

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.5
(1481 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
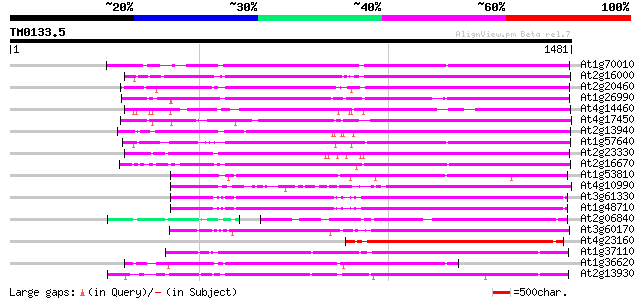
Score E
Sequences producing significant alignments: (bits) Value
At1g70010 hypothetical protein 947 0.0
At2g16000 putative retroelement pol polyprotein 943 0.0
At2g20460 putative retroelement pol polyprotein 929 0.0
At1g26990 polyprotein, putative 881 0.0
At4g14460 retrovirus-related like polyprotein 881 0.0
At4g17450 retrotransposon like protein 872 0.0
At2g13940 putative retroelement pol polyprotein 852 0.0
At1g57640 822 0.0
At2g23330 putative retroelement pol polyprotein 805 0.0
At2g16670 putative retroelement pol polyprotein 740 0.0
At1g53810 615 e-176
At4g10990 putative retrotransposon polyprotein 614 e-175
At3g61330 copia-type polyprotein 585 e-167
At1g48710 hypothetical protein 584 e-166
At2g06840 putative retroelement pol polyprotein 566 e-161
At3g60170 putative protein 559 e-159
At4g23160 putative protein 544 e-154
At1g37110 544 e-154
At1g36620 hypothetical protein 537 e-152
At2g13930 putative retroelement pol polyprotein 534 e-151
>At1g70010 hypothetical protein
Length = 1315
Score = 947 bits (2447), Expect = 0.0
Identities = 525/1245 (42%), Positives = 736/1245 (58%), Gaps = 114/1245 (9%)
Query: 257 PLIAARENSNDGRSSQSNSGYQSNSGYQSGS---GNYHSNSGRN--RYSNKK---CSYYG 308
P ++ N D +Q ++ + G S N S S N Y K+ CSY
Sbjct: 157 PSLSEVFNMIDQDETQRSARISTTPGMTSSVFPVSNQSSQSALNGDTYQKKERPVCSYCS 216
Query: 309 KMGHTVEDCYKKHGFPPGFKFKNPKYA--NRSANLVQTTKEDQESTSQGNTQEQEAARFG 366
+ GH + CYKKHG+P FK K K+ + SAN ++E +TS +
Sbjct: 217 RPGHVEDTCYKKHGYPTSFKSKQ-KFVKPSISANAAIGSEEVVNNTS--------VSTGD 267
Query: 367 FTADQYHHLLLLLPPSEQKNST----SQHTASINSCVQMLPGKNGNSISQISTKNGNPLD 422
T Q L+ L Q ST H+ S++S
Sbjct: 268 LTTSQIQQLVSFLSSKLQPPSTPVQPEVHSISVSS------------------------- 302
Query: 423 TTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLF 482
D ++ +C I GS+ L IL +VLF
Sbjct: 303 -----DPSSSSTVC---------------------------PISGSVHLGRHLILNDVLF 330
Query: 483 LPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPF 542
+P F+FNL+SV L KS+ R+ F + C++QD+ M+G + V +LYI++ DSLS
Sbjct: 331 IPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATRELMVGMGKQVANLYIVDLDSLSHP 390
Query: 543 PSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDL--FPYVQYTQDHVCEV 600
+ +S++ S + +LWH RLGHPS K Q + L FP + D C V
Sbjct: 391 GTDSSITVA---------SVTSHDLWHKRLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRV 441
Query: 601 CPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLL 660
C I+KQK L F N+ S F +IH+D WGP SV + +G+ YFLTIVDDYSR TWVYLL
Sbjct: 442 CHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYLL 501
Query: 661 KSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQN 720
++K +V T++ F V NQF ++K VRSDN E +QFY KGI+ + SC ETPQQN
Sbjct: 502 RNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELNFTQFYHSKGIVPYHSCPETPQQN 561
Query: 721 SIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQL 780
S+VERKHQHILNVAR+L FQ+H+P +W I+ +++LINRLP PIL+++CPF++L K +
Sbjct: 562 SVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRLPAPILEDKCPFEVLTKTV 621
Query: 781 PDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNV 840
P +KVFG LC+AST R KF RAK C F+G+ SG KGY + DL T I +SR+V
Sbjct: 622 PTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFKGYKLLDLETHSIIVSRHV 681
Query: 841 IFHENIFPYKLHRDDHECESSTFPQI---PCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQ 897
