS IE+GK + + +AR AIRWGTDYLLK ++ P +YV VGD
Sbjct: 81 VKFGFPMAFSVTMLSWSVIEFGKLIPADIDKAREAIRWGTDYLLKACSALPDAIYVQVGD 140
Query: 144 PNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVR 203
P +H+CWERPE+M T R+VY ++P+NPGSDVA ETAAALAAASLVF K+D Y+K L
Sbjct: 141 PTSEHQCWERPENMSTQRSVYKITPSNPGSDVAGETAAALAAASLVFHKIDHKYAKTLRE 200
Query: 204 TAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT 263
TA+K + FA ++GSYSDSL SAVCP+YCSYSGF+DELLW A WL++AT+ Y N ++
Sbjct: 201 TAKKAFTFAESHRGSYSDSLSSAVCPYYCSYSGFQDELLWAAGWLYKATHNASYLNFAQS 260
Query: 264 LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQY 323
L D D FSWD+K+ GA +LL++ ++N ++ Y+Q+AE F+C +LPNSP+ S +Y
Sbjct: 261 LNVDSNADTFSWDNKFPGADILLAQDFIINKNQAASSYKQQAEKFLCTVLPNSPTLSAKY 320
Query: 324 TQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDY 383
T GGL++K+ SNLQYVTS +FLL YAKY+ + TF+CGN++VTP+ L + ++Q+ Y
Sbjct: 321 TAGGLLYKMTPSNLQYVTSTSFLLGVYAKYLRISRRTFSCGNMVVTPSVLIRLVEKQIGY 380
Query: 384 ILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNIL 443
ILG NP +SYMV +G NFP IHHRGSS+PS+ A Q IGCD GFQ ++++SNPNPNIL
Sbjct: 381 ILGHNPQALSYMVDFGNNFPLHIHHRGSSIPSVHADAQAIGCDEGFQ-YYNTSNPNPNIL 439
Query: 444 VGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAG 486
GA+VGGP+++D F DDR +YS SEP TYIN +VG LA+ AG
Sbjct: 440 TGAVVGGPDENDSFADDRDNYSQSEPATYINAPLVGTLAFLAG 482
>pir||JC7226 endo-1,3(4)-beta-glucanase (EC 3.2.1.6) - garden pea
Length = 506
Score = 587 bits (1513), Expect = e-166
Identities = 284/461 (61%), Positives = 349/461 (75%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NYR+AL KS+LF QGQRSG+LP +Q++ WR DSGL DG +VDL GGYYDAGDNVKF F
Sbjct: 42 NYRDALTKSILFFQGQRSGKLPSNQRISWRRDSGLSDGSALHVDLVGGYYDAGDNVKFGF 101
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS IE+G M +L A+ A+RW TDYLLK AT+ P +YV VGD DH
Sbjct: 102 PMAFTTTMLSWSVIEFGGLMKSELPNAKKAVRWATDYLLK-ATAHPNIIYVQVGDAKKDH 160
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
CWERPEDMDT R+V+ V N PGS+VAAETAAALAAASLVF+K DPTY+K+LVR A +V
Sbjct: 161 ACWERPEDMDTPRSVFKVDANAPGSEVAAETAAALAAASLVFRKSDPTYAKILVRRAIRV 220
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+QFA +++ SYS++L VCPFYCSYSG++DELLWGAAWL +AT Y I+T L
Sbjct: 221 FQFADKHRRSYSNALKPFVCPFYCSYSGYQDELLWGAAWLHKATKNPMYLKYIQTNGQIL 280
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GA + + F WD+K+ GA +LLS+ L+ K+ Y+ ++NF+C ++P + SSS QYT
Sbjct: 281 GAAEFDNTFGWDNKHVGARILLSKEFLVQNVKSLHDYKGHSDNFVCSLIPGAGSSSAQYT 340
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL+FK+ DSN+QYVTS TFLL TYAKY++ NCG VTP LR++AKRQVDY+
Sbjct: 341 PGGLLFKMSDSNMQYVTSTTFLLVTYAKYLTKSHSVVNCGGTTVTPKRLRTLAKRQVDYL 400
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYGP +P+RIHHRGSSLPS+A HP I C GF +S +PNPNIL+
Sbjct: 401 LGDNPLKMSYMVGYGPRYPQRIHHRGSSLPSMAVHPGKIQCSAGF-GVMNSKSPNPNILM 459
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP+Q D FPD RSDY SEP TY+N +VG LAY A
Sbjct: 460 GAVVGGPDQHDRFPDQRSDYEQSEPATYVNAPLVGTLAYLA 500
>gb|AAF02887.1| endo-1,4-beta glucanase [Arabidopsis thaliana]
gi|15218612|ref|NP_171779.1| endo-1,4-beta-glucanase /
cellulase (CEL2) [Arabidopsis thaliana]
gi|25350166|pir||A86158 endo-1,4-beta glucanase
