

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0400a.3
(419 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
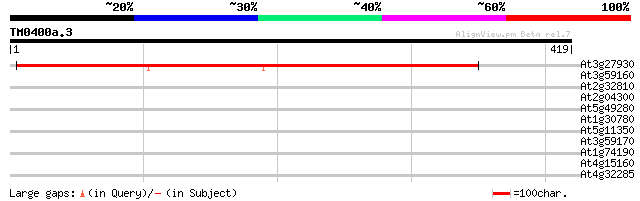
Score E
Sequences producing significant alignments: (bits) Value
At3g27930 unknown protein 449 e-126
At3g59160 putative protein 34 0.13
At2g32810 putative beta-galactosidase (BGAL9) 32 0.50
At2g04300 putative receptor-like protein kinase 32 0.65
At5g49280 predicted GPI-anchored protein 30 2.5
At1g30780 hypothetical protein 30 2.5
At5g11350 unknown protein 29 4.2
At3g59170 putative protein 29 4.2
At1g74190 disease resistance protein, putative 29 5.5
At4g15160 cell wall protein like 28 7.2
At4g32285 unknown protein 28 9.4
>At3g27930 unknown protein
Length = 425
Score = 449 bits (1155), Expect = e-126
Identities = 222/367 (60%), Positives = 271/367 (73%), Gaps = 22/367 (5%)
Query: 6 VKPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTTRIM 65
VK PPP VVLVPPLFD+PPL+AR RMLESSY+++FGKLALRCLF DYF+ FT + +
Sbjct: 6 VKKEPPPPVVLVPPLFDYPPLSARTRMLESSYNLLFGKLALRCLFEDYFEEANRFTGKFL 65
Query: 66 FKPIDEPHVDLIATVSGPLDHKPEESITGNALFRWQR------------------ILRMR 107
KP D+PHVDL+A+VSG +D + E GNA FRWQ +L MR
Sbjct: 66 LKPTDDPHVDLVASVSGAVDGRVEGDFVGNAEFRWQSDVDDPHTFVDLSVSTSNPVLLMR 125
Query: 108 SCAYYPRYGFGAFGVFPLLLR-KREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKS 166
S AYYP+YG GAF V+PL+ + + SE+Y +MGLRYGS NLS G T+ PF+ +ELPK
Sbjct: 126 SSAYYPKYGIGAFAVYPLISKITGKSSEEYRIMGLRYGSTNLSVGATVTPFSANNELPKH 185
Query: 167 AWLVSKMGRLTAGVQYEPQHGN---AKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVK 223
AWLVSKMG LT GVQYEP HG+ AK ++ NWSCA YGVGS SPL+PSFN +EL +
Sbjct: 186 AWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQSPLTPSFNIGIELAR 245
Query: 224 SSQFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRI 283
SSQF+ASFYQH+VVQRRV+NP EEN V+GITNYIDFGFELQ+ VDD+ N DS ++
Sbjct: 246 SSQFIASFYQHVVVQRRVQNPFEENQVVGITNYIDFGFELQSRVDDSKTPPNAPDSLLQV 305
Query: 284 GASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSES 343
ASWQANKNFLLK KVG S+++LAFKSWWKPSF F+ISAT + G +Q GFG++ ++
Sbjct: 306 AASWQANKNFLLKGKVGAHSSTLSLAFKSWWKPSFAFNISATTNHRTGNVQCGFGLRVDN 365
Query: 344 LREASFQ 350
LREAS+Q
Sbjct: 366 LREASYQ 372
>At3g59160 putative protein
Length = 464
Score = 34.3 bits (77), Expect = 0.13
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYID 258
S S LS + + L V + +F S Y H +RVKNPL E G++G +D
Sbjct: 37 STSLLSRRWRYLLAFVPNLEFDDSAYLHR--DKRVKNPLHEKGLVGFVLTVD 86
>At2g32810 putative beta-galactosidase (BGAL9)
Length = 887
Score = 32.3 bits (72), Expect = 0.50
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 15/98 (15%)
Query: 120 FGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKD-ELPKSAWLVSKMGRLTA 178
FG FP+ LR + G+ + + N F + F K +L + A L G
Sbjct: 136 FGGFPVWLRD--------IPGIEFRTDNEPFKKEMQKFVTKIVDLMREAKLFCWQGGPII 187
Query: 179 GVQYEPQHGNAKLS------NLMNWSCAMAYGVGSGSP 210
+Q E ++G+ + S + + W+ +MA G+G+G P
Sbjct: 188 MLQIENEYGDVEKSYGQKGKDYVKWAASMALGLGAGVP 225
>At2g04300 putative receptor-like protein kinase
Length = 851
Score = 32.0 bits (71), Expect = 0.65