+FHE +FP+ L D + E + FP + P + + D+ P + V+I P S+ P+
Sbjct: 682 VFHEELFPF-LGSDLSQEEQNFFPDLNPTPPMQRQSSDHVNPSDSSSSVEILP-SANPTN 739
Query: 898 NSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPS 957
N +P V + S R K+P YLQDY+C +S T H + + +SY +++
Sbjct: 740 NVPEPSV-QTSHRKAKKPAYLQDYYC-----------HSVVSSTPHEIRKFLSYDRINDP 787
Query: 958 YHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWV 1017
Y F+ + T EP+ Y+EA K + WR AM E + LE +TW + P DK IGC+W+
Sbjct: 788 YLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWI 847
Query: 1018 YRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQ 1077
++IKY DG+++RYKARLV +GYTQ EGID+ +TFSPVAK+ ++++LL + + L Q
Sbjct: 848 FKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQ 907
Query: 1078 LDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PNQVCLLQKSLYGLKQASRQWFSTLCQ 1132
LD+ NAFL+ LDEEIYM LPQG S + PN VC L+KSLYGLKQASRQW+
Sbjct: 908 LDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSS 967
Query: 1133 ALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIK 1192
L GLGF QS+ DHT ++K S G F +L+Y+DD+++A N+ + ++K + F+++
Sbjct: 968 TLLGLGFIQSYCDHTCFLKIS-DGIFLCVLVYIDDIIIASNNDAAVDILKSQMKSFFKLR 1026
Query: 1193 DLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTP 1252
DLGE K+FLGL I RS KGI ++QRKYAL+LL ++G LG K ++ PMD S + SG
Sbjct: 1027 DLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGD 1086
Query: 1253 LSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCG 1312
++ YRRLIGRL+YL TRPDI + VN+L+QF AP H A +++L+YIKG+ G G
Sbjct: 1087 FVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQG 1146
Query: 1313 LFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEY 1372
LFY A S L ++++D C D+R+S +GYCMFLG SL+ W+S+KQ S+SS EAEY
Sbjct: 1147 LFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEY 1206
Query: 1373 RAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIV 1432
R+++ E+ WL L++LQV + P +FCDN++A+HIA+N +HERTKHIE DCH V
Sbjct: 1207 RSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSV 1266
Query: 1433 REKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDI 1477
RE++ +GL L + + LQ+AD FTKPL P+ F + SK+ L +I
Sbjct: 1267 RERLLKGLFELYHINTELQIADPFTKPLYPSHFHRLISKMGLLNI 1311
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 943 bits (2438), Expect = 0.0
Identities = 513/1192 (43%), Positives = 711/1192 (59%), Gaps = 65/1192 (5%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGF--------KFKNPKYANRSANLVQTTKEDQESTSQ- 354
CS+Y ++GH E CYKKHGFPPGF K + PK +AN+ ++++ + S
Sbjct: 309 CSFYNRVGHIAERCYKKHGFPPGFTPKGKAGEKLQKPKPL--AANVAESSEVNTSLESMV 366
Query: 355 GNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQIS 414
GN +++ +F + L PPS +++ + ++ C + I ++
Sbjct: 367 GNLSKEQLQQF---IAMFSSQLQNTPPSTYATASTSQSDNLGICFSP---STYSFIGILT 420
Query: 415 TKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPS 474
TW++D+GAT H+ + S FSS V+LP G + G+++L+
Sbjct: 421 VARHTLSSATWVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPTGPTVKISGVGTLKLNDD 480
Query: 475 FILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYIL 534
+L NVLF+P F NLIS+ L + R+ F + C IQD +M+G R V +LY+L
Sbjct: 481 ILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRMLGQGRRVANLYLL 540
Query: 535 NNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQ 594
+ S S N V + S +WH RLGH S + +I D ++
Sbjct: 541 DVGDQSI-----------SVNAVVDIS-----MWHRRLGHASLQRLDAISDSLGTTRHKN 584
Query: 595 --DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYS 652
C VC +AKQ++L FP SN IF ++H+D+WGP SV ++ G+ YFLTIVDD+S
