

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.8
(334 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
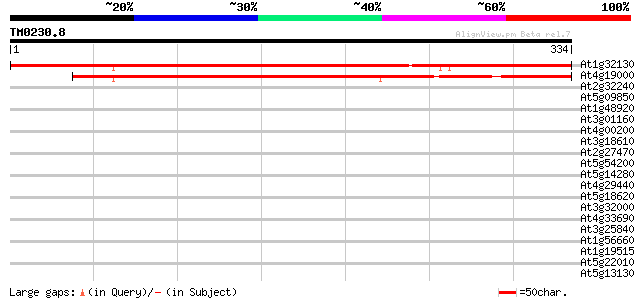
Score E
Sequences producing significant alignments: (bits) Value
At1g32130 unknown protein 498 e-141
At4g19000 putative protein 274 6e-74
At2g32240 putative myosin heavy chain 37 0.015
At5g09850 unknown protein 37 0.020
At1g48920 unknown protein 33 0.17
At3g01160 unknown protein 33 0.22
At4g00200 putative transcription factor 33 0.28
At3g18610 unknown protein 32 0.37
At2g27470 putative CCAAT-binding transcription factor subunit 32 0.37
At5g54200 putative protein 32 0.63
At5g14280 unknown protein 31 0.83
At4g29440 putative protein 31 1.1
At5g18620 chromatin remodelling complex ATPase chain ISWI -like ... 30 1.4
At3g32000 unknown protein 30 1.4
At4g33690 hypothetical protein 30 1.8
At3g25840 protein kinase like protein 30 1.8
At1g56660 hypothetical protein 30 1.8
At1g19515 unknown protein 30 1.8
At5g22010 replication factor C large subunit-like protein 30 2.4
At5g13130 putative protein 30 2.4
>At1g32130 unknown protein
Length = 404
Score = 498 bits (1281), Expect = e-141
Identities = 255/343 (74%), Positives = 295/343 (85%), Gaps = 10/343 (2%)
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGF-YGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK 59
+D+EGVR +DDDNFIDDTG++P YG SP PQAEEGE++DE+N+LFKMGKKK
Sbjct: 62 NDEEGVRTMDDDNFIDDTGLDPSERYGGDAGDRSPTHYPQAEEGEDEDEVNNLFKMGKKK 121
Query: 60 N--ERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLE 117
ER+PAEIALLVENV+AELEVTAEEDAELNRQGKPAINKLKKL LLT+VL KKQLQ E
Sbjct: 122 KKTERNPAEIALLVENVMAELEVTAEEDAELNRQGKPAINKLKKLSLLTDVLGKKQLQTE 181
Query: 118 FLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVI 177
FLDHGVLTLLKNWLEPLPDGSLPNINIR AIL++L DFPIDL+Q DRREQLK+SGLGKVI
Sbjct: 182 FLDHGVLTLLKNWLEPLPDGSLPNINIRAAILRVLTDFPIDLDQYDRREQLKKSGLGKVI 241
Query: 178 MFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKA 237
MFLSKSDEE N NR+L K+LVDKWSRPIFNKSTRFEDMRN++++R P+RRP VKKP+NKA
Sbjct: 242 MFLSKSDEETNSNRRLAKDLVDKWSRPIFNKSTRFEDMRNLDEDRVPYRRPPVKKPSNKA 301
Query: 238 PGMQSRDSDLDLDLPQPR----SGQSS--SRQHASRPEATPMDFVIRPQSKVDPEEVRAR 291
M+SRD D DL++ + + SGQSS RQ RPEATP+DF+IRPQSK+DP+E+ AR
Sbjct: 302 T-MESRDGDFDLEIRERKTGLTSGQSSRGDRQMTMRPEATPLDFLIRPQSKIDPDEIIAR 360
Query: 292 AKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
AKQ S DQ+R+KMNKKLQQL+ KK++LQATK+SVEGRGMIKY
Sbjct: 361 AKQVSQDQRRVKMNKKLQQLKGTKKKRLQATKVSVEGRGMIKY 403
>At4g19000 putative protein
Length = 406
Score = 274 bits (700), Expect = 6e-74
Identities = 155/301 (51%), Positives = 201/301 (66%), Gaps = 11/301 (3%)
Query: 38 PQAEEGEEDDEINDLFKMGKKKN--ERSPAEIALLVENVVAELEVTAEEDAELNRQGKPA 95
P ++ E+ +EI LF + KKK+ +++ EI + VE V+A LE+ E+D NR+GKPA
Sbjct: 112 PPKKKDEDAEEIKKLFSLRKKKSKCDKTSMEIGMQVEQVMANLEIAVEDDVICNREGKPA 171
