

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0181.13
(507 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
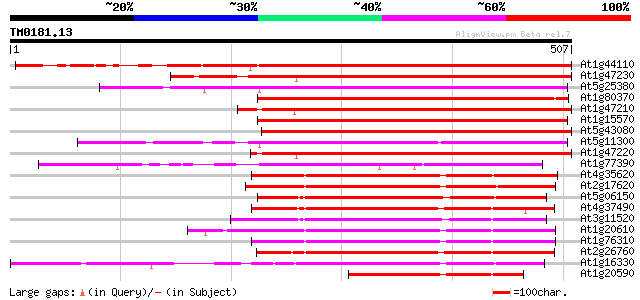
Score E
Sequences producing significant alignments: (bits) Value
At1g44110 mitotic cyclin a2-type, putative 544 e-155
At1g47230 cyclin like protein 304 9e-83
At5g25380 cyclin 3a 301 6e-82
At1g80370 putative cyclin 299 3e-81
At1g47210 cyclin like protein 295 3e-80
At1g15570 putative cyclin (At1g15570) 295 3e-80
At5g43080 cyclin A-type 294 9e-80
At5g11300 cyclin 3b 286 2e-77
At1g47220 cyclin, putative 277 9e-75
At1g77390 276 3e-74
At4g35620 cyclin 2b protein 194 1e-49
At2g17620 putative cyclin 2 189 3e-48
At5g06150 mitosis-specific cyclin 1b 187 1e-47
At4g37490 cyclin cyc1 181 1e-45
At3g11520 cyclin box 179 2e-45
At1g20610 hypothetical protein 179 4e-45
At1g76310 putative G2/mitotic-specific cyclin 1 (B-like cyclin) 177 2e-44
At2g26760 putative cyclin 176 3e-44
At1g16330 putative mitotic cyclin 158 6e-39
At1g20590 hypothetical protein 113 3e-25
>At1g44110 mitotic cyclin a2-type, putative
Length = 460
Score = 544 bits (1401), Expect = e-155
Identities = 302/505 (59%), Positives = 365/505 (71%), Gaps = 55/505 (10%)
Query: 6 HRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNLTN 65
+RRSSFSSST SSLAKR A ++ N +VK A + KKR PLSN+TN
Sbjct: 7 NRRSSFSSSTKSSLAKRQAPSSSEN------------SVKLMPAMT---KKRAPLSNITN 51
Query: 66 H-IASRNSSSQSLVPCVSKFAKTKKEAPPKVAPALPNVKSAAAAVVFPKVATATASFTER 124
IASR +S S V C +K AK K+AP++ ASF+
Sbjct: 52 QKIASRLQNSDS-VHCSNKSAKL------KIAPSV----------------CVNASFSSN 88
Query: 125 NEDDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPAEIEYVDNSDV 184
+ V + SP KSD SVSMDE+ SS DS+KSP ++EY++N DV
Sbjct: 89 LQQSI------------VPHKVASSPSKSDDGSVSMDETRSSSDSYKSP-QVEYIENDDV 135
Query: 185 SAVDSIERKTFSILNISDAKEQAGNICSRDIL--VEELEKGEKIVNIDNDHMDPQLCASF 242
SAV SIERK S L I+ E N CSRD+L +++++K + IVNID+++ DPQLCA+F
Sbjct: 136 SAVVSIERKALSNLFITPNSETIDNYCSRDVLSDMKKMDKNQ-IVNIDSNNGDPQLCATF 194
Query: 243 ACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVN 302
ACDIYKHLRASEAKKRP D+ME+VQKD+N+SMR IL+DWL+EV+EEYRLVP+TLYLTVN
Sbjct: 195 ACDIYKHLRASEAKKRPDVDYMERVQKDVNSSMRGILVDWLIEVSEEYRLVPETLYLTVN 254
Query: 303 YIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESE 362
YIDRYLSGN +SRQKLQLLGVA MMIA+KYEEICAPQVEEFCYITDNTY K+EVL MES+
Sbjct: 255 YIDRYLSGNVISRQKLQLLGVACMMIAAKYEEICAPQVEEFCYITDNTYLKDEVLDMESD 314
Query: 363 VLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSL 422
VLN+LKFEMTAPT KCFLRRFVRAA GV E P +QLE + NYIAELSL+EY+ML ++PSL
Sbjct: 315 VLNYLKFEMTAPTTKCFLRRFVRAAHGVHEAPLMQLECMANYIAELSLLEYTMLSHSPSL 374
Query: 423 VAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEK 482
VAASAIFLAK+IL P+ +PW+STLQHYT Y+ +L CVK+L RL + S LPA++EK
Sbjct: 375 VAASAIFLAKYILDPTRRPWNSTLQHYTQYKAMELRGCVKDLQRLCSTAHGSTLPAVREK 434
Query: 483 YSQHKYKYVAKKYCPPSIPSEFFQN 507
YSQHKYK+VAKK+CP IP EFF N
Sbjct: 435 YSQHKYKFVAKKFCPSVIPQEFFNN 459
>At1g47230 cyclin like protein
Length = 369
Score = 304 bits (778), Expect = 9e-83
Identities = 167/365 (45%), Positives = 241/365 (65%), Gaps = 15/365 (4%)
Query: 146 MDVSPCKSDGCSVSMDESMSSCDSFKSPAEIEYVDNSDVSAVDSIERKTFSI-LNISDAK 204
M + K S+++DE+ S K E + S+V AV + ER+T +++ +K
Sbjct: 10 MTRAAAKRKASSMALDENPVSK---KRVVLGELPNMSNVVAVPNQERETLKAKTSVNTSK 66
Query: 205 EQAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKK--RPSTD 262
Q + + L E V I++ +DPQ+C FA DI +LR E K RP D