Sbjct: 585 KGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHS 644
Query: 653 RYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTS 712
R TW+YLLK+K EV T+ F V NQ+ V VK VRSDN E + FYAEKGI+ S
Sbjct: 645 RATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFYAEKGIVSFHS 704
Query: 713 CVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCP 772
C ETP+QNS+VERKHQHILNVARAL+FQ+ +P W ++ ++FLINR P+ +L N+ P
Sbjct: 705 CPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTP 764
Query: 773 FQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTR 832
+++L P L+ FG LC++ST R KF R++ C+FLG+ SG KGY + DL +
Sbjct: 765 YEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESN 824
Query: 833 DISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLS 892
+ ISRNV FHE +FP + P +P D + P P QI L
Sbjct: 825 TVFISRNVQFHEEVFPLAKNPGSESSLKLFTPMVPVSSGIISDTTHS-PSSLPSQISDL- 882
Query: 893 SGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYH 952
P Q S SQR RK P +L DYHC N+ +P+S ISY
Sbjct: 883 --PPQIS--------SQRVRKPPAHLNDYHC-----------NTMQSDHKYPISSTISYS 921
Query: 953 KLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPI 1012
K+ PS+ +I NIT PT Y+EA + W A++ EI A+E+ NTW + P K +
Sbjct: 922 KISPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAV 981
Query: 1013 GCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKN 1072
GCKWV+ +K+ DG ++RYKARLV KGYTQ EG+D+ DTFSPVAKMTT+++LL + ++K
Sbjct: 982 GCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKK 1041
Query: 1073 WFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PNQVCLLQKSLYGLKQASRQWF 1127
WFL QLDV NAFL+ +L+EEI+M +P+G K N V L++S+YGLKQASRQWF
Sbjct: 1042 WFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWF 1101
Query: 1128 STLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQ 1187
+L LGF ++ DHTL++ K G F +L+YVDD+++A + L Q
Sbjct: 1102 KKFSSSLLSLGFKKTHGDHTLFL-KMYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQ 1160
Query: 1188 KFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSA 1247
+F+++DLG+ K+FLGL +AR+ GI + QRKYALELL +G+L K + PM + K+
Sbjct: 1161 RFKLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRK 1220
Query: 1248 SSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKG 1307
G + DI YRR++G+L+YLT TRPDI + VN+L QF SAP H AA+RVL+YIKG
Sbjct: 1221 DDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKG 1280
Query: 1308 SPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSS 1367
+ G GLFY A+S L F+DSD A C D+R+S T + MF+G SL+SWRSKKQ T SRSS
Sbjct: 1281 TVGQGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSS 1340
Query: 1368 CEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIEL 1427
EAEYRA+A CE+ WL LL LQ S P+ ++ D+ +A++IA NP +HERTKHI+L
Sbjct: 1341 AEAEYRALALATCEMVWLFTLLVSLQASPPVPI-LYSDSTAAIYIATNPVFHERTKHIKL 1399
Query: 1428 DCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHS 1479
DCH VRE++ G + LL V + Q+ADI TKPL P F H+ SK+ + +I S
Sbjct: 1400 DCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSILNIFS 1451
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 929 bits (2400), Expect = 0.0
Identities = 510/1210 (42%), Positives = 715/1210 (58%), Gaps = 83/1210 (6%)
Query: 294 SGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFK---NPKYANRSANLVQTTKEDQE 350
SG N+ CS+ ++GH E CYKKHGFPPGF K + K A Q T +
Sbjct: 304 SGPNK-GRPTCSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPDK 362
Query: 351 STSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQ------KNSTSQHTASINSCVQ---- 400
