

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0246c.3
(286 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
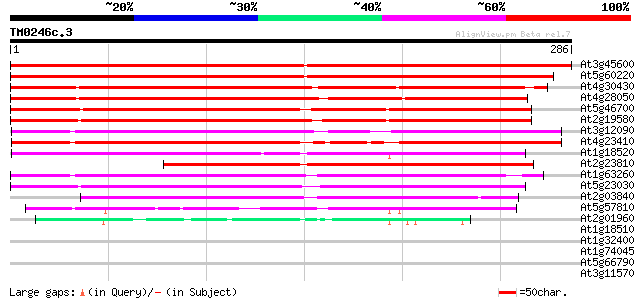
Score E
Sequences producing significant alignments: (bits) Value
At3g45600 unknown protein 472 e-134
At5g60220 senescence-associated protein - like 424 e-119
At4g30430 senescence-associated protein homolog 254 3e-68
At4g28050 senescence-associated protein -like 249 9e-67
At5g46700 senescence-associated protein 5-like protein 243 1e-64
At2g19580 putative senescence-associated protein 5 241 2e-64
At3g12090 putative protein 231 3e-61
At4g23410 unknown protein 213 1e-55
At1g18520 unknown protein 201 3e-52
At2g23810 similar to senescence-associated protein 190 9e-49
At1g63260 unknown protein 182 2e-46
At5g23030 senescence-associated protein 5-like protein 159 2e-39
At2g03840 putative senescence-associated protein 128 3e-30
At5g57810 unknown protein 105 2e-23
At2g01960 unknown protein 44 8e-05
At1g18510 hypothetical protein 39 0.003
At1g32400 unknown protein 37 0.009
At1g74045 putative protein 35 0.061
At5g66790 unknown protein 31 0.67
At3g11570 hypothetical protein 28 4.3
>At3g45600 unknown protein
Length = 285
Score = 472 bits (1215), Expect = e-134
Identities = 222/286 (77%), Positives = 252/286 (87%), Gaps = 1/286 (0%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MRTSNHLIG++NFLTFLLSIPILGGGIWLSSRAN+TDCL+FLQWPLI+IG+SIMVVSLAG
Sbjct: 1 MRTSNHLIGLVNFLTFLLSIPILGGGIWLSSRANSTDCLRFLQWPLIVIGISIMVVSLAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
FAGACYRN FLM LYLVVM L+IA LIGFIIFAY VTDKGSGR VLNRGY+DYYL+DYSG
Sbjct: 61 FAGACYRNKFLMWLYLVVMLLIIAALIGFIIFAYAVTDKGSGRTVLNRGYLDYYLEDYSG 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV+ SYWGKI+SC+RDS C K+GR + NG+PE AD+F+LR+L+ V+SGCCKPPT
Sbjct: 121 WLKDRVSDDSYWGKISSCLRDSGACRKIGR-NFNGVPETADMFFLRRLSPVESGCCKPPT 179
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCG+ Y NET W+ G++ N DC WSNDQ LCY+C SCKAGVL SLKKSWRKVSVI
Sbjct: 180 DCGFSYVNETGWDTRGGMIGPNQDCMVWSNDQSMLCYQCSSCKAGVLGSLKKSWRKVSVI 239
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 286
NIVV+IILVI Y+IAYAAYRN KR+DNDEP GEARMTKSHPSHF L
Sbjct: 240 NIVVLIILVIFYVIAYAAYRNVKRIDNDEPAGEARMTKSHPSHFHL 285
>At5g60220 senescence-associated protein - like
Length = 327
Score = 424 bits (1090), Expect = e-119
Identities = 203/277 (73%), Positives = 230/277 (82%), Gaps = 1/277 (0%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MR+ ++LIG++NF TFLLSIPILGGGIWLSSRAN+TDCL+FLQWPLIIIG+SIMV+SLAG
Sbjct: 1 MRSRSNLIGLINFFTFLLSIPILGGGIWLSSRANSTDCLRFLQWPLIIIGISIMVISLAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
AGACY+N FLM LYL MF VIA LIGF IFAYVVTDKGSGR V+NR Y+DYYL DYSG
Sbjct: 61 IAGACYQNKFLMWLYLFTMFFVIAALIGFTIFAYVVTDKGSGRFVMNRRYLDYYLNDYSG 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV + YW I SCVRDS VC K+GR D NG+PE A +FY R L+ V+SGCCKPPT
