

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.5
(471 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
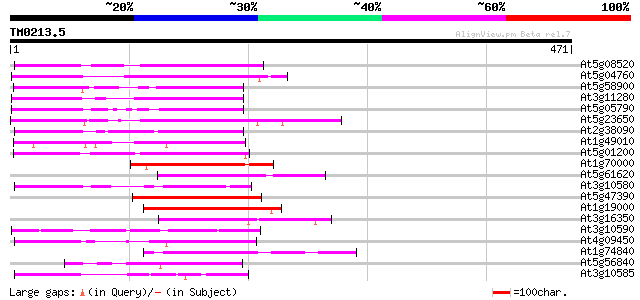
Score E
Sequences producing significant alignments: (bits) Value
At5g08520 unknown protein 134 1e-31
At5g04760 I-box binding factor-like protein 133 2e-31
At5g58900 I-box binding factor - like protein 124 1e-28
At3g11280 unknown protein 122 3e-28
At5g05790 unknown protein 122 6e-28
At5g23650 putative protein 115 4e-26
At2g38090 MYB family transcription factor like protein 113 2e-25
At1g49010 hypothetical protein 112 3e-25
At5g01200 putative protein 112 4e-25
At1g70000 unknown protein 100 1e-21
At5g61620 transcriptional activator - like protein 100 2e-21
At3g10580 hypothetical protein 99 7e-21
At5g47390 Myb-related transcription activator-like 97 2e-20
At1g19000 unknown protein 97 2e-20
At3g16350 putative MYB family transcription factor 96 3e-20
At3g10590 hypothetical protein 95 9e-20
At4g09450 unknown protein (At4g09450) 94 1e-19
At1g74840 MYB family transcription factor like protein 91 1e-18
At5g56840 putative protein 82 6e-16
At3g10585 putative protein 74 2e-13
>At5g08520 unknown protein
Length = 298
Score = 134 bits (337), Expect = 1e-31
Identities = 82/211 (38%), Positives = 112/211 (52%), Gaps = 23/211 (10%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
W+ D A E ALA D++ +RWE+IAA V G+ +I E Y LLV V IE G
Sbjct: 12 WSREDDIAFERALANNTDESEERWEKIAADVPGKSVEQIKEHYELLVEDVTRIESGC--- 68
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
VP P P S + ++G + S + K S
Sbjct: 69 -----VPLPAYGSPEGSNGHAGDEGASSKKGGN-------------SHAGESNQAGKSKS 110
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
+ K + WT+DEHR FL GL KYGKG W++ISR F+ ++TPTQ+ASHAQKY++RLNS
Sbjct: 111 DQERRKGIAWTEDEHRLFLLGLDKYGKGDWRSISRNFVVTRTPTQVASHAQKYFIRLNSM 170
Query: 184 PKRRKRASIHDLT-IDDTDLVPQQNQVPQLN 213
K R+R+SIHD+T + + D+ Q + N
Sbjct: 171 NKDRRRSSIHDITSVGNADVSTPQGPITGQN 201
>At5g04760 I-box binding factor-like protein
Length = 215
Score = 133 bits (334), Expect = 2e-31
Identities = 79/235 (33%), Positives = 118/235 (49%), Gaps = 38/235 (16%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPAEIMERYRLLVLHVDNIEEGPA 61
S WT ++D+ E AL P+ +P+RWE+IA + + E+ E Y +LV V I+ G
Sbjct: 3 SSQWTRSEDKMFEQALVLFPEGSPNRWERIADQLHKSAGEVREHYEVLVHDVFEIDSGRV 62
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
PD++ D D + K
Sbjct: 63 D---------------------------------VPDYMDDSAAAAAGWDSAGQISFGSK 89
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
++ ++ WT++EH+ FL GLK+YGKG W++ISR + ++TPTQ+ASHAQKY+LR N
Sbjct: 90 HGESERKRGTPWTENEHKLFLIGLKRYGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQN 149
Query: 182 STPKRRKRASIHDL-TIDDTDLVPQQNQ--VPQLNAPMQELSPQQNGVQLDAPMQ 233
S K RKR+SIHD+ T+D T +P N Q +P+Q +PQQ + + Q
Sbjct: 150 SVKKERKRSSIHDITTVDATLAMPGSNMDWTGQHGSPVQ--APQQQQIMSEFGQQ 202
>At5g58900 I-box binding factor - like protein