Sbjct: 67 RQ---------MKKALMIPEASVLIESRSVDPQMCEPFASDICAYLREMEGKPKHRPLPD 117
Query: 263 FMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLG 322
++EKVQ D+ MRA+L+DWLVEVAEEY+LV DTLYLT++Y+DR+LS ++RQKLQL+G
Sbjct: 118 YIEKVQSDLTPHMRAVLVDWLVEVAEEYKLVSDTLYLTISYVDRFLSVKPINRQKLQLVG 177
Query: 323 VASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRR 382
V++M+IASKYEEI P+VE+FCYITDNT+ K+EV+ ME+++L L+FE+ +PTIK FLRR
Sbjct: 178 VSAMLIASKYEEIGPPKVEDFCYITDNTFTKQEVVSMEADILLALQFELGSPTIKTFLRR 237
Query: 383 FVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPW 442
F R AQ + LQ+E L Y++ELS+++Y+ + Y PSL++ASA+FLA+FI+ P PW
Sbjct: 238 FTRVAQEDFKDSQLQIEFLCCYLSELSMLDYTCVKYLPSLLSASAVFLARFIIRPKQHPW 297
Query: 443 SSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPS 502
+ L+ YT Y+ +DL VCV +H L+ + + L A++ KY QHKYK VA P +P
Sbjct: 298 NQMLEEYTKYKAADLQVCVGIIHDLYLSRRGNTLEAVRNKYKQHKYKCVATMPVSPELPL 357
Query: 503 EFFQN 507
FF++
Sbjct: 358 AFFED 362
>At5g25380 cyclin 3a
Length = 444
Score = 301 bits (771), Expect = 6e-82
Identities = 181/442 (40%), Positives = 256/442 (56%), Gaps = 24/442 (5%)
Query: 82 SKFAKTKKEAPPKVAPALPNVKSAAAAVVFPKVATATASFTERNEDDAAAAASGARVSVP 141
SK KKEA NV+ + V+ + + ++E A S R++
Sbjct: 6 SKHTNAKKEAISTSKIRDNNVRVTRSRAKALGVSNSPSKPAFKHETKRVARPSNKRMA-- 63
Query: 142 VSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPA------------EIEYVDNSDVSAVDS 189
S +++ C +V D + + +S S E + ++ + VD
Sbjct: 64 ---SDNITVCNQKRRAVLKDVTNTLAESIISTEGNVKVACKRGGKETKQIEEDGLVDVDG 120
Query: 190 IERKTFSILNISDAKEQAGNICSRDILVEELEKGE-------KIVNIDNDHMDPQLCASF 242
+ K L+ E S+ LV+ E+ +IV+ID+ DPQ C+ +
Sbjct: 121 EKSKLAEDLSKIRMVESLDASASKQKLVDCAEEDRSDVTDCVQIVDIDSGVQDPQFCSLY 180
Query: 243 ACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVN 302
A IY + +E ++RPST +M +VQ+DI+ +MR ILIDWLVEV+EEY+LV DTLYLTVN
Sbjct: 181 AASIYDSINVAELEQRPSTSYMVQVQRDIDPTMRGILIDWLVEVSEEYKLVSDTLYLTVN 240
Query: 303 YIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESE 362
IDR++S N + +QKLQLLG+ M+IASKYEEI AP++EEFC+ITDNTY + EVL ME +
Sbjct: 241 LIDRFMSHNYIEKQKLQLLGITCMLIASKYEEISAPRLEEFCFITDNTYTRLEVLSMEIK 300
Query: 363 VLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSL 422
VLN L F ++ PT K FLRRF+RAAQ ++VP +++E L NY AEL+L EY+ L + PSL
Sbjct: 301 VLNSLHFRLSVPTTKTFLRRFIRAAQASDKVPLIEMEYLANYFAELTLTEYTFLRFLPSL 360
Query: 423 VAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEK 482
+AASA+FLA++ L S PW+ TLQHYT Y+ S L V + L N+ S L AI K
Sbjct: 361 IAASAVFLARWTLDQSNHPWNQTLQHYTRYETSALKNTVLAMEELQLNTSGSTLIAIHTK 420
Query: 483 YSQHKYKYVAKKYCPPSIPSEF 504
Y+Q K+K VA P + + F
Sbjct: 421 YNQQKFKRVATLTSPERVNTLF 442
>At1g80370 putative cyclin
Length = 461
Score = 299 bits (765), Expect = 3e-81
Identities = 152/281 (54%), Positives = 201/281 (71%), Gaps = 1/281 (0%)
Query: 225 KIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLV 284
K V+ID+D DP LC+ +A DIY +LR +E K+RP DFMEK Q+D+ +MR IL+DWLV
Sbjct: 180 KFVDIDSDDKDPLLCSLYAPDIYYNLRVAELKRRPFPDFMEKTQRDVTETMRGILVDWLV 239
Query: 285 EVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFC 344
EV+EEY LVPDTLYLTV ID +L GN + RQ+LQLLG+ M+IASKYEEI AP++EEFC
Sbjct: 240 EVSEEYTLVPDTLYLTVYLIDWFLHGNYVERQRLQLLGITCMLIASKYEEIHAPRIEEFC 299
Query: 345 YITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNY 404
+ITDNTY +++VL+MES+VL F++ PT K FLRRF+RAAQ SL++E L NY
Sbjct: 300 FITDNTYTRDQVLEMESQVLKHFSFQIYTPTSKTFLRRFLRAAQVSFPNQSLEMEFLANY 359
Query: 405 IAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKEL 464