Sbjct: 121 WLKDRVTDNGYWRDIGSCVRDSGVCKKIGR-DLNGVPETAHMFYFRNLSPVESGCCKPPT 179
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCGY Y NETVW G ++ NPDC W+NDQ LCY+C SCKAGVL SLKKSWRKVSVI
Sbjct: 180 DCGYTYVNETVWIPGGEMVGPNPDCMLWNNDQRLLCYQCSSCKAGVLGSLKKSWRKVSVI 239
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMT 277
NIVV+IILVI Y+IA AAY+N KRM NDEP GEARMT
Sbjct: 240 NIVVVIILVIFYVIACAAYQNVKRMYNDEPVGEARMT 276
>At4g30430 senescence-associated protein homolog
Length = 272
Score = 254 bits (650), Expect = 3e-68
Identities = 121/274 (44%), Positives = 176/274 (64%), Gaps = 9/274 (3%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN L+G+LNF FLLS+PIL GIWLS +A T C +FL P+I +GV +M++++AG
Sbjct: 2 VRFSNSLVGILNFFVFLLSVPILSTGIWLSLKAT-TQCERFLDKPMIALGVFLMIIAIAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
G+C R T+L+ YL VMF +I +++ F IFA+VVT KGSG + + Y +Y L+ YS
Sbjct: 61 VVGSCCRVTWLLWSYLFVMFFLILIVLCFTIFAFVVTSKGSGETIQGKAYKEYRLEAYSD 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL+ RV + +W I SC+ +SK C + V +N FY LT+ +SGCCKP
Sbjct: 121 WLQRRVNNAKHWNSIRSCLYESKFCYNLELVTAN---HTVSDFYKEDLTAFESGCCKPSN 177
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DC + Y T WN SG N DC W N++ LCY C +CKAG L +LK +W++V+++
Sbjct: 178 DCDFTYITSTTWNKTSG-THKNSDCQLWDNEKHKLCYNCKACKAGFLDNLKAAWKRVAIV 236
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEA 274
NI+ +++LV+VY + A+RNNK ++ YG +
Sbjct: 237 NIIFLVLLVVVYAMGCCAFRNNK----EDRYGRS 266
>At4g28050 senescence-associated protein -like
Length = 263
Score = 249 bits (637), Expect = 9e-67
Identities = 121/265 (45%), Positives = 174/265 (65%), Gaps = 7/265 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
++ SN+L+G+LNF TFLLSIPIL GIWL A T+C +FL P++++G+ +M VS+AG
Sbjct: 2 VQCSNNLLGILNFFTFLLSIPILSAGIWLGKNAA-TECERFLDKPMVVLGIFLMFVSIAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
GAC R + L+ LYL MFL+I + F IFA+ VT++G+G + +RGY +Y++ DYS
Sbjct: 61 LVGACCRVSCLLWLYLFAMFLLILLGFCFTIFAFAVTNRGAGEVISDRGYKEYHVADYSN 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAK-MGRVDSNGIPEPADVFYLRKLTSVQSGCCKPP 179
WL++RV + W +I SC+ S VC+ R S + + FY L ++QSGCCKP
Sbjct: 121 WLQKRVNNAKNWERIRSCLMYSDVCSTYRTRYASINVED----FYKSNLNALQSGCCKPS 176
Query: 180 TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
DC + Y N T W G N DC+ W N G LCY C++CKAG+L ++K SW+KV+
Sbjct: 177 NDCNFTYVNPTTWTKTPGPYK-NEDCNVWDNKPGTLCYDCEACKAGLLDNIKNSWKKVAK 235
Query: 240 INIVVMIILVIVYIIAYAAYRNNKR 264
+NIV +I L+IVY + A+RNN++
Sbjct: 236 VNIVFLIFLIIVYSVGCCAFRNNRK 260
>At5g46700 senescence-associated protein 5-like protein
Length = 269
Score = 243 bits (619), Expect = 1e-64
Identities = 121/266 (45%), Positives = 172/266 (64%), Gaps = 7/266 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++IG +NF+T LLSIP++G GIWL+ N+ C+K LQWP+II+GV I++V LAG
Sbjct: 1 MPLSNNVIGCINFITVLLSIPVIGAGIWLAIGTVNS-CVKLLQWPVIILGVLILLVGLAG 59