Length = 288
Score = 124 bits (311), Expect = 1e-28
Identities = 75/197 (38%), Positives = 111/197 (56%), Gaps = 27/197 (13%)
Query: 4 SWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG--P 60
+WT A+++A E ALA D+ PDRW+++AA + G+ ++++ +Y L V +IE G P
Sbjct: 33 TWTAAENKAFENALAVYDDNTPDRWQKVAAVIPGKTVSDVIRQYNDLEADVSSIEAGLIP 92
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P +I P D + P + V N S + +
Sbjct: 93 VPGYITS---PPFTLDWAGGGGGCNGFKPGHQ---------------VCNKRSQAGR--- 131
Query: 121 KISPN-QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLR 179
SP + +K V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R
Sbjct: 132 --SPELERKKGVPWTEEEHKLFLMGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIR 189
Query: 180 LNSTPKRRKRASIHDLT 196
S K ++RASIHD+T
Sbjct: 190 QLSGGKDKRRASIHDIT 206
>At3g11280 unknown protein
Length = 263
Score = 122 bits (307), Expect = 3e-28
Identities = 73/196 (37%), Positives = 102/196 (51%), Gaps = 34/196 (17%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SWT +++ E ALA +D+PDRW ++A+ + G+ ++M++Y L V +IE G
Sbjct: 30 SGSWTKEENKMFERALAIYAEDSPDRWFKVASMIPGKTVFDVMKQYSKLEEDVFDIEAGR 89
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P+P P++S P D +
Sbjct: 90 V----------PIPGYPAASS-----------------------PLGFDTDMCRKRPSGA 116
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ S +K V WT++EHR FL GL KYGKG W+ ISR F+ SKTPTQ+ASHAQKYY R
Sbjct: 117 RGSDQDRKKGVPWTEEEHRRFLLGLLKYGKGDWRNISRNFVVSKTPTQVASHAQKYYQRQ 176
Query: 181 NSTPKRRKRASIHDLT 196
S K ++R SIHD+T
Sbjct: 177 LSGAKDKRRPSIHDIT 192
>At5g05790 unknown protein
Length = 277
Score = 122 bits (305), Expect = 6e-28
Identities = 77/196 (39%), Positives = 104/196 (52%), Gaps = 28/196 (14%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SWT +++ E ALA DD PDRW ++AA + G+ +++M +Y L + +IE G
Sbjct: 28 SSSWTKEENKKFERALAVYADDTPDRWFKVAAMIPGKTISDVMRQYSKLEEDLFDIEAGL 87
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P +P P ++ +SP DF R +PN +
Sbjct: 88 VP------IPGYRSVTPCGFDQV---VSPR-------DFDAYR---KLPNGARGFDQ--- 125
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
K V WT++EHR FL GL KYGKG W+ ISR F+ SKTPTQ+ASHAQKYY R
Sbjct: 126 -----DRRKGVPWTEEEHRRFLLGLLKYGKGDWRNISRNFVGSKTPTQVASHAQKYYQRQ 180
Query: 181 NSTPKRRKRASIHDLT 196
S K ++R SIHD+T
Sbjct: 181 LSGAKDKRRPSIHDIT 196
>At5g23650 putative protein
Length = 337
Score = 115 bits (289), Expect = 4e-26
Identities = 92/290 (31%), Positives = 128/290 (43%), Gaps = 35/290 (12%)
Query: 1 MSDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
+ SW+ D A E ALA D RW++IA V G+ +++E Y +L V IE G
Sbjct: 9 VGSSWSKDDDIAFEKALAIYNDKTEIRWKKIATVVPGKTLEQVIEHYNILARDVMLIESG 68
Query: 60 PA--PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQK 117
PD+ D + EP N + I +E G ND K
Sbjct: 69 CVRLPDYD-DFLEEPNHNAFGKERSI-------LEGG---------------NDRKYESK 105
Query: 118 ITKKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKY 176