+ EL+LM+Y L + PS++AASA+FLAK+ L S PW+ TL+HYT Y+ SDL V L
Sbjct: 360 LTELTLMDYPFLKFLPSIIAASAVFLAKWTLNQSSHPWNPTLEHYTTYKASDLKASVHAL 419
Query: 465 HRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFF 505
L N+ +L +I+ KY Q K+K VA + +P + F
Sbjct: 420 QDLQLNTKGCSLNSIRMKYRQDKFKSVA-VFSSGELPDKLF 459
>At1g47210 cyclin like protein
Length = 372
Score = 295 bits (756), Expect = 3e-80
Identities = 147/303 (48%), Positives = 215/303 (70%), Gaps = 5/303 (1%)
Query: 207 AGNICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAK--KRPSTDFM 264
A I S + + +LE +ID+ DPQ+C + DIY++LR E K +RP D++
Sbjct: 70 AKQIKSAPVAIIDLESKS---DIDSRSDDPQMCGPYVADIYEYLRQLEVKPKQRPLPDYI 126
Query: 265 EKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVA 324
EKVQKD+ SMR +L+DWLVEVAEEY+L +TLYLTV++IDR+LS +++QKLQL+GV+
Sbjct: 127 EKVQKDVTPSMRGVLVDWLVEVAEEYKLGSETLYLTVSHIDRFLSLKTVNKQKLQLVGVS 186
Query: 325 SMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFV 384
+M+IASKYEEI P+V++FCYITDNT+ K++V++ME+++L L+FE+ PTI F+RRF
Sbjct: 187 AMLIASKYEEISPPKVDDFCYITDNTFSKQDVVKMEADILLALQFELGRPTINTFMRRFT 246
Query: 385 RAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSS 444
R AQ +VP LQLE L Y++ELS+++Y + + PSL+AASA+FLA+FI+ P PW+
Sbjct: 247 RVAQDDFKVPHLQLEPLCCYLSELSILDYKTVKFVPSLLAASAVFLARFIIRPKQHPWNQ 306
Query: 445 TLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEF 504
L+ YT Y+ +DL VCV +H L+ + L A++EKY HK++ VA P +P F
Sbjct: 307 MLEEYTKYKAADLQVCVGIIHDLYLSRRGGALQAVREKYKHHKFQCVATMPVSPELPVTF 366
Query: 505 FQN 507
+++
Sbjct: 367 WED 369
>At1g15570 putative cyclin (At1g15570)
Length = 450
Score = 295 bits (756), Expect = 3e-80
Identities = 150/280 (53%), Positives = 198/280 (70%)
Query: 225 KIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLV 284
K V+ID+D DP LC +A +I+ +LR SE K+RP DFME++QKD+ SMR IL+DWLV
Sbjct: 171 KFVDIDSDDKDPLLCCLYAPEIHYNLRVSELKRRPLPDFMERIQKDVTQSMRGILVDWLV 230
Query: 285 EVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFC 344
EV+EEY L DTLYLTV ID +L GN + RQ+LQLLG+ M+IASKYEEI AP++EEFC
Sbjct: 231 EVSEEYTLASDTLYLTVYLIDWFLHGNYVQRQQLQLLGITCMLIASKYEEISAPRIEEFC 290
Query: 345 YITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNY 404
+ITDNTY +++VL+ME++VL F++ PT K FLRRF+RAAQ PSL++E L +Y
Sbjct: 291 FITDNTYTRDQVLEMENQVLKHFSFQIYTPTPKTFLRRFLRAAQASRLSPSLEVEFLASY 350
Query: 405 IAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKEL 464
+ EL+L++Y L + PS+VAASA+FLAK+ + S PW+ TL+HYT Y+ SDL V L
Sbjct: 351 LTELTLIDYHFLKFLPSVVAASAVFLAKWTMDQSNHPWNPTLEHYTTYKASDLKASVHAL 410
Query: 465 HRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEF 504
L N+ L AI+ KY Q KYK VA P + + F
Sbjct: 411 QDLQLNTKGCPLSAIRMKYRQEKYKSVAVLTSPKLLDTLF 450
>At5g43080 cyclin A-type
Length = 355
Score = 294 bits (752), Expect = 9e-80
Identities = 141/280 (50%), Positives = 199/280 (70%)
Query: 228 NIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVA 287
+ID DPQ+C + I+++LR E K RP D++EK+QKD+ ++MR +L+DWLVEVA
Sbjct: 73 DIDTRSDDPQMCGPYVTSIFEYLRQLEVKSRPLVDYIEKIQKDVTSNMRGVLVDWLVEVA 132
Query: 288 EEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYIT 347
EEY+L+ DTLYL V+YIDR+LS +++Q+LQLLGV SM+IASKYEEI P V++FCYIT
Sbjct: 133 EEYKLLSDTLYLAVSYIDRFLSLKTVNKQRLQLLGVTSMLIASKYEEITPPNVDDFCYIT 192
Query: 348 DNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAE 407
DNTY K+E+++ME+++L L+FE+ PT FLRRF R AQ E+ LQ+E L +Y++E
Sbjct: 193 DNTYTKQEIVKMEADILLALQFELGNPTSNTFLRRFTRVAQEDFEMSHLQMEFLCSYLSE 252
Query: 408 LSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRL 467