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F G +R T+L+ +YL+ M ++I +L + F Y+VT +GSG +R Y++Y LQD+SG
Sbjct: 60 FIGGFWRITWLLVVYLIAMLILIVLLGCLVGFIYMVTIRGSGHPEPSRAYLEYSLQDFSG 119
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL RV W +I +C+ + +C ++ N A F+ L +QSGCCKPPT
Sbjct: 120 WLRRRVQRSYKWERIRTCLSTTTICPEL-----NQRYTLAQDFFNAHLDPIQSGCCKPPT 174
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
CG+ + N T W + MSA+ DC WSNDQ LCY CDSCKAG+LA++K W K +
Sbjct: 175 KCGFTFVNPTYW-ISPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVDWLKADIF 233
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMD 266
++ +I L+IVYII A+RN + D
Sbjct: 234 LLLALIGLIIVYIIGCCAFRNAETED 259
>At2g19580 putative senescence-associated protein 5
Length = 270
Score = 241 bits (616), Expect = 2e-64
Identities = 118/267 (44%), Positives = 173/267 (64%), Gaps = 8/267 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M +N+L +LN L L SIPI GIWL+S+ +N +C+ L+WP++++GV I+VVS G
Sbjct: 1 MALANNLTAILNLLALLCSIPITASGIWLASKPDN-ECVNLLRWPVVVLGVLILVVSATG 59
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F GA L+ +YL M ++I +L+ +IFA+VVT RV RGY +Y L+ +S
Sbjct: 60 FIGAYKYKETLLAVYLCCMAILIGLLLVVLIFAFVVTRPDGSYRVPGRGYKEYRLEGFSN 119
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFY-LRKLTSVQSGCCKPP 179
WL+E V WG++ +C+ D+ VC K+ + AD F+ K+T +QSGCCKPP
Sbjct: 120 WLKENVVDSKNWGRLRACLADTNVCPKLNQEFIT-----ADQFFSSSKITPLQSGCCKPP 174
Query: 180 TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
T CGY + N T+W L M+A+ DC WSNDQ LCY C+SCKAG+L +L+K WRK ++
Sbjct: 175 TACGYNFVNPTLW-LNPTNMAADADCYLWSNDQSQLCYNCNSCKAGLLGNLRKEWRKANL 233
Query: 240 INIVVMIILVIVYIIAYAAYRNNKRMD 266
I I+ +++L+ VY+IA +A+RN + D
Sbjct: 234 ILIITVVVLIWVYVIACSAFRNAQTED 260
>At3g12090 putative protein
Length = 282
Score = 231 bits (589), Expect = 3e-61
Identities = 120/281 (42%), Positives = 166/281 (58%), Gaps = 20/281 (7%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IGVLN LT L SIPI+G ++ + ++T C FLQ PL++IG I++VSLAGF
Sbjct: 3 RFSNTVIGVLNLLTLLASIPIIGTALYKAR--SSTTCENFLQTPLLVIGFIILIVSLAGF 60
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC+ + + +YLVVM +IA L+G +F VVT +G G V R Y +Y L DY W
Sbjct: 61 IGACFNVAWALWVYLVVMIFLIATLMGLTLFGLVVTSQGGGVEVPGRIYKEYRLGDYHPW 120
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L ERV YW I SC+ SK C K+ + ++ R +TSVQSGCCKPPT
Sbjct: 121 LRERVRDPEYWNSIRSCILSSKTCTKIESWTTLD-------YFQRDMTSVQSGCCKPPTA 173
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C Y +G++ DC +W+N LCY CD+CKAGVL ++ WRK+SV+N
Sbjct: 174 CTY----------EAGVVDGGGDCFRWNNGVEMLCYECDACKAGVLEEIRLDWRKLSVVN 223
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDND-EPYGEARMTKSHP 281
I+V+++L+ VY A+ N + + P + RMT+ P
Sbjct: 224 ILVLVLLIAVYAAGCCAFHNTRHAAHPYHPSDDNRMTRVRP 264
>At4g23410 unknown protein
Length = 281
Score = 213 bits (541), Expect = 1e-55