K Q +R V W EHR FL GLKKYGKG W++ISR + ++T TQ+ASHAQKY
Sbjct: 106 HKGKSKLKQKRRRGVPWKPFEHRQFLHGLKKYGKGDWRSISRHCVVTRTSTQVASHAQKY 165
Query: 177 YLRLNSTPKRRKRASIHDLTIDDTDLVPQQ------NQVPQLNAPMQELSPQQNGVQ--L 228
+ +NS K+RKR SIHD+TI + + + ++ A Q +Q L
Sbjct: 166 FAHINSEDKKRKRPSIHDITIAENKSISTKQRPITWQKINNNGATASNTQANQTTLQPSL 225
Query: 229 DAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPL 278
D P+ + A Q D P+ + QV Q + P+
Sbjct: 226 DIPIYGTHNNIWNAQATQVISQPSQNHPTYDAPIIWNTQVASQPSANIPM 275
>At2g38090 MYB family transcription factor like protein
Length = 298
Score = 113 bits (283), Expect = 2e-25
Identities = 69/193 (35%), Positives = 104/193 (53%), Gaps = 16/193 (8%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
WT +++ E ALA D PDRW ++AA + G+ +++++YR L V +IE G P
Sbjct: 29 WTAEENKKFENALAFYDKDTPDRWSRVAAMLPGKTVGDVIKQYRELEEDVSDIEAGLIP- 87
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
+P S S T F ++ S T +
Sbjct: 88 ---------IPGYASDS--FTLDWGGYDGASGNNGFNMNGYYFSAAGGKRGSAARTAE-- 134
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
++ +K V WT++EHR FL GLKKYGKG W+ I+R F+ ++TPTQ+ASHAQKY++R +
Sbjct: 135 -HERKKGVPWTEEEHRQFLMGLKKYGKGDWRNIARNFVTTRTPTQVASHAQKYFIRQVNG 193
Query: 184 PKRRKRASIHDLT 196
K ++R+SIHD+T
Sbjct: 194 GKDKRRSSIHDIT 206
>At1g49010 hypothetical protein
Length = 441
Score = 112 bits (281), Expect = 3e-25
Identities = 74/213 (34%), Positives = 111/213 (51%), Gaps = 36/213 (16%)
Query: 4 SWTPAQDRALELALA---TVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
+W+ +++A E A+A + D+W ++++ V + E+ + Y++L+ V IE G
Sbjct: 7 TWSREEEKAFENAIALHCVEEEITEDQWNKMSSMVPSKALEEVKKHYQILLEDVKAIENG 66
Query: 60 PAP--------DFIVDRVP----EPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEP 107
P IVD P D SS KK P P
Sbjct: 67 QVPLPRYHHRKGLIVDEAAAAATSPANRDSHSSGSSEKK------------------PNP 108
Query: 108 VPNDPSSSQKITKKISPNQNEKR--VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKT 165
+ SSS S + E+R + WT++EHR FL GL K+GKG W++ISR F+ S+T
Sbjct: 109 GTSGISSSNGGRSGGSRAEQERRKGIPWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRT 168
Query: 166 PTQIASHAQKYYLRLNSTPKRRKRASIHDLTID 198
PTQ+ASHAQKY++RLNS + R+R+SIHD+T +
Sbjct: 169 PTQVASHAQKYFIRLNSMNRDRRRSSIHDITTE 201
>At5g01200 putative protein
Length = 267
Score = 112 bits (280), Expect = 4e-25
Identities = 70/203 (34%), Positives = 110/203 (53%), Gaps = 17/203 (8%)
Query: 4 SWTPAQDRALELALATVPD-DAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPA 61
+WT +++ E ALA + D D + W +IA + G+ A++++RY+ L V +IE G
Sbjct: 29 TWTAEENKRFEKALAYLDDKDNLESWSKIADLIPGKTVADVIKRYKELEDDVSDIEAG-- 86
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
+P P +SS + D++V + P+ +
Sbjct: 87 ------LIPIPGYGGDASSAANSDYFFGLENSSYGYDYVVGGKR----SSPAMTDCFRSP 136
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
+ + +K V WT+DEH FL GLKKYGKG W+ I++ F+ ++TPTQ+ASHAQKY+LR