[imported] - Arabidopsis thaliana
Length = 501
Score = 587 bits (1512), Expect = e-166
Identities = 284/463 (61%), Positives = 357/463 (76%), Gaps = 8/463 (1%)
Query: 27 NPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKF 86
N NY++AL+KS+LF +GQRSG+LPP+Q++ WRS+SGL DG NVDL GGYYDAGDN+KF
Sbjct: 41 NHNYKDALSKSILFFEGQRSGKLPPNQRMTWRSNSGLSDGSALNVDLVGGYYDAGDNMKF 100
Query: 87 NFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
FPMAFTTTMLSWS IE+G M +L A+ AIRW TD+LLK ATS P +YV VGDPN+
Sbjct: 101 GFPMAFTTTMLSWSLIEFGGLMKSELPNAKDAIRWATDFLLK-ATSHPDTIYVQVGDPNM 159
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DH CWERPEDMDT R+V+ V NNPGSD+A E AAALAAAS+VF+K DP+YS L++ A
Sbjct: 160 DHACWERPEDMDTPRSVFKVDKNNPGSDIAGEIAAALAAASIVFRKCDPSYSNHLLQRAI 219
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT--- 263
V+ FA +Y+G YS L VCPFYCSYSG++DELLWGAAWL +ATN Y N IK
Sbjct: 220 TVFTFADKYRGPYSAGLAPEVCPFYCSYSGYQDELLWGAAWLQKATNNPTYLNYIKANGQ 279
Query: 264 -LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQ 322
LGAD+ ++FSWD+K+ GA +LLS+ L+ K+ ++Y++ A++F+C +LP +SS+Q
Sbjct: 280 ILGADEFDNMFSWDNKHVGARILLSKEFLIQKVKSLEEYKEHADSFICSVLPG--ASSSQ 337
Query: 323 YTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVD 382
YT GGL+FK+ +SN+QYVTS +FLL TYAKY+++ + CG +VTP LRS+AK+QVD
Sbjct: 338 YTPGGLLFKMGESNMQYVTSTSFLLLTYAKYLTSARTVAYCGGSVVTPARLRSIAKKQVD 397
Query: 383 YILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNI 442
Y+LG NPL+MSYMVGYG +P+RIHHRGSSLPS+A HP I C GF F S +PNPN
Sbjct: 398 YLLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRIQCHDGFS-LFTSQSPNPND 456
Query: 443 LVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
LVGA+VGGP+Q+D FPD+RSDY SEP TYIN +VG LAY A
Sbjct: 457 LVGAVVGGPDQNDQFPDERSDYGRSEPATYINAPLVGALAYLA 499
>gb|AAC16418.1| endo-1,4-beta glucanase; ATCEL2 [Arabidopsis thaliana]
gi|25350173|pir||T52135 cellulase (EC 3.2.1.4)
[imported] - Arabidopsis thaliana
Length = 501
Score = 586 bits (1510), Expect = e-166
Identities = 283/463 (61%), Positives = 357/463 (76%), Gaps = 8/463 (1%)
Query: 27 NPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKF 86
N NY++AL+KS+LF +GQRSG+LPP+Q++ WRS+SGL DG NVDL GGYYDAGDN+KF
Sbjct: 41 NHNYKDALSKSILFFEGQRSGKLPPNQRMTWRSNSGLSDGSALNVDLVGGYYDAGDNMKF 100
Query: 87 NFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
FPMAFTTTMLSWS IE+G M +L A+ AIRW TD+LLK ATS P +YV VGDPN+
Sbjct: 101 GFPMAFTTTMLSWSLIEFGGLMKSELPNAKDAIRWATDFLLK-ATSHPDTIYVQVGDPNM 159
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DH CWERPEDMDT R+V+ V NNPGSD+A E AAALAAAS+VF+K DP+YS L++ A
Sbjct: 160 DHACWERPEDMDTPRSVFKVDKNNPGSDIAGEIAAALAAASIVFRKCDPSYSNHLLQRAI 219
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT--- 263
V+ FA +Y+G YS L VCPFYCSYSG++DELLWGAAWL +ATN Y N IK
Sbjct: 220 TVFTFADKYRGPYSAGLAPEVCPFYCSYSGYQDELLWGAAWLQKATNNPTYLNYIKANGQ 279
Query: 264 -LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQ 322
LGAD+ ++FSWD+K+ GA +LLS+ L+ K+ ++Y++ A++F+C ++P +SS+Q
Sbjct: 280 ILGADEFDNMFSWDNKHVGARILLSKEFLIQKVKSLEEYKEHADSFICSVIPG--ASSSQ 337
Query: 323 YTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVD 382