T Q E F+ DQ +L+ L Q + ++SQH AS + V
Sbjct: 363 MTGQ-----LETLAGNFSPDQIQNLIALFSSQLQPQIVSPQTASSQHEASSSQSVAPSGI 417
Query: 401 MLPGKNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNI 460
+ I ++ + + TW++D+GAT H+ + F + V+LP G
Sbjct: 418 LFSPSTYCFIGILAVSHNSLSSDTWVIDSGATHHVSHDRKLFQTLDTSIVSFVNLPTGPN 477
Query: 461 ESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACK 520
+ G++ ++ IL NVLF+P F NLIS+ L L R+ F C IQD
Sbjct: 478 VRISGVGTVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGL 537
Query: 521 MIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKG 580
+G + + +LY+L D+ SP S N+V ++WH RLGHPSF +
Sbjct: 538 TLGEGKRIGNLYVL--DTQSPAISVNAVVDV--------------SVWHKRLGHPSFSRL 581
Query: 581 QSIKDLFPYVQYT--QDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSL 638
S+ ++ ++ + C VC +AKQK+L FP +N+ + F+++H+D+WGP SV ++
Sbjct: 582 DSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVETV 641
Query: 639 NGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVL 698
G+ YFLTIVDD+SR TW+YLLKSK +V T+ F V NQ+ VK VRSDN KE
Sbjct: 642 EGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKELAF 701
Query: 699 SQFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFL 758
++FY KGI+ SC ETP+QNS+VERKHQHILNVARAL+FQ+++ +W ++ ++FL
Sbjct: 702 TEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFL 761
Query: 759 INRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFK 818
INR P+ +L N+ PF++L +LPD + LK FG LC++ST + R KF R++ CVFLG+
Sbjct: 762 INRTPSALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYP 821
Query: 819 SGTKGYLVYDLNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEY 878
G KGY + DL + + ISRNV FHE +FP + S F P D
Sbjct: 822 FGFKGYKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVF--------TPMD--- 870
Query: 879 PIPPDNPVQIQPLSSGPSQNSE------QPIVHRVSQRPRKQPTYLQDYHCTLAATSTVV 932
PLSSG S S P +R K P +LQDYHC
Sbjct: 871 -----------PLSSGNSITSHLPSPQISPSTQISKRRITKFPAHLQDYHCYFV------ 913
Query: 933 PANSSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIE 992
+K SHP+S +SY ++ PS+ +I NI+ P Y EA + W A++ EI
Sbjct: 914 -----NKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIG 968
Query: 993 ALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTF 1052
A+ER +TW + PP K +GCKWV+ +K+ DG+++R+KAR+V KGYTQ EG+D+ +TF
Sbjct: 969 AMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETF 1028
Query: 1053 SPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PN 1107
SPVAKM T+++LL + ++K W+L+QLD+ NAFL+ L+E IYM LP G K PN
Sbjct: 1029 SPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPN 1088
Query: 1108 QVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDD 1167
VC L+KS+YGLKQASRQWF +L LGF + DHTL+V + F LL+YVDD
Sbjct: 1089 VVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFV-RCIGSEFIVLLVYVDD 1147
Query: 1168 VLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDS 1227
+++A + + +L F++++LG K+FLGL +AR+ +GI L+QRKYALELL+ +
Sbjct: 1148 IVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSA 1207
Query: 1228 GLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFL 1287
+L K ++ PM + +LS + G L D YRRL+G+L+YLT TRPDI + VN+L QF
Sbjct: 1208 DMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFS 1267
Query: 1288 SAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMF 1347
SAP H AA ++VL+YIKG+ G GLFY A L ++D+D C D+R+S TG+ MF
Sbjct: 1268 SAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMF 1327
Query: 1348 LGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQ 1407
+GSSL+SWRSKKQ T SRSS EAEYRA+A CE+ WL LL L+V P+ ++ D+