LS+++Y + + PS VAASA+FLA+FI+ P PW+ L+ YT Y+ DL CV +H L
Sbjct: 253 LSMLDYQSVKFLPSTVAASAVFLARFIIRPKQHPWNVMLEEYTRYKAGDLKECVAMIHDL 312
Query: 468 FCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFFQN 507
+ + L AI+EKY QHK+K VA P +P F++
Sbjct: 313 YLSRKCGALEAIREKYKQHKFKCVATMPVSPELPLTVFED 352
>At5g11300 cyclin 3b
Length = 434
Score = 286 bits (732), Expect = 2e-77
Identities = 170/449 (37%), Positives = 247/449 (54%), Gaps = 32/449 (7%)
Query: 62 NLTNHIASRNSSSQSLVPCVSKFAKTKKEAPPKVAPALPNVKSAAAAVVFPKVATATASF 121
N S + +S V AK + P P+ K V V+ +A
Sbjct: 10 NANKENISTSDVQESFVRITRSRAKKAMGRGVSIPPTKPSFKQQKRRAVLKDVSNTSADI 69
Query: 122 TERNEDDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPAEIEYVDN 181
+ G + + +G + +MD + AE
Sbjct: 70 IY------SELRKGGNIKANRKCLKEPKKAAKEGANSAMDILVDMHTEKSKLAE------ 117
Query: 182 SDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGE------KIVNIDNDHMD 235
D+S + E + S+ N D + + E+ E G ++V+ID++ D
Sbjct: 118 -DLSKIRMAEAQDVSLSNFKDEE-----------ITEQQEDGSGVMELLQVVDIDSNVED 165
Query: 236 PQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPD 295
PQ C+ +A DIY ++ +E ++RP ++ME VQ+DI+ MR ILIDWLVEV+++Y+LVPD
Sbjct: 166 PQCCSLYAADIYDNIHVAELQQRPLANYMELVQRDIDPDMRKILIDWLVEVSDDYKLVPD 225
Query: 296 TLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEE 355
TLYLTVN IDR+LS + + RQ+LQLLGV+ M+IASKYEE+ AP VEEFC+IT NTY + E
Sbjct: 226 TLYLTVNLIDRFLSNSYIERQRLQLLGVSCMLIASKYEELSAPGVEEFCFITANTYTRPE 285
Query: 356 VLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSM 415
VL ME ++LNF+ F ++ PT K FL + +VP ++LE L NY+AEL+L+EYS
Sbjct: 286 VLSMEIQILNFVHFRLSVPTTKTFLSALFLII--ILQVPFIELEYLANYLAELTLVEYSF 343
Query: 416 LCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSN 475
L + PSL+AASA+FLA++ L + PW+ TLQHYT Y+ ++L V + L N+
Sbjct: 344 LRFLPSLIAASAVFLARWTLDQTDHPWNPTLQHYTRYEVAELKNTVLAMEDLQLNTSGCT 403
Query: 476 LPAIKEKYSQHKYKYVAKKYCPPSIPSEF 504
L A +EKY+Q K+K VAK P + S F
Sbjct: 404 LAATREKYNQPKFKSVAKLTSPKRVTSLF 432
>At1g47220 cyclin, putative
Length = 327
Score = 277 bits (709), Expect = 9e-75
Identities = 142/292 (48%), Positives = 203/292 (68%), Gaps = 5/292 (1%)
Query: 218 EELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKK--RPSTDFMEKVQKDINTSM 275
E ++ G +ID DPQ+C + DIY++LR E K RP D++EK+Q+DI S
Sbjct: 35 ENIQSGS---DIDARSDDPQMCGLYVSDIYEYLRELEVKPKLRPLHDYIEKIQEDITPSK 91
Query: 276 RAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEI 335
R +L+DWLVEVAEE+ LV +TLYLTV+YIDR+LS ++ LQL+GV++M IASKYEE
Sbjct: 92 RGVLVDWLVEVAEEFELVSETLYLTVSYIDRFLSLKMVNEHWLQLVGVSAMFIASKYEEK 151
Query: 336 CAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPS 395
P+VE+FCYIT NTY K++VL+ME ++L L+FE+ PT FLRRF+R AQ +VP+
Sbjct: 152 RRPKVEDFCYITANTYTKQDVLKMEEDILLALEFELGRPTTNTFLRRFIRVAQEDFKVPN 211
Query: 396 LQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPS 455
LQLE L Y++ELS+++YS + + PSL+AASA+FLA+FI+ P+ PWS L+ T Y+ +
Sbjct: 212 LQLEPLCCYLSELSMLDYSCVKFVPSLLAASAVFLARFIILPNQHPWSQMLEECTKYKAA 271
Query: 456 DLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFFQN 507
DL VCV+ + L+ + A++EKY QHK++YVA +P F+++
Sbjct: 272 DLQVCVEIMLDLYLSRSEGASKAVREKYKQHKFQYVAAIPVYQELPVTFWED 323
>At1g77390
Length = 477
Score = 276 bits (705), Expect = 3e-74
Identities = 187/473 (39%), Positives = 258/473 (54%), Gaps = 70/473 (14%)
Query: 27 NNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNLTNHI-ASRNSS-SQSLVPCVSKF 84
+++ N + N + K + +R PL ++TN SRN S S +LV C +K