Sbjct: 137 MPEKERKKGVPWTEDEHLRFLMGLKKYGKGDWRNIAKSFVTTRTPTQVASHAQKYFLRQL 196
Query: 182 STPKRRKRASIHDLT---IDDTD 201
+ K ++R+SIHD+T I D D
Sbjct: 197 TDGKDKRRSSIHDITTVNIPDAD 219
>At1g70000 unknown protein
Length = 261
Score = 100 bits (250), Expect = 1e-21
Identities = 51/123 (41%), Positives = 75/123 (60%), Gaps = 5/123 (4%)
Query: 102 DRVPEPVPNDPS--SSQKITKKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISR 158
D+ P+P P D +S + N+ KR WT++EHR FL GL K GKG W+ ISR
Sbjct: 66 DQTPDPNPTDDGGYASDDVVHASGRNRERKRGTPWTEEEHRLFLTGLHKVGKGDWRGISR 125
Query: 159 EFLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQE 218
F+ ++TPTQ+ASHAQKY+LR + +RR+R+S+ D+T D + + QL P++
Sbjct: 126 NFVKTRTPTQVASHAQKYFLRRTNQNRRRRRSSLFDITPD--SFIGSSKEENQLQTPLEL 183
Query: 219 LSP 221
+ P
Sbjct: 184 IRP 186
>At5g61620 transcriptional activator - like protein
Length = 317
Score = 100 bits (248), Expect = 2e-21
Identities = 52/142 (36%), Positives = 83/142 (57%), Gaps = 6/142 (4%)
Query: 125 NQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTP 184
++ +K WT++EHRNFL GL K GKG W+ I++ F+ ++TPTQ+ASHAQKY++RLN
Sbjct: 102 HEKKKGKPWTEEEHRNFLIGLNKLGKGDWRGIAKSFVSTRTPTQVASHAQKYFIRLNVND 161
Query: 185 KRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDA 244
KR++RAS+ D++++D + +Q P P+Q + P+ Q +Q +
Sbjct: 162 KRKRRASLFDISLEDQKEKERNSQDASTKTP-----PKQPITGIQQPVVQGHTQTEISNR 216
Query: 245 L-DFPMQQLSFQQNGDPPLNFP 265
+ M+ + Q P NFP
Sbjct: 217 FQNLSMEYMPIYQPIPPYYNFP 238
>At3g10580 hypothetical protein
Length = 287
Score = 98.6 bits (244), Expect = 7e-21
Identities = 71/200 (35%), Positives = 98/200 (48%), Gaps = 43/200 (21%)
Query: 5 WTPAQDRALELALATVP-DDAPDRWEQIAAAVGRLPAEIMERYRLLVLHVDNIEEGPAPD 63
W D+ ELAL P + +PD E IA + + E+ Y+ LV V IE G P
Sbjct: 8 WKRDDDKRFELALVRFPAEGSPDFLENIAQFLQKPLKEVYSYYQALVDDVTLIESGKYP- 66
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
+P+ +D S + TK S +Q KK
Sbjct: 67 -----LPKYPEDDYVSLPEATK---------------------------SKTQGTGKK-- 92
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
K + W+ +EHR FL GL KYGKG W++ISRE + S++P Q+ASHAQKY+LR +
Sbjct: 93 -----KGIPWSPEEHRLFLDGLNKYGKGDWKSISRECVTSRSPMQVASHAQKYFLRQKN- 146
Query: 184 PKRRKRASIHDLTIDDTDLV 203
K+ KR SIHD+T+ D + V
Sbjct: 147 -KKGKRFSIHDMTLGDAENV 165
>At5g47390 Myb-related transcription activator-like
Length = 365
Score = 97.1 bits (240), Expect = 2e-20
Identities = 44/109 (40%), Positives = 72/109 (65%), Gaps = 1/109 (0%)
Query: 104 VPEPVPNDPSSSQK-ITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLP 162
VP+ V D +S+ + S + +K WT++EHR FL GL+K GKG W+ ISR ++
Sbjct: 68 VPDHVAGDGYASEDFVAGSSSSRERKKGTPWTEEEHRMFLLGLQKLGKGDWRGISRNYVT 127
Query: 163 SKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQ 211
++TPTQ+ASHAQKY++R ++ +R++R+S+ D+ D+ +P Q P+
Sbjct: 128 TRTPTQVASHAQKYFIRQSNVSRRKRRSSLFDMVPDEVGDIPMDLQEPE 176
>At1g19000 unknown protein
Length = 285
Score = 97.1 bits (240), Expect = 2e-20