Identities = 110/282 (39%), Positives = 170/282 (60%), Gaps = 21/282 (7%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IG LN LT + SI +LG +W+ + T C FLQ PL+I+G++I+++S+AG
Sbjct: 3 RMSNTVIGFLNILTLISSIVLLGSALWMGR--SKTTCEHFLQKPLLILGLAILILSVAGL 60
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC +++ +YL M +I L+G +F ++VT G V R Y ++ L+ Y W
Sbjct: 61 VGACCDVAWVLWVYLFFMVFIIVALMGLTLFGFIVTSHSGGVVVDGRVYKEFKLEAYHPW 120
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRK-LTSVQSGCCKPPT 180
L+ RV +YW I +C+ S C+K+ + P D YL+K L+ +QSGCCKPPT
Sbjct: 121 LKTRVVDTNYWVTIKTCLLGSVTCSKL------ALWTPLD--YLQKDLSPLQSGCCKPPT 172
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
C +Y +TV + +PDC +W+N LCY CD+C+AGVL ++++ W K+S++
Sbjct: 173 SC--VYNTDTV-------IQQDPDCYRWNNAATVLCYDCDTCRAGVLETVRRDWHKLSLV 223
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDN-DEPYGEARMTKSHP 281
N++V+I L+ VY + A++N KR + PYG M+KS P
Sbjct: 224 NVIVVIFLIAVYCVGCCAFKNAKRPQHYGFPYGRYGMSKSRP 265
>At1g18520 unknown protein
Length = 271
Score = 201 bits (512), Expect = 3e-52
Identities = 100/265 (37%), Positives = 154/265 (57%), Gaps = 7/265 (2%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN ++G+ N L L+ +G I++ TDC ++ PL+ G+ + +VSL G
Sbjct: 3 RVSNFMVGLANTLVMLVGASAIGYSIYMFVHQGVTDCESAIRIPLLTTGLILFLVSLLGV 62
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
G+C++ M YL+++F I L+ F IF + VT+KG+GR V RGY +Y D+S W
Sbjct: 63 IGSCFKENLAMVSYLIILFGGIVALMIFSIFLFFVTNKGAGRVVSGRGYKEYRTVDFSTW 122
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L V W I SC+ ++ VC + + + AD FY + L+ +QSGCCKPP+D
Sbjct: 123 LNGFVGG-KRWVGIRSCLAEANVCDDL---SDGRVSQIADAFYHKNLSPIQSGCCKPPSD 178
Query: 182 CGYIYQNETVW---NLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVS 238
C + ++N T W + ++ N DC WSN Q LC+ C++CKAGVLA++++ WR +
Sbjct: 179 CNFEFRNATFWIPPSKNETAVAENGDCGTWSNVQTELCFNCNACKAGVLANIREKWRNLL 238
Query: 239 VINIVVMIILVIVYIIAYAAYRNNK 263
V NI ++I+L+ VY A RNN+
Sbjct: 239 VFNICLLILLITVYSCGCCARRNNR 263
>At2g23810 similar to senescence-associated protein
Length = 195
Score = 190 bits (482), Expect = 9e-49
Identities = 84/189 (44%), Positives = 126/189 (66%), Gaps = 3/189 (1%)
Query: 79 MFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASC 138
MFL+I ++ +FA+VVT+KG+G + +GY +Y L DYS WL++RV + W KI SC
Sbjct: 1 MFLLILLVFCITVFAFVVTNKGAGEAIEGKGYKEYKLGDYSTWLQKRVENGKNWNKIRSC 60
Query: 139 VRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETVWNLGSGL 198
+ +SKVC+K+ ++ + P + FY LT++QSGCCKP +CG+ Y N T W +
Sbjct: 61 LVESKVCSKL---EAKFVNVPVNSFYKEHLTALQSGCCKPSDECGFEYVNPTTWTKNTTG 117
Query: 199 MSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAA 258
NPDC W N + LC+ C SCKAG+L ++K +W+KV+++NIV ++ L+IVY + A
Sbjct: 118 THTNPDCQTWDNAKEKLCFDCQSCKAGLLDNVKSAWKKVAIVNIVFLVFLIIVYSVGCCA 177
Query: 259 YRNNKRMDN 267
+RNNKR D+
Sbjct: 178 FRNNKRDDS 186
>At1g63260 unknown protein
Length = 284