Sbjct: 3 SSSRNLSQENPIPRPNLAKTRTSLRDVGNRRAPLGDITNQKNGSRNPSPSSTLVNCSNKI 62
Query: 85 AKTKKEAPPKVA---------PALPNVKSAAAAVVFPKVATATASFTERNEDDAAAAASG 135
++KK P ++ LP +A + ++ P T DD+ +S
Sbjct: 63 GQSKKAPKPALSRNWNLGILDSGLPPKPNAKSNIIVPYEDTELLQ-----SDDSLLCSSP 117
Query: 136 ARVSVPVSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPAEIEYVDNSDVSAVDSIERKTF 195
A S+D SP +SD S+S +S+++ VD + T
Sbjct: 118 A-------LSLDASPTQSDP-SISTHDSLTN------------------HVVDYMVESTT 151
Query: 196 SILNISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEA 255
N D E IVNID+D MDPQLCASFACDIY+HLR SE
Sbjct: 152 DDGNDDDDDE--------------------IVNIDSDLMDPQLCASFACDIYEHLRVSEV 191
Query: 256 KKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSR 315
KRP+ D+ME+ Q IN SMR+ILIDWLVEVAEEYRL P+TLYL VNY+DRYL+GNA+++
Sbjct: 192 NKRPALDYMERTQSSINASMRSILIDWLVEVAEEYRLSPETLYLAVNYVDRYLTGNAINK 251
Query: 316 QKLQLLGVASMMIASKY----EEICAPQVEEFCYITDNTYFKEEVLQMESEVL---NFLK 368
Q LQLLGV MMIA+ + + ++ E N+ EV +V+ +FL
Sbjct: 252 QNLQLLGVTCMMIAAFLTAFGDGVFCSELLEVRINNSNSKMFLEVSSCHIDVIFIHHFLW 311
Query: 369 FEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAI 428
F K +RRF+RAAQG +EVPSL E L Y+ ELSL++Y+ML YAPSLVAASA+
Sbjct: 312 FPADL-DFKNVVRRFLRAAQGRKEVPSLLSECLACYLTELSLLDYAMLRYAPSLVAASAV 370
Query: 429 FLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKE 481
FLA++ L PS KPW++TL+HYT Y+ + CVK L +L +S++ AI++
Sbjct: 371 FLAQYTLHPSRKPWNATLEHYTSYRAKHMEACVKNLLQLCNEKLSSDVVAIRK 423
>At4g35620 cyclin 2b protein
Length = 429
Score = 194 bits (493), Expect = 1e-49
Identities = 107/278 (38%), Positives = 172/278 (61%), Gaps = 8/278 (2%)
Query: 219 ELEKGEKIVNIDNDHMDPQLCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRA 277
E E+ E +++ID + L A + D+Y R +E D+M + Q DI+ MRA
Sbjct: 148 EEEQEEPVLDIDEYDANNSLAAVEYVQDLYDFYRKTERFSCVPLDYMAQ-QFDISDKMRA 206
Query: 278 ILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICA 337
ILIDWL+EV +++ L+ +TL+LTVN IDR+LS A++R+KLQL+G+ ++++A KYEE+
Sbjct: 207 ILIDWLIEVHDKFELMNETLFLTVNLIDRFLSKQAVARKKLQLVGLVALLLACKYEEVSV 266
Query: 338 PQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQ 397
P VE+ I+D Y + +VL+ME +L+ L+F M+ PT FL+RF++AAQ +
Sbjct: 267 PIVEDLVVISDKAYTRTDVLEMEKIMLSTLQFNMSLPTQYPFLKRFLKAAQS-----DKK 321
Query: 398 LESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDL 457
LE L +++ EL+L++Y M+ Y PSL+AA+A++ A+ + W+ST + + Y + L
Sbjct: 322 LEILASFLIELALVDYEMVRYPPSLLAATAVYTAQCTIH-GFSEWNSTCEFHCHYSENQL 380
Query: 458 CVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKY 495
C + + RL + L + KYS K+ Y+A KY
Sbjct: 381 LECCRRMVRLHQKAGTDKLTGVHRKYSSSKFGYIATKY 418
>At2g17620 putative cyclin 2
Length = 429
Score = 189 bits (480), Expect = 3e-48
Identities = 107/281 (38%), Positives = 171/281 (60%), Gaps = 8/281 (2%)
Query: 214 DILVEELEKGEKIVNIDN-DHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDIN 272
++ +E++ E IV+ID D + + D+Y R E D+M + Q D+N
Sbjct: 142 EVEMEDVTVEEPIVDIDVLDSKNSLAAVEYVQDLYAFYRTMERFSCVPVDYMMQ-QIDLN 200
Query: 273 TSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKY 332
MRAILIDWL+EV +++ L+ +TL+LTVN IDR+LS + R+KLQL+G+ ++++A KY
Sbjct: 201 EKMRAILIDWLIEVHDKFDLINETLFLTVNLIDRFLSKQNVMRKKLQLVGLVALLLACKY 260
Query: 333 EEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEE 392
EE+ P VE+ I+D Y + +VL+ME +L+ L+F ++ PT FL+RF++AAQ
Sbjct: 261 EEVSVPVVEDLVLISDKAYTRNDVLEMEKTMLSTLQFNISLPTQYPFLKRFLKAAQA--- 317
Query: 393 VPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLY 452
+ E L +++ EL+L+EY ML + PSL+AA++++ A+ L S K W+ST + + Y