Sbjct: 1328 VGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPI-LYSDST 1386
Query: 1408 SAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRH 1467
+A++IA NP +HERTKHIE+DCH VREK+ G + LL V + Q+ADI TKPL P F H
Sbjct: 1387 AAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAH 1446
Query: 1468 IFSKLRLYDI 1477
+ SK+ + +I
Sbjct: 1447 LLSKMSIQNI 1456
>At1g26990 polyprotein, putative
Length = 1436
Score = 881 bits (2277), Expect = 0.0
Identities = 505/1235 (40%), Positives = 707/1235 (56%), Gaps = 117/1235 (9%)
Query: 296 RNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQG 355
+ + KC++ ++GHTV+ CYK HG+PPG S NL T + + ++
Sbjct: 263 QGNFKKPKCTHCNRIGHTVDKCYKVHGYPPGHPRAKENTYVGSTNLASTDQIETQAPPTM 322
Query: 356 NTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGN---SISQ 412
+ E + D L+ L ST + SI SC + N SISQ
Sbjct: 323 SATGHET----MSNDHIQQLISYL-------STKLQSPSITSCFDKAIASSSNPVPSISQ 371
Query: 413 ISTK----NGNPL--------------DTT-------------------WILDTGATDHI 435
I+ K + NP+ D+T W++D+GA+ H+
Sbjct: 372 ITDKAIASSSNPVPSISQITGTFFSLYDSTYYEMLTSSIPIETELSLRAWVIDSGASHHV 431
Query: 436 CNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHK 495
+ + + +YK + V LPNG+ G IQL+ + L NVLF+P F+FNL+SV
Sbjct: 432 THERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQLTDALSLHNVLFIPEFKFNLLSVSV 491
Query: 496 LVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDS----LSPFPSCNSVSTC 551
L K+L+ +++F+ D+C+IQ M+G V +LYILN D +S FP + S+
Sbjct: 492 LTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDVSSFPGKSVCSSV 551
Query: 552 KSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKD--LFPYVQYTQDHV-CEVCPIAKQKR 608
K+++ +WH RLGHPSF K ++ D + P + +D C VC ++KQK
Sbjct: 552 KNES----------EMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQKH 601
Query: 609 LKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQT 668
L F N + F+++H+D WGP SV +++ + YFLTIVDD+SR TW+YLLK K +V T
Sbjct: 602 LPFKSVNHIREKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKSDVLT 661
Query: 669 LVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNSIVERKHQ 728
+ F V Q+ V VRSDN E ++ +A++GI C ETP+QN +VERKHQ
Sbjct: 662 VFPSFLKMVETQYHTKVCSVRSDNAHELKFNELFAKEGIKADHPCPETPEQNFVVERKHQ 721
Query: 729 HILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITFLKV 788
H+LNVARAL+FQ+ +P +W ++ ++FLINRL +P+++N+ P++ L K PD + LK
Sbjct: 722 HLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDYSSLKA 781
Query: 789 FGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHENIFP 848
FG LC+ ST RTKFD RAK C+FLG+ G KGY + D+ T +SISR+VIF+E+IFP
Sbjct: 782 FGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFYEDIFP 841
Query: 849 YKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQPIVHRV- 907
+ + + FP I LP+ D P+ +Q S P + E + V
Sbjct: 842 FA-SSNITDAAKDFFPHI-YLPAPNNDEHLPL-------VQSSSDAPHNHDESSSMIFVP 892
Query: 908 ----SQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIM 963
S R RK P++LQD+HC +T +K + +PL+ ISY L + AFI
Sbjct: 893 SEPKSTRQRKLPSHLQDFHCYNNTPTT-------TKTSPYPLTNYISYSYLSEPFGAFIN 945
Query: 964 NITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYK 1023
IT T P +YSEA + W AM EI A R TW + D P K +GCKW+ IK+
Sbjct: 946 IITATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIITIKFL 1005
Query: 1024 QDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNA 1083
DG+I+R+KARLV KGYTQ EGIDF +TFSPVAKM T++VLLSL W+LHQLD+ NA
Sbjct: 1006 ADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLHQLDISNA 1065