Query: 96 INKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDF 155
INKL KLPLL E LSKK LQ EFLDHGVL LLKNWLEPLPDGSLPNINIR+A+L ILNDF
Sbjct: 172 INKLMKLPLLNETLSKKPLQGEFLDHGVLNLLKNWLEPLPDGSLPNINIRSAVLMILNDF 231
Query: 156 PIDLEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDM 215
IDL+Q RREQL +SGLGKVIMFLSKSDEE NR+L ++++KW R I+NKSTR+++M
Sbjct: 232 RIDLDQDSRREQLIKSGLGKVIMFLSKSDEETTPNRRLANDIINKWGRIIYNKSTRYDNM 291
Query: 216 RNIE--DERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPM 273
E DE+ K A K G ++RD + D+DL + G + R A P M
Sbjct: 292 FTQEELDEQRQILLRRQTKTAPKVSGTRARDFNTDIDLYE--LGTWTGRARAKIPTTMSM 349
Query: 274 DFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIK 333
DF IRP SKVD + ++ + +M+ K +Q + +K +QA KLSV+GR M+K
Sbjct: 350 DFKIRPPSKVDINQ-----EEEPCSKWQMEKRHKNKQQKNIRKGGMQALKLSVDGRTMLK 404
Query: 334 Y 334
Y
Sbjct: 405 Y 405
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 37.0 bits (84), Expect = 0.015
Identities = 55/257 (21%), Positives = 97/257 (37%), Gaps = 47/257 (18%)
Query: 32 SSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTAEEDAELNRQ 91
+S GE Q++ +E N + M + E + IA L E + E +++ D ++
Sbjct: 1061 TSEGEKLQSQISSHTEENNQVNAMFQSTKEELQSVIAKLEEQLTVE---SSKADTLVSE- 1116
Query: 92 GKPAINKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILK- 150
I KL+ + VL +LE V LK +E S+ + + + +
Sbjct: 1117 ----IEKLRAVAAEKSVLESHFEELEKTLSEVKAQLKENVENAATASVKVAELTSKLQEH 1172
Query: 151 --------ILNDFPIDLEQ--------IDRREQLKRSGLGKVIMFLSKSDEEINVNRKLT 194
+LN+ + L++ ID ++Q ++ L KS EEI +K
Sbjct: 1173 EHIAGERDVLNEQVLQLQKELQAAQSSIDEQKQAHSQKQSELESALKKSQEEIEAKKKAV 1232
Query: 195 KELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP 254
T FE M +++ K+ G++SRD DL P
Sbjct: 1233 ---------------TEFESMVKDLEQKVQLADAKTKETEAMDVGVKSRDIDLSFSSP-- 1275
Query: 255 RSGQSSSRQHASRPEAT 271
+ R+ +PEA+
Sbjct: 1276 -----TKRKSKKKPEAS 1287
>At5g09850 unknown protein
Length = 353
Score = 36.6 bits (83), Expect = 0.020
Identities = 43/169 (25%), Positives = 73/169 (42%), Gaps = 18/169 (10%)
Query: 158 DLEQIDRREQ-LKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPI-----FNKSTR 211
+LE +D Q L+ + +G+ + + K N R+L K+LV KW + FN+
Sbjct: 150 NLEDMDITFQALQETDIGRHVNRVRKHPS--NNVRRLAKQLVKKWKETVDEWVKFNQPGD 207
Query: 212 FEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHA--SRPE 269
E I DE +P V+K + Q D P P++G SSS +++ + PE
Sbjct: 208 LEPPSLIADEDSP-----VQKALHNGSRQQVPDFGYS---PVPQNGYSSSSKNSNITEPE 259
Query: 270 ATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQ 318
P +P+ + +R + R K +K++ A K+ Q
Sbjct: 260 RKPRPVAPQPRRESPSPAKPSRPSPSQQTIPRDKEHKEVDFDTARKRLQ 308
>At1g48920 unknown protein
Length = 557
Score = 33.5 bits (75), Expect = 0.17
Identities = 33/150 (22%), Positives = 59/150 (39%), Gaps = 7/150 (4%)