Sbjct: 318 --DKKCEVLASFLIELALVEYEMLRFPPSLLAATSVYTAQCTLDGSRK-WNSTCEFHCHY 374
Query: 453 QPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAK 493
L C ++L L + NL + KYS K+ Y+AK
Sbjct: 375 SEDQLMECSRKLVSLHQRAATGNLTGVYRKYSTSKFGYIAK 415
>At5g06150 mitosis-specific cyclin 1b
Length = 445
Score = 187 bits (475), Expect = 1e-47
Identities = 106/262 (40%), Positives = 163/262 (61%), Gaps = 9/262 (3%)
Query: 225 KIVNIDNDHMDPQLCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWL 283
KI++ID D L A + D+Y + E + +P +M +Q ++N MRAILIDWL
Sbjct: 164 KIIDIDESDKDNHLAAVEYVDDMYSFYKEVEKESQPKM-YMH-IQTEMNEKMRAILIDWL 221
Query: 284 VEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEF 343
+EV ++ L +TLYLTVN IDR+LS A+ +++LQL+G+++++IASKYEEI PQV +
Sbjct: 222 LEVHIKFELNLETLYLTVNIIDRFLSVKAVPKRELQLVGISALLIASKYEEIWPPQVNDL 281
Query: 344 CYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTN 403
Y+TDN Y ++L ME +L L++ +T PT FL RF++A+ E +E++ +
Sbjct: 282 VYVTDNAYSSRQILVMEKAILGNLEWYLTVPTQYVFLVRFIKASMSDPE-----MENMVH 336
Query: 404 YIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKE 463
++AEL +M Y L + PS++AASA++ A+ L S W+ TLQ +T Y S++ C K
Sbjct: 337 FLAELGMMHYDTLTFCPSMLAASAVYTARCSLNKS-PAWTDTLQFHTGYTESEIMDCSKL 395
Query: 464 LHRLFCNSPNSNLPAIKEKYSQ 485
L L S L A+ +KYS+
Sbjct: 396 LAFLHSRCGESRLRAVYKKYSK 417
>At4g37490 cyclin cyc1
Length = 428
Score = 181 bits (458), Expect = 1e-45
Identities = 107/281 (38%), Positives = 179/281 (63%), Gaps = 15/281 (5%)
Query: 219 ELEKGEKIVNIDNDHMDPQLCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRA 277
E ++ EKIV+ID+ ++ L A + DIY ++ E++ RP D+M Q DIN MR
Sbjct: 141 EKKQKEKIVDIDSADVENDLAAVEYVEDIYSFYKSVESEWRPR-DYMAS-QPDINEKMRL 198
Query: 278 ILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICA 337
IL++WL++V + L P+T YLTVN +DR+LS + R++LQL+G++++++++KYEEI
Sbjct: 199 ILVEWLIDVHVRFELNPETFYLTVNILDRFLSVKPVPRKELQLVGLSALLMSAKYEEIWP 258
Query: 338 PQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQ 397
PQVE+ I D+ Y +++L ME +L+ L++ +T PT FL RF++A+ + +
Sbjct: 259 PQVEDLVDIADHAYSHKQILVMEKTILSTLEWYLTVPTHYVFLARFIKAS-----IADEK 313
Query: 398 LESLTNYIAELSLMEY-SMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSD 456
+E++ +Y+AEL +M Y +M+ ++PS+VAASAI+ A+ L + W+STL+H+T Y +
Sbjct: 314 MENMVHYLAELGVMHYDTMIMFSPSMVAASAIYAARSSL-RQVPIWTSTLKHHTGYSETQ 372
Query: 457 LCVCVKEL-----HRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
L C K L + S +S A+++KYS+ + VA
Sbjct: 373 LMDCAKLLAYQQWKQQEEGSESSTKGALRKKYSKDERFAVA 413
>At3g11520 cyclin box
Length = 414
Score = 179 bits (455), Expect = 2e-45
Identities = 107/287 (37%), Positives = 169/287 (58%), Gaps = 9/287 (3%)
Query: 200 ISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCA-SFACDIYKHLRASEAKKR 258
++ AKE + +L + K ++ID + L A + D+Y + + +
Sbjct: 116 VAKAKENKKKVTYSSVLDARSKAASKTLDIDYVDKENDLAAVEYVEDMYIFYKEVVNESK 175
Query: 259 PSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKL 318
P +M Q +I+ MR+ILIDWLVEV ++ L P+TLYLTVN IDR+LS + R++L
Sbjct: 176 PQM-YMH-TQPEIDEKMRSILIDWLVEVHVKFDLSPETLYLTVNIIDRFLSLKTVPRREL 233
Query: 319 QLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKC 378
QL+GV++++IASKYEEI PQV + Y+TDN+Y ++L ME +L L++ +T PT
Sbjct: 234 QLVGVSALLIASKYEEIWPPQVNDLVYVTDNSYNSRQILVMEKTILGNLEWYLTVPTQYV 293
Query: 379 FLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPS 438
FL RF++A+ + +LE+L +++AEL LM + L + PS++AASA++ A+ L
Sbjct: 294 FLVRFIKASGSDQ-----KLENLVHFLAELGLMHHDSLMFCPSMLAASAVYTARCCL-NK 347
