

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.14
(75 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
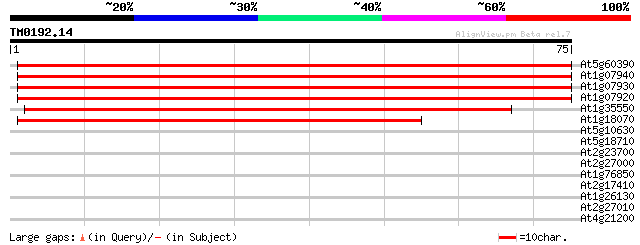
Score E
Sequences producing significant alignments: (bits) Value
At5g60390 translation elongation factor eEF-1 alpha chain (gene A4) 132 4e-32
At1g07940 elongation factor 1-alpha 132 4e-32
At1g07930 elongation factor 1-alpha 132 4e-32
At1g07920 elongation factor 1-alpha 132 4e-32
At1g35550 elongation factor, putative 101 6e-23
At1g18070 unknown protein 42 7e-05
At5g10630 putative protein 33 0.034
At5g18710 putative protein 28 1.1
At2g23700 unknown protein 27 2.4
At2g27000 putative cytochrome P450 26 3.1
At1g76850 unknown protein 26 3.1
At2g17410 unknown protein 26 4.1
At1g26130 P-type transporting ATPase like protein 25 5.4
At2g27010 putative cytochrome P450 25 7.0
At4g21200 gibberellin 20-oxidase - like protein 25 9.1
>At5g60390 translation elongation factor eEF-1 alpha chain (gene A4)
Length = 449
Score = 132 bits (331), Expect = 4e-32
Identities = 65/74 (87%), Positives = 69/74 (92%)
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSV+KK
Sbjct: 373 KEIEKEPKFLKNGDAGMVKMTPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVDKK 432
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA+KK
Sbjct: 433 DPTGAKVTKAAVKK 446
>At1g07940 elongation factor 1-alpha
Length = 449
Score = 132 bits (331), Expect = 4e-32
Identities = 65/74 (87%), Positives = 69/74 (92%)
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSV+KK
Sbjct: 373 KEIEKEPKFLKNGDAGMVKMTPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVDKK 432
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA+KK
Sbjct: 433 DPTGAKVTKAAVKK 446
>At1g07930 elongation factor 1-alpha
Length = 449
Score = 132 bits (331), Expect = 4e-32
Identities = 65/74 (87%), Positives = 69/74 (92%)
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSV+KK
Sbjct: 373 KEIEKEPKFLKNGDAGMVKMTPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVDKK 432
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA+KK
Sbjct: 433 DPTGAKVTKAAVKK 446
>At1g07920 elongation factor 1-alpha
Length = 449
Score = 132 bits (331), Expect = 4e-32
Identities = 65/74 (87%), Positives = 69/74 (92%)
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSV+KK
Sbjct: 373 KEIEKEPKFLKNGDAGMVKMTPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVDKK 432
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA+KK
Sbjct: 433 DPTGAKVTKAAVKK 446
>At1g35550 elongation factor, putative
Length = 104
Score = 101 bits (252), Expect = 6e-23
Identities = 49/65 (75%), Positives = 54/65 (82%)
Query: 3 EIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKE 62
EIEKEPKFLKN +A + M PTKPMVVE +S YP LGRFA+RDMRQTV GVIKSV KK+
Sbjct: 40 EIEKEPKFLKNSEAAIINMTPTKPMVVEAYSAYPPLGRFAIRDMRQTVGVGVIKSVVKKD 99
Query: 63 PSGAK 67
PSGAK
Sbjct: 100 PSGAK 104
>At1g18070 unknown protein
Length = 532
Score = 41.6 bits (96), Expect = 7e-05
Identities = 19/54 (35%), Positives = 33/54 (60%)
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVI 55
K ++K+ F+KNG A ++ T + +E FS++P LGRF +R +T+ G +
Sbjct: 469 KPMKKKVLFVKNGAAVVCRIQVTNSICIEKFSDFPQLGRFTLRTEGKTIAVGKV 522
>At5g10630 putative protein