Identities = 47/118 (39%), Positives = 75/118 (62%), Gaps = 2/118 (1%)
Query: 113 SSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASH 172
S++ + S ++ V WT++EH+ FL GL+K GKG W+ ISR F+ S+TPTQ+ASH
Sbjct: 84 SANDAVQISSSSGGRKRGVPWTENEHKRFLIGLQKVGKGDWKGISRNFVKSRTPTQVASH 143
Query: 173 AQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQE--LSPQQNGVQL 228
AQKY+LR + +RR+R+S+ D+T + + + Q N+P+ E +S Q +Q+
Sbjct: 144 AQKYFLRRTNLNRRRRRSSLFDITTETVTEMAMEQDPTQENSPLPETNISSGQQAMQV 201
>At3g16350 putative MYB family transcription factor
Length = 387
Score = 96.3 bits (238), Expect = 3e-20
Identities = 53/151 (35%), Positives = 85/151 (56%), Gaps = 7/151 (4%)
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ ++ V WT++EHR FL GL+K GKG W+ ISR ++ S+TPTQ+ASHAQKY++R S+ +
Sbjct: 132 ERKRGVPWTEEEHRLFLVGLQKLGKGDWRGISRNYVTSRTPTQVASHAQKYFIRHTSSSR 191
Query: 186 RRKRASIHDLTIDD--TDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVD 243
R++R+S+ D+ D+ TD P Q + LN P++ + ++ + +
Sbjct: 192 RKRRSSLFDMVTDEMVTDSSPTQEE-QTLNGSSPSKEPEKKSYLPSLELSLNNTTEAEEV 250
Query: 244 ALDFPMQQLSFQ----QNGDPPLNFPHQVFP 270
P Q+ S + NG P+ P FP
Sbjct: 251 VATAPRQEKSQEAIEPSNGVSPMLVPGGFFP 281
>At3g10590 hypothetical protein
Length = 206
Score = 94.7 bits (234), Expect = 9e-20
Identities = 75/211 (35%), Positives = 105/211 (49%), Gaps = 26/211 (12%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPAEIMERYRLLVLHVDNIEEGPA 61
S WT +R + AL+ P D R +A + + E+ Y LV V
Sbjct: 4 SPRWTEDDNRRFKSALSQFPPDNK-RLVNVAQHLPKPLEEVKYYYEKLVNDV-------- 54
Query: 62 PDFIVDRVPEPVPNDPSSSQK-ITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
+P+P+ N QK + + + E A D V+++PE VP SS K K
Sbjct: 55 ------YLPKPLENVTQHLQKPMEMEEMKYMYEKMAND--VNQMPEYVPLAESSQSKRRK 106
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
K +PN WT++EHR FL+GLKKYG+G S F+ +KTP Q++SHAQ YY R
Sbjct: 107 KDTPNP------WTEEEHRLFLQGLKKYGEGASTLTSTNFVKTKTPRQVSSHAQ-YYKRQ 159
Query: 181 NSTPKRRKRASIHDLTIDDTDLVPQQ-NQVP 210
S K+ KR SI D+T++ T+ P NQ P
Sbjct: 160 KSDNKKEKRRSIFDITLESTEGNPDSGNQNP 190
>At4g09450 unknown protein (At4g09450)
Length = 200
Score = 94.4 bits (233), Expect = 1e-19
Identities = 63/205 (30%), Positives = 94/205 (45%), Gaps = 46/205 (22%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPAEIMERYRLLVLHVDNIEEGPAPDF 64
WT D+ ELAL +P+ +P+ E IA + + E+ Y LV ++ IE G +
Sbjct: 7 WTRVDDKRFELALLQIPEGSPNFIENIAYYLQKPVKEVEYYYCALVHDIERIESGK---Y 63
Query: 65 IVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISP 124
++ + PE D++ K+T+
Sbjct: 64 VLPKYPED-------------------------DYV----------------KLTEAGES 82
Query: 125 NQNEKR--VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
N K+ + W+++E R FL GL K+GKG W+ ISR + S+T TQ+ASHAQKY+ R
Sbjct: 83 KGNGKKTGIPWSEEEQRLFLEGLNKFGKGDWKNISRYCVKSRTSTQVASHAQKYFARQKQ 142
Query: 183 TPKRRKRASIHDLTIDDTDLVPQQN 207
KR SIHD+T+ VP N
Sbjct: 143 ESTNTKRPSIHDMTLGVAVNVPGSN 167