Query: 439 IKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQ 485
W+ TL+ +T Y S L C K L + + S L + +KYS+
Sbjct: 348 TPTWTDTLKFHTGYSESQLMDCSKLLAFIHSKAGESKLRGVLKKYSK 394
>At1g20610 hypothetical protein
Length = 429
Score = 179 bits (453), Expect = 4e-45
Identities = 114/338 (33%), Positives = 185/338 (54%), Gaps = 15/338 (4%)
Query: 161 DESMSSCDSFKSPAE---IEYVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILV 217
DE DS S I VD SD DS E + ++A + ++I +
Sbjct: 94 DEETKKPDSVSSEEPETIIIDVDESDKEGGDSNEPM---FVQHTEAMLEEIEQMEKEIEM 150
Query: 218 EELEKGEK-IVNIDN-DHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSM 275
E+ +K E+ +++ID D +P + D++ + E ++M+ Q+D+N M
Sbjct: 151 EDADKEEEPVIDIDACDKNNPLAAVEYIHDMHTFYKNFEKLSCVPPNYMDN-QQDLNERM 209
Query: 276 RAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEI 335
R ILIDWL+EV ++ L+ +TLYLT+N IDR+L+ + + R+KLQL+GV ++++A KYEE+
Sbjct: 210 RGILIDWLIEVHYKFELMEETLYLTINVIDRFLAVHQIVRKKLQLVGVTALLLACKYEEV 269
Query: 336 CAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPS 395
P V++ I+D Y + EVL ME + N L+F + PT F++RF++AAQ
Sbjct: 270 SVPVVDDLILISDKAYSRREVLDMEKLMANTLQFNFSLPTPYVFMKRFLKAAQS-----D 324
Query: 396 LQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPS 455
+LE L+ ++ EL L+EY ML Y PS +AASAI+ A+ L + WS T + +T Y
Sbjct: 325 KKLEILSFFMIELCLVEYEMLEYLPSKLAASAIYTAQCTL-KGFEEWSKTCEFHTGYNEK 383
Query: 456 DLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAK 493
L C +++ + L + KY+ K+ + A+
Sbjct: 384 QLLACARKMVAFHHKAGTGKLTGVHRKYNTSKFCHAAR 421
>At1g76310 putative G2/mitotic-specific cyclin 1 (B-like cyclin)
Length = 418
Score = 177 bits (448), Expect = 2e-44
Identities = 103/277 (37%), Positives = 162/277 (58%), Gaps = 9/277 (3%)
Query: 219 ELEKGEKIVNIDN-DHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRA 277
+ E E +++ID+ D +P + DIY + +E + ++ME Q DIN MR
Sbjct: 140 DAEVEESVMDIDSCDKNNPLSVVEYINDIYCFYKKNECRSCVPPNYMEN-QHDINERMRG 198
Query: 278 ILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNA-MSRQKLQLLGVASMMIASKYEEIC 336
IL DWL+EV ++ L+ +TLYLT+N IDR+L+ + ++R+KLQL+GV +M++A KYEE+
Sbjct: 199 ILFDWLIEVHYKFELMEETLYLTINLIDRFLAVHQHIARKKLQLVGVTAMLLACKYEEVS 258
Query: 337 APQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSL 396
P V++ I+D Y + E+L ME + N L+F PT F+RRF++AAQ
Sbjct: 259 VPVVDDLILISDKAYTRTEILDMEKLMANTLQFNFCLPTPYVFMRRFLKAAQS-----DK 313
Query: 397 QLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSD 456
+LE L+ ++ EL L+EY ML Y PS +AASAI+ A+ L + WS T + ++ Y
Sbjct: 314 KLELLSFFMIELCLVEYEMLQYTPSQLAASAIYTAQSTL-KGYEDWSKTSEFHSGYTEEA 372
Query: 457 LCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAK 493
L C +++ L + L + KY+ K+ Y A+
Sbjct: 373 LLECSRKMVGLHHKAGTGKLTGVHRKYNTSKFGYAAR 409
>At2g26760 putative cyclin
Length = 387
Score = 176 bits (446), Expect = 3e-44
Identities = 108/272 (39%), Positives = 171/272 (62%), Gaps = 11/272 (4%)
Query: 224 EKIVNIDNDHMDPQLCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDW 282
+ +++ID + +L A + DI+K R E ++ D++ Q +IN MR+ILIDW
Sbjct: 111 DAVIDIDAVDANNELAAVEYVEDIFKFYRTVE-EEGGIKDYIGS-QPEINEKMRSILIDW 168
Query: 283 LVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEE 342
LV+V ++ L+P+TLYLT+N +DR+LS + R++LQLLG+ +M+IA KYEEI AP+V +
Sbjct: 169 LVDVHRKFELMPETLYLTINLVDRFLSLTMVHRRELQLLGLGAMLIACKYEEIWAPEVND 228
Query: 343 FCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVP-SLQLESL 401
F I+DN Y +++VL ME +L +++ +T PT FL R+V+AA VP ++E L