Score = 182 bits (462), Expect = 2e-46
Identities = 89/272 (32%), Positives = 158/272 (57%), Gaps = 15/272 (5%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M TS +I +N LT LL++ ++ G+W+S+ +N C + L +P+I +G I ++S+ G
Sbjct: 3 MGTSTFVIRWVNLLTMLLAVAVIIFGVWMST--HNDGCRRSLTFPVIALGGFIFLISIIG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F GAC R+ L+ +YL V+ +V+ ++ F + A++VT+ GSG Y +Y L DYS
Sbjct: 61 FLGACKRSVALLWIYLAVLLIVLIAILVFTVLAFIVTNNGSGHTNPGLRYKEYKLNDYSS 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
W +++ + S W ++ SC+ S+ C K+ + + +LT +++GCC+PP+
Sbjct: 121 WFLKQLNNTSNWIRLKSCLVKSEQCRKLSKK-----YKTIKQLKSAELTPIEAGCCRPPS 175
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
+CGY N + ++L +S+N DC + N + CY CDSCKAGV +K WR V++
Sbjct: 176 ECGYPAVNASYYDLSFHSISSNKDCKLYKNLRTIKCYNCDSCKAGVAQYMKTEWRLVAIF 235
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYG 272
N+V+ ++L+ + + R D+++ +G
Sbjct: 236 NVVLFVVLISSLL--------STRFDSEQSFG 259
>At5g23030 senescence-associated protein 5-like protein
Length = 264
Score = 159 bits (401), Expect = 2e-39
Identities = 82/265 (30%), Positives = 141/265 (52%), Gaps = 12/265 (4%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN + N + L+ + L +++ + + C +F+Q PLI+ + +S G
Sbjct: 2 LRLSNAAVITTNAILALIGLAALSFSVYVYVQGPS-QCQRFVQNPLIVTAALLFFISSLG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
A Y + ++ LYL +FL I +L+ +F ++VT+ +G+ + RG + DY
Sbjct: 61 LIAALYGSHIIITLYLFFLFLSILLLLVLSVFIFLVTNPTAGKALSGRGIGNVKTGDYQN 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADV-FYLRKLTSVQSGCCKPP 179
W+ W I C+ DS+VC + G P D+ F + L++VQ GCC+PP
Sbjct: 121 WIGNHFLRGKNWEGITKCLSDSRVCKRFG---------PRDIDFDSKHLSNVQFGCCRPP 171
Query: 180 TDCGYIYQNETVWNLGSGLMSA-NPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVS 238
+CG+ +N T W + + +A DC WSN Q LCY C+SCK GVL ++K WR +
Sbjct: 172 VECGFESKNATWWTVPATATTAIIGDCKAWSNTQRQLCYACESCKIGVLKGIRKRWRILI 231
Query: 239 VINIVVMIILVIVYIIAYAAYRNNK 263
V+N+++++++V +Y +NN+
Sbjct: 232 VVNLLLILLVVFLYSCGCCVRKNNR 256
>At2g03840 putative senescence-associated protein
Length = 278
Score = 128 bits (322), Expect = 3e-30
Identities = 65/223 (29%), Positives = 111/223 (49%), Gaps = 7/223 (3%)
Query: 37 DCLKFLQWPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVV 96
+C +F+ P I I S++ +SL GF A +++ L R++ + FL + V++ IF +
Sbjct: 54 ECNRFVTTPGIFISFSLLAMSLTGFYAAYFKSDCLFRIHFFIFFLWMFVVVSKAIFVIFL 113
Query: 97 TDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGI 156
+ + R ++ +DYSGW+ V W + C+ VC ++
Sbjct: 114 HKETNPRLFPGTKIYEFRYEDYSGWVSRLVIKDDEWYRTRRCLVKDNVCNRLNH------ 167
Query: 157 PEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLC 216
PA FY LT +QSGCCKPP CG Y+ W + + DC +W+N LC
Sbjct: 168 KMPASEFYQMNLTPIQSGCCKPPLSCGLNYEKPNNWTVSRYYNNLEVDCKRWNNSADTLC 227
Query: 217 YRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAY 259
+ CDSCKA ++A + + ++V NI+ +I + + + + A+
