

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0249.1
(68 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
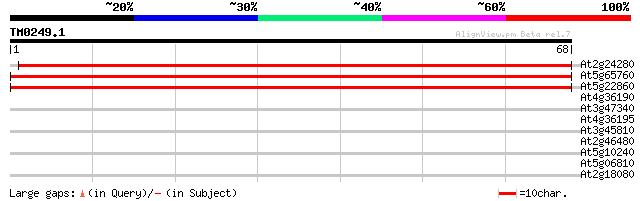
Sequences producing significant alignments: (bits) Value
At2g24280 putative prolylcarboxypeptidase 95 6e-21
At5g65760 lysosomal Pro-X carboxypeptidase 95 7e-21
At5g22860 prolylcarboxypeptidase-like protein 72 5e-14
At4g36190 prolyl carboxypeptidase like protein 26 4.2
At3g47340 glutamine-dependent asparagine synthetase 25 5.5
At4g36195 prolyl carboxypeptidase like protein 25 7.1
At3g45810 respiratory burst oxidase - like protein 25 7.1
At2g46480 hypothetical protein 25 7.1
At5g10240 asparagine synthetase ASN3 25 9.3
At5g06810 putative protein 25 9.3
At2g18080 unknown protein 25 9.3
>At2g24280 putative prolylcarboxypeptidase
Length = 494
Score = 95.1 bits (235), Expect = 6e-21
Identities = 44/67 (65%), Positives = 55/67 (81%)
Query: 2 IEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWL 61
IE LKR SNIIF NG++DPWS GGVLKNIS ++VA+V K+GAHH DLR +TK+DP+WL
Sbjct: 409 IETVLKRFGSNIIFSNGMQDPWSRGGVLKNISSSIVALVTKKGAHHADLRAATKDDPEWL 468
Query: 62 KDVRIKE 68
K+ R +E
Sbjct: 469 KEQRRQE 475
>At5g65760 lysosomal Pro-X carboxypeptidase
Length = 515
Score = 94.7 bits (234), Expect = 7e-21
Identities = 47/68 (69%), Positives = 50/68 (73%)
Query: 1 DIEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKW 60
DI LK SNIIF NGL DPWSGG VLKN+S T+VA+V KEGAHH DLR ST EDPKW
Sbjct: 422 DIATTLKSFGSNIIFSNGLLDPWSGGSVLKNLSDTIVALVTKEGAHHLDLRPSTPEDPKW 481
Query: 61 LKDVRIKE 68
L D R E
Sbjct: 482 LVDQREAE 489
>At5g22860 prolylcarboxypeptidase-like protein
Length = 502
Score = 72.0 bits (175), Expect = 5e-14
Identities = 36/68 (52%), Positives = 48/68 (69%)
Query: 1 DIEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKW 60
+++ L++ SNIIF NGL DP+S GGVL++IS TLVAI K G+H D+ +KEDP+W
Sbjct: 413 EVKLILQKFGSNIIFSNGLSDPYSVGGVLEDISDTLVAITTKNGSHCLDITLKSKEDPEW 472
Query: 61 LKDVRIKE 68
L R KE
Sbjct: 473 LVIQREKE 480
>At4g36190 prolyl carboxypeptidase like protein
Length = 482
Score = 25.8 bits (55), Expect = 4.2
Identities = 10/17 (58%), Positives = 13/17 (75%)
Query: 7 KRSASNIIFYNGLRDPW 23
K +A+ IIF NG +DPW
Sbjct: 395 KIAATKIIFTNGSQDPW 411
>At3g47340 glutamine-dependent asparagine synthetase
Length = 584
Score = 25.4 bits (54), Expect = 5.5
Identities = 15/35 (42%), Positives = 18/35 (50%)
Query: 21 DPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTK 55
DP SG L N KT+V V E +H +LR K
Sbjct: 55 DPASGDQPLFNEDKTIVVTVNGEIYNHEELRKRLK 89
>At4g36195 prolyl carboxypeptidase like protein
Length = 488
Score = 25.0 bits (53), Expect = 7.1
Identities = 9/15 (60%), Positives = 12/15 (80%)
Query: 9 SASNIIFYNGLRDPW 23
+A+ IIF NG +DPW
Sbjct: 397 AATKIIFTNGSQDPW 411
>At3g45810 respiratory burst oxidase - like protein
Length = 835
Score = 25.0 bits (53), Expect = 7.1
Identities = 9/13 (69%), Positives = 12/13 (92%)
Query: 24 SGGGVLKNISKTL 36
+GGG+LKN+SK L
Sbjct: 47 AGGGILKNVSKNL 59
>At2g46480 hypothetical protein
Length = 528
Score = 25.0 bits (53), Expect = 7.1
Identities = 12/27 (44%), Positives = 17/27 (62%), Gaps = 2/27 (7%)
Query: 4 KDLKRSASNIIFYNGLRDPWSGGGVLK 30
K ++RSA +I YNG PW+ G+ K
Sbjct: 481 KKIERSA--VIHYNGHMKPWTEMGISK 505
>At5g10240 asparagine synthetase ASN3
Length = 578
Score = 24.6 bits (52), Expect = 9.3
Identities = 14/35 (40%), Positives = 18/35 (51%)
Query: 21 DPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTK 55
DP SG L N KT+ V E +H+ LR + K
Sbjct: 55 DPTSGDQPLYNEDKTIAVTVNGEIYNHKALRENLK 89
>At5g06810 putative protein
Length = 1141
Score = 24.6 bits (52), Expect = 9.3
Identities = 15/64 (23%), Positives = 29/64 (44%)
Query: 4 KDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKD 63
KD++ I L W G LK S L+ + +G + ++ + +E KW+
Sbjct: 907 KDIEMDDDEIGKIFRLHSLWIGVSRLKQTSTLLINLKGGKGRLCQVIQENPEEMKKWIMG 966
Query: 64 VRIK 67
+R++
Sbjct: 967 LRVQ 970
>At2g18080 unknown protein
Length = 365
Score = 24.6 bits (52), Expect = 9.3
Identities = 9/15 (60%), Positives = 11/15 (73%)
Query: 9 SASNIIFYNGLRDPW 23
+A+ IIF NG DPW
Sbjct: 274 AATKIIFTNGSEDPW 288
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,529,500
Number of Sequences: 26719
Number of extensions: 47622
Number of successful extensions: 95
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 85
Number of HSP's gapped (non-prelim): 11
length of query: 68
length of database: 11,318,596
effective HSP length: 44
effective length of query: 24
effective length of database: 10,142,960
effective search space: 243431040
effective search space used: 243431040
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0249.1