Identities = 28/97 (28%), Positives = 44/97 (44%), Gaps = 14/97 (14%)
Query: 207 SGSPLSPS--FNFSLELVKSSQFVASFYQHMVVQRRVKNPLEE-----NG------VIGI 253
+ +P+S + FNF+ L+ S+ S+ +Q N E NG + +
Sbjct: 256 ASTPISKNAPFNFTWSLIPSTAKFYSYMHFADIQTLQANETREFDMMLNGNLALERALEV 315
Query: 254 TNYIDFGFELQTSVDDAIATNNISDSTFRIGASWQAN 290
IDF EL+T+ DD IA NI ++ SWQ +
Sbjct: 316 FTVIDFP-ELETNQDDVIAIKNIQNTYGVSKTSWQGD 351
>At5g49280 predicted GPI-anchored protein
Length = 162
Score = 30.0 bits (66), Expect = 2.5
Identities = 13/39 (33%), Positives = 20/39 (50%)
Query: 8 PPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLAL 46
PPP P+V P + PP + + SSY ++F A+
Sbjct: 119 PPPNPIVPYFPFYYHTPPPGSGSDRFMSSYSIIFALFAV 157
>At1g30780 hypothetical protein
Length = 481
Score = 30.0 bits (66), Expect = 2.5
Identities = 15/27 (55%), Positives = 18/27 (66%), Gaps = 2/27 (7%)
Query: 2 FRKEVKPPPPP--LVVLVPPLFDFPPL 26
F++ V PPPPP L +L PPL D P L
Sbjct: 158 FQEAVAPPPPPPDLPLLAPPLPDVPLL 184
>At5g11350 unknown protein
Length = 754
Score = 29.3 bits (64), Expect = 4.2
Identities = 15/44 (34%), Positives = 23/44 (52%), Gaps = 4/44 (9%)
Query: 372 SYNILGVDSASKHQDLYSNIRHSFQNNGFKGVGKDYTGDARDGC 415
S +I+ + K QDL ++H G+ + K TG+A DGC
Sbjct: 229 SADIMCLQEVDKFQDLEEEMKH----RGYSAIWKMRTGNAVDGC 268
>At3g59170 putative protein
Length = 473
Score = 29.3 bits (64), Expect = 4.2
Identities = 18/46 (39%), Positives = 24/46 (52%), Gaps = 2/46 (4%)
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIG 252
S S LS + + L V + +F S Y H +RVKN L E G +G
Sbjct: 31 STSLLSRRWRYLLAFVPNLEFDDSVYLHR--DKRVKNTLHEKGFVG 74
>At1g74190 disease resistance protein, putative
Length = 965
Score = 28.9 bits (63), Expect = 5.5
Identities = 17/53 (32%), Positives = 29/53 (54%), Gaps = 2/53 (3%)
Query: 274 NNISDSTFRIGASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATR 326
+N + +F G+ AN + L+ K+ + SS+ + +S WKP F S+ A R
Sbjct: 301 DNDFEGSFSFGSL--ANLSNLMVLKLCSKSSSLQVLSESSWKPKFQLSVIALR 351
>At4g15160 cell wall protein like
Length = 428
Score = 28.5 bits (62), Expect = 7.2
Identities = 12/20 (60%), Positives = 13/20 (65%)
Query: 6 VKPPPPPLVVLVPPLFDFPP 25
VKPPPPP V PP + PP
Sbjct: 95 VKPPPPPTVKPPPPPYVKPP 114
>At4g32285 unknown protein
Length = 635
Score = 28.1 bits (61), Expect = 9.4
Identities = 15/58 (25%), Positives = 28/58 (47%)
Query: 157 FAKKDELPKSAWLVSKMGRLTAGVQYEPQHGNAKLSNLMNWSCAMAYGVGSGSPLSPS 214
+ + E+ K +L+++ +L Q E G A L+ + AM YG+ + + PS
Sbjct: 568 YVQMAEMDKKQYLLTQEQQLWQQYQQEGMRGQASLAKMNTAQTAMPYGMPPVNGMGPS 625
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,999,827
Number of Sequences: 26719
Number of extensions: 434919
Number of successful extensions: 1865
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1834
Number of HSP's gapped (non-prelim): 24
length of query: 419
length of database: 11,318,596
effective HSP length: 102
effective length of query: 317
effective length of database: 8,593,258
effective search space: 2724062786
effective search space used: 2724062786
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0400a.3