Sbjct: 228 FDCDSCKAVIIADVHNTSFSITV-NIIHIIFSLCIGMTGWFAW 269
>At5g57810 unknown protein
Length = 317
Score = 105 bits (263), Expect = 2e-23
Identities = 78/268 (29%), Positives = 120/268 (44%), Gaps = 36/268 (13%)
Query: 9 GVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLI--IIGVSIMVVSLAGFAGACY 66
GVL TF+LS+ +LG +WL + DC L P + + V ++ V + A
Sbjct: 60 GVLPIFTFVLSLTLLGYAVWLLYM-RSYDCEDILGLPRVQTLASVGLLAVFVVSNAALFL 118
Query: 67 RNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRV-LNRGYMDYYLQDYSGWLEER 125
R F M LVVM +V+ +++ FI AY ++ RR R + + D
Sbjct: 119 RRKFPMPA-LVVMVVVLLLML-FIGLAYAGVNEMQSRRFPATRMWFKLKIMD-------- 168
Query: 126 VASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCGYI 185
H W I SCV D C + N P + RK+ +++GCC PP C
Sbjct: 169 --DHVTWNNIKSCVYDKGACNDLIYGSPNEKP-----YNRRKMPPIKNGCCMPPETCNMD 221
Query: 186 YQNETVW------------NLGSG---LMSANPDCSKWSNDQGFLCYRCDSCKAGVLASL 230
N T W NL G ++ DC W ND LCY C SCK G + S+
Sbjct: 222 AINATFWYRRKDEGPPSSMNLMYGDEMMVGRISDCQLWRNDWSILCYDCRSCKFGFIRSV 281
Query: 231 KKSWRKVSVINIVVMIILVIVYIIAYAA 258
++ W ++ + IV+ I+L++ +++ + A
Sbjct: 282 RRKWWQLGIFLIVISILLLMSHLLIFLA 309
>At2g01960 unknown protein
Length = 260
Score = 44.3 bits (103), Expect = 8e-05
Identities = 66/245 (26%), Positives = 100/245 (39%), Gaps = 40/245 (16%)
Query: 14 LTFLLSIPILGGGIWLSSRANNTDCLKFLQWPL----IIIGVSIMVVSLAGFAGACYRNT 69
+TF LS P++G ++L N+ + Q L ++ VS++ + L G
Sbjct: 20 ITFFLSAPLVGHALYLFCMRNDHVYYRDFQSTLPRVQTLVSVSLLALFLLSNIG------ 73
Query: 70 FLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGWLEERVASH 129
+R + FLVI IGF AY K RR + +Y+ ER
Sbjct: 74 MFLRPRRLSYFLVIVFFIGF---AYSGVYKMESRRFSPTPMC--FKGEYNNGQGERKTEQ 128
Query: 130 SYWGKIA-SCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQN 188
KI S R +V + V+S +P P D R L SV++GCC P +C N
Sbjct: 129 YQVVKIEQSQGRLQRVHLRF--VNSYALP-PYD---RRLLPSVKTGCCNRPGNCKLETVN 182
Query: 189 ETVW----NLGSGLMSA--------NPDC----SKWSNDQGFLCYRCDSCKAGVLAS--L 230
T+W G L +A N D W ++ L Y C +C+ ++ S L
Sbjct: 183 ATLWVTRNREGPPLETAMIYDRYGGNADIKDYYDMWRHELSVLYYDCMTCQVRIIKSPRL 242
Query: 231 KKSWR 235
+K W+
Sbjct: 243 RKWWQ 247
>At1g18510 hypothetical protein
Length = 238
Score = 38.9 bits (89), Expect = 0.003
Identities = 43/244 (17%), Positives = 85/244 (34%), Gaps = 34/244 (13%)
Query: 17 LLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYL 76
L+ I + G L R + C++ ++IG+ ++V+ C + + +Y+
Sbjct: 6 LICIGLTMTGTGLYYRKTVSKCIRETDGSFVVIGLLLLVIPQFALYAICCHSKRMFTIYI 65
Query: 77 VVMFLVIAVLIGFIIFAYVV-TDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKI 135
M V VL G+ + ++ T G + + + L R+ S K+
Sbjct: 66 YAMIFVSIVLGGYSLKCFIYNTTFGIAKNPAEE-------KRTAKQLVGRLVPESKLAKV 118
Query: 136 ASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCG----YIYQNETV 191
C+ + C +SN V CC P CG + E
Sbjct: 119 TECIIHNHDCNFNASQNSN----------------VWRYCCAQPRGCGVTTMFGQPGEWS 162