Length = 804
Score = 32.7 bits (73), Expect = 0.034
Identities = 17/54 (31%), Positives = 28/54 (51%)
Query: 5 EKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSV 58
+K P+ L + +++ P+ VETFSE LGR +R +TV G + +
Sbjct: 747 KKSPRCLTAKQSAMLEVSLQNPVCVETFSESRALGRVFLRSSGRTVAMGKVTRI 800
>At5g18710 putative protein
Length = 519
Score = 27.7 bits (60), Expect = 1.1
Identities = 15/46 (32%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Query: 29 VETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEPSGA---KVTKA 71
V+ F + P++ + V ++ TV+ GVI+ ++K+ P+G + TKA
Sbjct: 376 VQAFEQDPVV-KLHVGRLKATVIKGVIQWIDKRMPTGCGSFRATKA 420
>At2g23700 unknown protein
Length = 707
Score = 26.6 bits (57), Expect = 2.4
Identities = 14/40 (35%), Positives = 23/40 (57%)
Query: 26 PMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEPSG 65
P ++E+FS+ LG+ A+ +M Q + +K KK SG
Sbjct: 642 PKIIESFSKDSGLGQAALMEMIQECLPETMKKTIKKLNSG 681
>At2g27000 putative cytochrome P450
Length = 514
Score = 26.2 bits (56), Expect = 3.1
Identities = 11/35 (31%), Positives = 21/35 (59%)
Query: 19 VKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTG 53
+++ PT P+V+ TF + +G F++ + VV G
Sbjct: 368 LRLHPTIPLVLRTFQDGCTIGGFSIPKKTKLVVNG 402
>At1g76850 unknown protein
Length = 1090
Score = 26.2 bits (56), Expect = 3.1
Identities = 14/33 (42%), Positives = 21/33 (63%), Gaps = 1/33 (3%)
Query: 43 VRDMRQTVVTGVIKSVEKKEPS-GAKVTKAALK 74
VRDMR++ V++ VE K P+ G KV +L+
Sbjct: 147 VRDMRESRTAPVVQKVEGKAPAPGKKVALTSLQ 179
>At2g17410 unknown protein
Length = 287
Score = 25.8 bits (55), Expect = 4.1
Identities = 9/23 (39%), Positives = 16/23 (69%)
Query: 29 VETFSEYPLLGRFAVRDMRQTVV 51
V+ F YPL+G+ V+DM +++
Sbjct: 45 VDIFKNYPLIGQLFVQDMYNSIM 67
>At1g26130 P-type transporting ATPase like protein
Length = 1184
Score = 25.4 bits (54), Expect = 5.4
Identities = 12/32 (37%), Positives = 19/32 (58%)
Query: 44 RDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
RDM+Q ++ +++ E SG K AALK+
Sbjct: 751 RDMKQIIINLETPEIQQLEKSGEKDAIAALKE 782
>At2g27010 putative cytochrome P450
Length = 498
Score = 25.0 bits (53), Expect = 7.0
Identities = 12/35 (34%), Positives = 19/35 (54%)
Query: 19 VKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTG 53
+++ P P+V+ TF E +G F V + VV G
Sbjct: 346 LRLHPPVPLVLRTFKEGCTIGGFYVPEKTTLVVNG 380
>At4g21200 gibberellin 20-oxidase - like protein
Length = 293
Score = 24.6 bits (52), Expect = 9.1
Identities = 13/36 (36%), Positives = 18/36 (49%)
Query: 21 MIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIK 56
M P+ V+E S+ P F+ R+ RQ V V K
Sbjct: 243 MCPSYDAVIECSSDRPAYRNFSFREFRQQVQEDVKK 278
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.133 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,573,780
Number of Sequences: 26719
Number of extensions: 50044
Number of successful extensions: 87
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 73
Number of HSP's gapped (non-prelim): 15
length of query: 75
length of database: 11,318,596
effective HSP length: 51
effective length of query: 24
effective length of database: 9,955,927
effective search space: 238942248
effective search space used: 238942248
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0192.14