YT GGL+FK+ +SN+QYVTS +FLL TYAKY+++ + CG +VTP LRS+AK+QVD
Sbjct: 338 YTPGGLLFKMGESNMQYVTSTSFLLLTYAKYLTSARTVAYCGGSVVTPARLRSIAKKQVD 397
Query: 383 YILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNI 442
Y+LG NPL+MSYMVGYG +P+RIHHRGSSLPS+A HP I C GF F S +PNPN
Sbjct: 398 YLLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRIQCHDGFS-LFTSQSPNPND 456
Query: 443 LVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
LVGA+VGGP+Q+D FPD+RSDY SEP TYIN +VG LAY A
Sbjct: 457 LVGAVVGGPDQNDQFPDERSDYGRSEPATYINAPLVGALAYLA 499
>dbj|BAA85150.1| endo-1,4-beta-glucanase [Pisum sativum]
Length = 506
Score = 584 bits (1506), Expect = e-165
Identities = 283/461 (61%), Positives = 348/461 (75%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NYR+AL KS+LF QGQRSG+LP +Q++ WR DSGL DG +VDL GGYYDAGDNVKF F
Sbjct: 42 NYRDALTKSILFFQGQRSGKLPSNQRISWRRDSGLSDGSALHVDLVGGYYDAGDNVKFGF 101
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS IE+G M +L A+ A+RW TDYLLK AT+ P +YV VGD DH
Sbjct: 102 PMAFTTTMLSWSVIEFGGLMKSELPNAKKAVRWATDYLLK-ATAHPNIIYVQVGDAKKDH 160
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
CWERPEDMDT R+V+ V N PGS+VAAETAAALAAASLVF+K DPTY+K+LVR A +V
Sbjct: 161 ACWERPEDMDTPRSVFKVDANAPGSEVAAETAAALAAASLVFRKSDPTYAKILVRRAIRV 220
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+QFA +++ SYS++L VCPFYCSYSG++D LLWGAAWL +AT Y I+T L
Sbjct: 221 FQFADKHRRSYSNALKPFVCPFYCSYSGYQDGLLWGAAWLHKATKNPMYLKYIQTNGQIL 280
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GA + + F WD+K+ GA +LLS+ L+ K+ Y+ ++NF+C ++P + SSS QYT
Sbjct: 281 GAAEFDNTFGWDNKHVGARILLSKEFLVQNVKSLHDYKGHSDNFVCSLIPGAGSSSAQYT 340
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL+FK+ DSN+QYVTS TFLL TYAKY++ NCG VTP LR++AKRQVDY+
Sbjct: 341 PGGLLFKMSDSNMQYVTSTTFLLVTYAKYLTKSHSVVNCGGTTVTPKRLRTLAKRQVDYL 400
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYGP +P+RIHHRGSSLPS+A HP I C GF +S +PNPNIL+
Sbjct: 401 LGDNPLKMSYMVGYGPRYPQRIHHRGSSLPSMAVHPGKIQCSAGF-GVMNSKSPNPNILM 459
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP+Q D FPD RSDY SEP TY+N +VG LAY A
Sbjct: 460 GAVVGGPDQHDRFPDQRSDYEQSEPATYVNAPLVGTLAYLA 500
>gb|AAB65155.1| acidic cellulase [Citrus sinensis] gi|7488904|pir||T07883 cellulase
(EC 3.2.1.4) - sweet orange
Length = 505
Score = 579 bits (1493), Expect = e-164
Identities = 279/461 (60%), Positives = 343/461 (73%), Gaps = 5/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
+Y +AL KS+LF +GQRSGRLPP+QQL WR +SGL DG +VDL GGYYDAGDNVKF
Sbjct: 40 DYSDALGKSILFFEGQRSGRLPPNQQLTWRGNSGLSDGSSYHVDLVGGYYDAGDNVKFGL 99
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTT+LSWS IE+G M L+ A+AAIRWGTDYLLK +T+TPG LYV VGDPN+DH
Sbjct: 100 PMAFTTTLLSWSVIEFGSSMQNHLENAKAAIRWGTDYLLKASTATPGALYVQVGDPNMDH 159
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
CWERPEDMDT R VY VS NPGSDVAAETAAALAAAS+VFK DP+YS L++TA KV
Sbjct: 160 HCWERPEDMDTPRNVYKVSTQNPGSDVAAETAAALAAASVVFKDSDPSYSTKLLKTAMKV 219
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----L 264
+ FA +Y+GSYSDSL S VCP+YCSYSG+ DELLWGA+WL RA+ Y I++ L
Sbjct: 220 FDFADKYRGSYSDSLNSVVCPYYCSYSGYLDELLWGASWLHRASQNSSYLAYIQSNGHIL 279