Query: 183 SDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQS 242
SDEE+ V +K K K + ED + EDE A + KPA K
Sbjct: 131 SDEEVAVTKK--PAAAAKNGSVKAKKESSSEDDSSSEDEPAKKPAAKIAKPAAKDSSSSD 188
Query: 243 RDSDLDL--DLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPE---EVRARAKQASH 297
DSD D + P + ++ + AS +++ D + + + + +A K +S
Sbjct: 189 DDSDEDSEDEKPATKKAAPAAAKAASSSDSSDEDSDEESEDEKPAQKKADTKASKKSSSD 248
Query: 298 DQQRMKMNKKLQQLRAPKKRQLQATKLSVE 327
+ + ++ + PKK+ + E
Sbjct: 249 ESSESEEDESEDEEETPKKKSSDVEMVDAE 278
>At3g01160 unknown protein
Length = 713
Score = 33.1 bits (74), Expect = 0.22
Identities = 22/80 (27%), Positives = 40/80 (49%), Gaps = 6/80 (7%)
Query: 39 QAEEGEEDDEINDLFKMGKKKNERSPAEIALL-VENVVAELEVTAEEDAELNRQGKPAIN 97
+++ EEDD N++ KKK+++ AL+ E+V ++ ++ E D ++ +
Sbjct: 403 ESDSDEEDDLGNEVINQSKKKDKKKDKYRALIEAEDVDSDKDLEEENDQDMEVTFNTGLE 462
Query: 98 KLKKLPLLTEVLSKKQLQLE 117
L K E+L KK Q E
Sbjct: 463 DLSK-----EILKKKDNQSE 477
>At4g00200 putative transcription factor
Length = 345
Score = 32.7 bits (73), Expect = 0.28
Identities = 23/76 (30%), Positives = 35/76 (45%), Gaps = 5/76 (6%)
Query: 227 RPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPE 286
RPS PAN+ G+ S ++ P P SG++S ++ RP + P S V
Sbjct: 26 RPS-DSPANQFMGL----SLPPMEAPMPSSGEASGKKRRGRPRKYEANGAPLPSSSVPLV 80
Query: 287 EVRARAKQASHDQQRM 302
+ R R K D ++M
Sbjct: 81 KKRVRGKLNGFDMKKM 96
>At3g18610 unknown protein
Length = 636
Score = 32.3 bits (72), Expect = 0.37
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 181 SKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSV---KKPANKA 237
S SDEE V +K +V + +S+ E+ + +DE P ++P+V KPA K
Sbjct: 241 SSSDEETPVVKKKPTTVV----KDAKAESSSSEEESSSDDEPTPAKKPTVVKNAKPAAKD 296
Query: 238 PGMQSRDSD---LDLDLPQPRSGQSS---SRQHASRPEATPMDFVIRPQSKVDPEEVRAR 291
DSD D + P + + S S+Q +S E++ D + +SK E+V +
Sbjct: 297 SSSSEEDSDEEESDDEKPPTKKAKVSSKTSKQESSSDESS--DESDKEESK--DEKVTPK 352
Query: 292 AKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEG 328
K + + + +Q + P + +K G
Sbjct: 353 KKDSDVEMVDAEQKSNAKQPKTPTNQTQGGSKTLFAG 389
>At2g27470 putative CCAAT-binding transcription factor subunit
Length = 275
Score = 32.3 bits (72), Expect = 0.37
Identities = 24/93 (25%), Positives = 41/93 (43%), Gaps = 1/93 (1%)
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKN 60
+D+E D+++ DD E N+ + E EE E++DE N + + G ++
Sbjct: 175 NDEEDENGNDEEDENDDENTEENGNDEENDDENTEENGNDEENEKEDEENSMEENG-NES 233
Query: 61 ERSPAEIALLVENVVAELEVTAEEDAELNRQGK 93
E S E + EN E ED ++ G+
Sbjct: 234 EESGNEDHSMEENGSGVGEDNENEDGSVSGSGE 266
>At5g54200 putative protein
Length = 825
Score = 31.6 bits (70), Expect = 0.63
Identities = 29/101 (28%), Positives = 46/101 (44%), Gaps = 13/101 (12%)
Query: 229 SVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEV 288
S+K A+ G + R S D D P R GQ R ++ ++ M F DPE V
Sbjct: 289 SIKNVASSVTGYKERRSTDDRDSPSERGGQ---RFSSATDDSRDMSF-------HDPERV 338