Query: 192 WNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIV 251
W +CS C C C+ +L ++ W+ +S+ + + ++ +
Sbjct: 163 WKHQHVENHVPEECSY------EYCLSCRGCQMSILKAIVHQWKYLSMFSYPALFLVCLS 216
Query: 252 YIIA 255
I+
Sbjct: 217 LAIS 220
>At1g32400 unknown protein
Length = 280
Score = 37.4 bits (85), Expect = 0.009
Identities = 16/46 (34%), Positives = 27/46 (57%)
Query: 49 IGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAY 94
IGV++ V+S G G C R+ + Y +++ L+I V +GF F +
Sbjct: 88 IGVALFVISCCGCVGTCSRSVCCLSCYSLLLILLILVELGFAAFIF 133
>At1g74045 putative protein
Length = 215
Score = 34.7 bits (78), Expect = 0.061
Identities = 41/227 (18%), Positives = 80/227 (35%), Gaps = 34/227 (14%)
Query: 36 TDCLKFLQWPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYV 95
+ C++ ++G+ ++++ G G C R+ L + M ++I ++ + I +
Sbjct: 2 SSCIRETSSQFTLLGLLLLLIPQIGLYGICCRSKRLFNFFFYGMVVLIIIVSYYSIKCSI 61
Query: 96 V-TDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSN 154
T G + N + + G R+ S + K+ C+ C +SN
Sbjct: 62 YNTTFGIAK---NPAKDNRTVPQLLG----RLVSKEKFEKVTYCIIHKHDCNYNASKNSN 114
Query: 155 GIPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQ----NETVWNLGSGLMSANPDCSKWSN 210
V CC P CG I E W +CS
Sbjct: 115 ----------------VWKYCCAQPVGCGTITMFDKPGEWSWKHQYERNQVPEECSY--- 155
Query: 211 DQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYA 257
C C C+ +L ++ W+ +S+ +++ I IA++
Sbjct: 156 ---EYCLDCRGCQLSILKAIVHQWKYLSMFAYPALVLSCISLAIAWS 199
>At5g66790 unknown protein
Length = 622
Score = 31.2 bits (69), Expect = 0.67
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 16/84 (19%)
Query: 200 SANPDCSKWSNDQGFLCYRCDSCKAGV-------------LASLKKSWRKVSVINI--VV 244
S N DC+K D G L +RC +C+ G L +K K+ V+ ++
Sbjct: 199 SENADCAKVKLDDGGLGHRC-TCREGFSGKAFTVPGGCHRLVYKRKGLHKLVVLGTAGIL 257
Query: 245 MIILVIVYIIAYAAYRNNKRMDND 268
+ +LVIV +IA +RN + ++
Sbjct: 258 VGVLVIVVLIATYFFRNKQSASSE 281
>At3g11570 hypothetical protein
Length = 427
Score = 28.5 bits (62), Expect = 4.3
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 15/90 (16%)
Query: 79 MFLVIAVLIGFIIFAYVVTDKGSGRRVL-------------NRGYMDYYLQDYSGWLEER 125
+F+ I++LI F+IF+ +V D L RG D Y W+ R
Sbjct: 31 IFVGISLLITFLIFSVIVVDLAGFEPHLCFGFLLSPRTLTKERGNDDVCDYSYGRWVRRR 90
Query: 126 --VASHSYWGKIASCVRDSKVCAKMGRVDS 153
V SY+G+ + C GR DS
Sbjct: 91 RDVDETSYYGEECRFLDPGFRCLNNGRKDS 120
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,728,907
Number of Sequences: 26719
Number of extensions: 283312
Number of successful extensions: 933
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 878
Number of HSP's gapped (non-prelim): 33
length of query: 286
length of database: 11,318,596
effective HSP length: 98
effective length of query: 188
effective length of database: 8,700,134
effective search space: 1635625192
effective search space used: 1635625192
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0246c.3