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GADD FSWDDK AG VLLS+ L + F Y+ ++N++C ++P S S QYT
Sbjct: 280 GADDDDYSFSWDDKRAGTKVLLSKGFLEKNTQEFQLYKAHSDNYICSLIPGSSSFQAQYT 339
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL +K +SNLQYVT+ FLL TYAKY+S+ CG+ V L ++AK+QVDYI
Sbjct: 340 AGGLFYKASESNLQYVTTTAFLLLTYAKYLSSNGGVATCGSSTVKAENLIALAKKQVDYI 399
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NP +MSYMVG+G +P+ +HHRGSSLPS+ AHP I C+ GFQ + +S +PNPN+L
Sbjct: 400 LGDNPAKMSYMVGFGERYPQHVHHRGSSLPSIHAHPDHIACNDGFQ-YLYSRSPNPNVLT 458
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GAI+GGP+ D F DDR++Y SEP TYIN VG +A+F+
Sbjct: 459 GAILGGPDNRDNFADDRNNYQQSEPATYINAPFVGAVAFFS 499
>pir||S57808 cellulase (EC 3.2.1.4) precursor - tomato gi|924622|gb|AAA80495.1|
endo-1,4-beta-glucanase precursor
Length = 510
Score = 576 bits (1484), Expect = e-163
Identities = 272/461 (59%), Positives = 348/461 (75%), Gaps = 6/461 (1%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NYR+ALAKS+++ +GQRSG+LP Q++ WR DSGL DG+ VDL GGYYDAGDNVKF F
Sbjct: 46 NYRDALAKSIIYFEGQRSGKLPSSQRITWRKDSGLSDGKAMGVDLVGGYYDAGDNVKFGF 105
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDH 148
PMAFTTTMLSWS IE+G M +L A+ AI W T+YLLK AT+ P +YV VGD DH
Sbjct: 106 PMAFTTTMLSWSVIEFGGLMKGELLNAKQAIGWATEYLLK-ATAHPDTIYVQVGDAGSDH 164
Query: 149 KCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
CWERPEDMDT R+VY + N PG++VAAETAAALAAASLVF+K +P+YSK+L++ A +V
Sbjct: 165 SCWERPEDMDTPRSVYKIDKNTPGTEVAAETAAALAAASLVFRKCNPSYSKILIKRAIRV 224
Query: 209 YQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIK----TL 264
+ FA +Y+GSYS+ L VCP+YCS SG++DELLWGAAWL RAT Y N I+ TL
Sbjct: 225 FAFADKYRGSYSNGLRKVVCPYYCSVSGYEDELLWGAAWLHRATKNPTYLNYIQRNGQTL 284
Query: 265 GADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYT 324
GA + + F WD+K+ GA +LLS+ L+ + Y+ A+N++C ++P +P+S QYT
Sbjct: 285 GAAETDNTFGWDNKHVGARILLSKSFLVQKLQTLHDYKSHADNYICSLIPGTPASQAQYT 344
Query: 325 QGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYI 384
GGL+FK+ DSN+QYVTS +FLL TYAKY+++ + CG V++TP LR+VAK+QVDY+
Sbjct: 345 PGGLLFKMDDSNMQYVTSTSFLLVTYAKYLTSARMVVKCGGVVITPKRLRNVAKKQVDYL 404
Query: 385 LGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILV 444
LG+NPL+MSYMVGYG +P+RIHHRGSSLPS+A HP I C GF +S +PNPN+LV
Sbjct: 405 LGDNPLKMSYMVGYGARYPQRIHHRGSSLPSVANHPAKIQCRDGFS-VMNSQSPNPNVLV 463
Query: 445 GAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
GA+VGGP++ D FPD+RSDY SEP TYIN +VG L Y A
Sbjct: 464 GAVVGGPDEHDRFPDERSDYEQSEPATYINAPLVGTLTYLA 504
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 915,578,429
Number of Sequences: 2540612
Number of extensions: 41667671
Number of successful extensions: 78508
Number of sequences better than 10.0: 339
Number of HSP's better than 10.0 without gapping: 306
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 76730
Number of HSP's gapped (non-prelim): 395
length of query: 488
length of database: 863,360,394
effective HSP length: 132
effective length of query: 356
effective length of database: 527,999,610
effective search space: 187967861160
effective search space used: 187967861160
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0083.8