Sbjct: 229 FVCISDNAYNRKQVLAMEKSILGQVEWYITVPTPYVFLARYVKAA-----VPCDAEMEKL 283
Query: 402 TNYIAELSLMEYSMLCY-APSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVC 460
Y+AEL LM+Y ++ PS++AASA++ A+ IL W+ TL+H+T Y ++
Sbjct: 284 VFYLAELGLMQYPIVVLNRPSMLAASAVYAARQIL-KKTPFWTETLKHHTGYSEDEIMEH 342
Query: 461 VKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
K L +L ++ S L A+ +KYS + VA
Sbjct: 343 AKMLMKLRDSASESKLIAVFKKYSVSENAEVA 374
>At1g16330 putative mitotic cyclin
Length = 518
Score = 158 bits (400), Expect = 6e-39
Identities = 127/489 (25%), Positives = 228/489 (45%), Gaps = 60/489 (12%)
Query: 1 MSSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPL 60
+ + R S +++KR NNN + A + +A +++ K R +
Sbjct: 65 LKTGEMERCKNGKSEKENVSKRTTELGNNNLYKTASKASNQAVPQLSSAGTYTWKTRTSV 124
Query: 61 SNLTNHIASRNSSSQSLVPCVSKFAKTKKEAPPKVA-PALPNVKSAAAAVVFPKVATATA 119
++ + N S++ V + KT + + P + KS + + + +T
Sbjct: 125 GSIQS---DGNKQSKNNVRFIQTTVKTSLQNRSSLKKPPVGRSKSRSISSIPSSAVASTL 181
Query: 120 SFTERNE-----DDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPA 174
S E+ E +D +S + P + +DV+
Sbjct: 182 SLPEKVETKCLEEDTQGESSSSGNKDPTTKVLDVT------------------------- 216
Query: 175 EIEYVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHM 234
+ S RK+F+ L ++ +K N E + EK+ +ID++
Sbjct: 217 ----------AKPKSKRRKSFTSLLVNGSKFDEKN--------GETTEPEKLPSIDDESN 258
Query: 235 DPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVP 294
++ A + DIY+ +EA P+ +++ R ILI+WL+EV ++ L+
Sbjct: 259 QLEV-AEYVDDIYQFYWTAEALN-PALGHYLSAHAEVSPVTRGILINWLIEVHFKFDLMH 316
Query: 295 DTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKE 354
+TLYLT++ +DRYLS + + ++QL+G+ ++++ASKYE+ P++++ I+ +Y +E
Sbjct: 317 ETLYLTMDLLDRYLSQVPIHKNEMQLIGLTALLLASKYEDYWHPRIKDLISISAESYTRE 376
Query: 355 EVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYS 414
++L ME +L LKF + APT F+ RF++AAQ + +LE L Y+ EL L+EY
Sbjct: 377 QILGMERSMLKQLKFRLNAPTPYVFMLRFLKAAQS-----NKKLEQLAFYLIELCLVEYE 431
Query: 415 MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNS 474
L Y PSL+ ASAI++A+ L + W+S L ++T Y S + C + R +
Sbjct: 432 ALKYKPSLLCASAIYVARCTLHMT-PVWTSLLNNHTHYNVSQMKDCSDMILRFHKAAKTG 490
Query: 475 NLPAIKEKY 483
NL EKY
Sbjct: 491 NLRVTYEKY 499
>At1g20590 hypothetical protein
Length = 199
Score = 113 bits (282), Expect = 3e-25
Identities = 61/158 (38%), Positives = 97/158 (60%), Gaps = 6/158 (3%)
Query: 307 YLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNF 366
+L+ + + R+KLQL+GV ++++A KYEE+ P V++ I+D Y + EVL ME + N
Sbjct: 3 FLAVHQIVRKKLQLVGVTALLLACKYEEVSVPVVDDLILISDKAYSRREVLDMEKLMANT 62
Query: 367 LKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAAS 426
L+F + PT F++RF++AAQ +LE L+ ++ EL L+EY ML Y PS +AAS
Sbjct: 63 LQFNFSLPTPYVFMKRFLKAAQS-----DKKLEILSFFMIELCLVEYEMLEYLPSKLAAS 117
Query: 427 AIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKEL 464
AI+ A+ L + WS T + +T Y L C +++
Sbjct: 118 AIYTAQCTL-KGFEEWSKTCEFHTGYNEEQLLACARKM 154
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,992,392
Number of Sequences: 26719
Number of extensions: 459339
Number of successful extensions: 2913
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 2478
Number of HSP's gapped (non-prelim): 247
length of query: 507
length of database: 11,318,596
effective HSP length: 104
effective length of query: 403
effective length of database: 8,539,820
effective search space: 3441547460
effective search space used: 3441547460
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0181.13