Query: 289 RARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGR 329
+ R Q + + K Q+++A K + + K S++GR
Sbjct: 339 KVR--QYGKSCKELTALFKSQEIQA-HKGSIWSIKFSLDGR 376
>At5g14280 unknown protein
Length = 572
Score = 31.2 bits (69), Expect = 0.83
Identities = 27/90 (30%), Positives = 48/90 (53%), Gaps = 6/90 (6%)
Query: 36 EAPQAEEGEE---DDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTA-EEDAELNRQ 91
+A Q+E GEE +DE L G K+ +SP E ++V+ + + TA +E + +
Sbjct: 168 KAHQSESGEEVFEEDEEVALIDKGAAKSGKSPHEAVVVVDKITTKKNGTAGKESDDDDDD 227
Query: 92 GKPAINKLKKLPLLTEVLS--KKQLQLEFL 119
A+ + ++++ LS +K+LQLE L
Sbjct: 228 VLCAVRDAFETTMMSQGLSDYQKKLQLEKL 257
>At4g29440 putative protein
Length = 1071
Score = 30.8 bits (68), Expect = 1.1
Identities = 21/75 (28%), Positives = 36/75 (48%), Gaps = 2/75 (2%)
Query: 229 SVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEV 288
S K+P++ P S D + D++LP+ S + ++ SR T + + K EE+
Sbjct: 726 SDKRPSSIPPDSSSSDDESDMELPKRVSFRYQEKRTESRTRPTHLHSGV--SHKDLEEEI 783
Query: 289 RARAKQASHDQQRMK 303
RA S D++ K
Sbjct: 784 PTRASTRSQDRRTHK 798
>At5g18620 chromatin remodelling complex ATPase chain ISWI -like
protein
Length = 1069
Score = 30.4 bits (67), Expect = 1.4
Identities = 25/82 (30%), Positives = 38/82 (45%), Gaps = 2/82 (2%)
Query: 39 QAEEGEEDDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINK 98
QA E+DDE+ + + E + A + ++ V +E AEED E + K I+K
Sbjct: 26 QANVEEDDDELEAVARSAGSDEEDVAPDEAPVSDDEVVPVEDDAEEDEE--DEEKAEISK 83
Query: 99 LKKLPLLTEVLSKKQLQLEFLD 120
+K L KKQ + LD
Sbjct: 84 REKARLKEMQKMKKQKIQQILD 105
>At3g32000 unknown protein
Length = 839
Score = 30.4 bits (67), Expect = 1.4
Identities = 17/47 (36%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Query: 24 FYGNYNEPSSPGEAPQAEEG--EEDDEINDLFKMGKKKNERSPAEIA 68
F GNY +P PG APQ +G D E+ + + + S EIA
Sbjct: 344 FQGNYQQPPPPGFAPQQNQGPATPDAEMKQMVQQLLQGQASSSMEIA 390
>At4g33690 hypothetical protein
Length = 281
Score = 30.0 bits (66), Expect = 1.8
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 2/66 (3%)
Query: 255 RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARA--KQASHDQQRMKMNKKLQQLR 312
R+ + R+ +PE +P+ S D EEV RA K+ H ++ K +K ++ R
Sbjct: 212 RTSDTGDRKLVIQPERSPLLRRRTDSSSSDEEEVYKRAHRKRKEHKKKLSKKHKSKEKKR 271
Query: 313 APKKRQ 318
KKR+
Sbjct: 272 DRKKRK 277
>At3g25840 protein kinase like protein
Length = 935
Score = 30.0 bits (66), Expect = 1.8
Identities = 36/172 (20%), Positives = 64/172 (36%), Gaps = 33/172 (19%)
Query: 136 DGSLPNINIRTAILKILNDFPI------DLEQIDRREQLKRSGLGKVIMFLSKSDEEINV 189
+ +P+ T IL + PI + E ++ E L+ G+G V+M + SD E
Sbjct: 52 ENEIPSAGDETEILDVTPAAPIVVSNGCEEEDVEEGEILEEDGIGDVLMKTADSDGE--- 108
Query: 190 NRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDL 249
+ +FED + P + + G+ +R+S+ +
Sbjct: 109 -----------------SGEIKFED-----NNLPPLGEKGRQGEEKSSNGVLTRESERED 146