>At1g74840 MYB family transcription factor like protein
Length = 265
Score = 90.9 bits (224), Expect = 1e-18
Identities = 57/179 (31%), Positives = 97/179 (53%), Gaps = 27/179 (15%)
Query: 113 SSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASH 172
SS+ KI +K + V WT++EH+ FL GL++ GKG W+ ISR F+ ++T TQ+ASH
Sbjct: 85 SSNGKIERK-------RGVPWTEEEHKLFLLGLQRVGKGDWKGISRNFVKTRTSTQVASH 137
Query: 173 AQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPM 232
AQKY+LR ++ +RR+R+S+ D+T D + + +QV Q+N Q +P+
Sbjct: 138 AQKYFLRRSNLNRRRRRSSLFDMTTDTVIPMEEDHQV----------LIQENTSQSSSPV 187
Query: 233 QQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDP 291
++++ FP + P+ +Q + Q + +N+ L+FQ + P
Sbjct: 188 PEINNFSIHPVMQVFP----------EFPVPTGNQSYGQLTSSNLINLVPLTFQSSPAP 236
>At5g56840 putative protein
Length = 233
Score = 82.0 bits (201), Expect = 6e-16
Identities = 53/152 (34%), Positives = 77/152 (49%), Gaps = 25/152 (16%)
Query: 47 RLLVLHVDNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPE 106
RL +H+D P P P PS KK F +D +P
Sbjct: 27 RLFGVHLDTTSSSPPP-----------PPPPSILAAAIKK-----------SFSMDCLPA 64
Query: 107 PVPNDPSSSQKITKKISPN--QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSK 164
+ S + ++ ++ +K V WT +EHR FL GL+K GKG W+ ISR F+ +K
Sbjct: 65 CSSSSSSFAGYLSDGLAHKTPDRKKGVPWTAEEHRTFLIGLEKLGKGDWRGISRNFVVTK 124
Query: 165 TPTQIASHAQKYYLRLNST-PKRRKRASIHDL 195
+PTQ+ASHAQKY+LR +T +R+R S+ D+
Sbjct: 125 SPTQVASHAQKYFLRQTTTLHHKRRRTSLFDM 156
>At3g10585 putative protein
Length = 170
Score = 73.9 bits (180), Expect = 2e-13
Identities = 62/199 (31%), Positives = 89/199 (44%), Gaps = 46/199 (23%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPAEIMERY-RLLVLHVDNIEEGPAPD 63
WT D+ ELAL P+ +P E IA + + E+ Y +LV V IE G
Sbjct: 7 WTRDNDKRFELALVIFPEGSPYFLEYIAEFLQKPLEEVKYYYDAILVYDVVLIESGK--- 63
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
+ + + PE + + + +K
Sbjct: 64 -----------------------------------YALPKYPEAYYVSLTEATE-SKHGE 87
Query: 124 PNQNEKRVLWTDDEHRNFLRGLK--KYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
NQ + + WT++EHRN L G+ +YGKG W ISREF+ T TQ+ASHAQKY R
Sbjct: 88 TNQIPRIIPWTEEEHRN-LAGVDALRYGKGAWSMISREFV---TSTQVASHAQKYDKRQK 143
Query: 182 STPKRRKRASIHDLTIDDT 200
K+RKR S+ D+T++ T
Sbjct: 144 LDSKKRKRWSVLDITLEST 162
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,573,951
Number of Sequences: 26719
Number of extensions: 555779
Number of successful extensions: 1714
Number of sequences better than 10.0: 146
Number of HSP's better than 10.0 without gapping: 68
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 1476
Number of HSP's gapped (non-prelim): 246
length of query: 471
length of database: 11,318,596
effective HSP length: 103
effective length of query: 368
effective length of database: 8,566,539
effective search space: 3152486352
effective search space used: 3152486352
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0213.5