Query: 250 DLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQR 301
+G S R + F P + E RAR++ SHD++R
Sbjct: 147 KRWDKEAGGPSERVSKLSYDNGRSSF--SPSNSRQSNEGRARSRSKSHDRER 196
>At1g56660 hypothetical protein
Length = 522
Score = 30.0 bits (66), Expect = 1.8
Identities = 28/87 (32%), Positives = 41/87 (46%), Gaps = 16/87 (18%)
Query: 40 AEEGEEDDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTAEEDAE------------ 87
A E E DDE D K GKKK + A+ V + V E E ++D E
Sbjct: 309 ATEQEMDDEAAD-HKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDDEGETKQKKNKKKE 367
Query: 88 -LNRQGKPAI--NKLKKLPLLTEVLSK 111
+ +G+ + +K K+ PL TEV+S+
Sbjct: 368 KKSEKGEKDVKEDKKKENPLETEVMSR 394
>At1g19515 unknown protein
Length = 372
Score = 30.0 bits (66), Expect = 1.8
Identities = 17/73 (23%), Positives = 36/73 (49%)
Query: 148 ILKILNDFPIDLEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFN 207
I ++ N D+E+ ++E++K+ +M + + EI + L +EL KW+R +
Sbjct: 166 IWEVSNTVLKDMEKERKKEKMKQYVQSPEVMEMCRFAGEIGIRGDLLRELRFKWAREKMD 225
Query: 208 KSTRFEDMRNIED 220
+ +E + D
Sbjct: 226 DAEFYESLEQQRD 238
>At5g22010 replication factor C large subunit-like protein
Length = 956
Score = 29.6 bits (65), Expect = 2.4
Identities = 32/99 (32%), Positives = 41/99 (41%), Gaps = 8/99 (8%)
Query: 158 DLEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRN 217
DLE DRR+ K G K + K E I RKL E D +P K T+ D +
Sbjct: 47 DLETADRRKTSKYFGKDKTKVKDEKEVEAIPAKRKLKTE-SDDLVKPRPRKVTKVVD-DD 104
Query: 218 IEDERAPFRR------PSVKKPANKAPGMQSRDSDLDLD 250
+D P R PS K + G+ S+ D D D
Sbjct: 105 DDDFDVPISRKTRDTTPSKKLKSGSGRGIASKTVDNDDD 143
>At5g13130 putative protein
Length = 706
Score = 29.6 bits (65), Expect = 2.4
Identities = 38/122 (31%), Positives = 51/122 (41%), Gaps = 25/122 (20%)
Query: 30 EPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIALLVE----NVVAELEVTAEED 85
EPS + PQ E E +IN ++G K I+ E N+ AEL+ +E
Sbjct: 547 EPSGRNQIPQVETRERSFDINP--EIGAKNRSYYGLGISSFKETGSVNLEAELQKVKQES 604
Query: 86 A----ELNRQ--------GKPAINKLKKLPLLTEVL------SKKQLQ-LEFLDHGVLTL 126
A EL RQ K I L+K EVL SK ++Q LE GV T+
Sbjct: 605 AKLVSELQRQKQLLELQESKAKIQNLEKAQREKEVLELQLKESKARIQNLENRQEGVSTI 664
Query: 127 LK 128
+
Sbjct: 665 FQ 666
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.133 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,697,652
Number of Sequences: 26719
Number of extensions: 353210
Number of successful extensions: 1289
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 1248
Number of HSP's gapped (non-prelim): 71
length of query: 334
length of database: 11,318,596
effective HSP length: 100
effective length of query: 234
effective length of database: 8,646,696
effective search space: 2023326864
effective search space used: 2023326864
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0230.8