Query: 1084 FLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSF 1143
L+ L+EEIYM LP G + + +V K
Sbjct: 1066 LLNGDLEEEIYMKLPPGYSEIQGQEVSPNAKC---------------------------H 1098
Query: 1144 ADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGL 1203
DHTL+VK + G F +L+YVDD+L+A + L F+++DLGE KFFLG+
Sbjct: 1099 GDHTLFVK-AQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKFFLGI 1157
Query: 1204 AIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLI 1263
IAR+ GI L QRKY L+LL+ S K ++ PM+ +QKLS +GT L D YRR++
Sbjct: 1158 EIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQYRRIL 1217
Query: 1264 GRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVL 1323
G+L YL TRPDI + V++L+Q+ SAPTD+H A H++LRY+KG+ G GLFY A ++ L
Sbjct: 1218 GKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYGADTNFDL 1277
Query: 1324 TAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQ 1383
FSDSD C DTR+ +TG+ +F+G+SLVSWRSKKQ S SS EAEYRAM+ E+
Sbjct: 1278 RGFSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELI 1337
Query: 1384 WLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHL 1443
WL Y+L ++ T P ++CDN++A+HIA+N +HERTKHIE DCH VRE I G++
Sbjct: 1338 WLGYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILKT 1397
Query: 1444 LPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIH 1478
+ V + QLAD TKPL P PFR SKL L +I+
Sbjct: 1398 IFVRTDNQLADTLTKPLYPKPFRENNSKLGLLNIY 1432
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 881 bits (2276), Expect = 0.0
Identities = 519/1304 (39%), Positives = 718/1304 (54%), Gaps = 233/1304 (17%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPP-------GFKFK-NPK---------------------Y 334
C++ GK+GHT++ CYK HG+PP G+ +K NP+ Y
Sbjct: 286 CTHCGKVGHTIQKCYKVHGYPPGMKTGNTGYTYKPNPQLHVQPRMPMMPQPRMQFPAQPY 345
Query: 335 AN--RSANLV-QTTKEDQESTSQGNTQEQEAARFG------------------FTADQYH 373
N + AN+V Q E S+G +Q +G FT Q
Sbjct: 346 TNSMQKANVVAQVYAETGAYPSEGYSQAPMMNPYGSYPMPHITHGGNNLSLQDFTPQQIE 405
Query: 374 HLL------LLLPPSEQKNSTSQHTASINSCVQM-LPGKNGNSISQISTK---NGNPL-- 421
++ + +P +S A+++ M L +G I ST N L
Sbjct: 406 QMISQFQAQVQVPEPAASSSNPSPLATVSEHGFMALTSTSGTIIPFPSTSLKYENNDLKF 465
Query: 422 -------------DTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGS 468
WI+D+GA+ H+C+ L+ F K V+ G+
Sbjct: 466 QNHTLSALQKFLPSDAWIIDSGASSHVCSDLAMFRELKSVS-----------------GT 508
Query: 469 IQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAV 528
+ ++ IL NVL +P+F+FNL+SV LVK++ F D CLIQ+ + MIG R
Sbjct: 509 VHITQKLILHNVLHVPDFKFNLMSVSSLVKTISCSAHFYVDCCLIQELSQGLMIGRGRLY 568
Query: 529 RSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQN---LWHYRLGHPSFVKGQSIKD 585
+LYIL ++ SP S + C F+ SV N LWH RLGHPS V Q
Sbjct: 569 HNLYILETENTSPSTSTPAA---------CLFTGSVLNDGHLWHQRLGHPSSVVLQ---- 615
Query: 586 LFPYVQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFL 645
K KRL + N+ + F ++H+DIWGP S+ S+ GF YFL
Sbjct: 616 -------------------KLKRLAYISHNNLASNPFDLVHLDIWGPFSIESIEGFRYFL 656
Query: 646 TIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEK 705
T+VDD +R TWVY+L++K +V ++ +F V+ QF +K +RSDN E ++ E
Sbjct: 657 TVVDDCTRTTWVYMLRNKKDVSSVFPEFIKLVSTQFNAKIKAIRSDNAPELGFTEIVKEH 716
Query: 706 GIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTP 765
G++HH SC TPQQNS+VERKHQHILNVARALLFQ+++P +W+ + ++FLINRLP+P
Sbjct: 717 GMLHHFSCAYTPQQNSVVERKHQHILNVARALLFQSNIPMQYWSDCVTTAVFLINRLPSP 776
Query: 766 ILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYL 825
+L+N+ P++L+ + PD + LK FG LCF ST A RTKF RA+ CVFLG+ SG KGY
Sbjct: 777 LLNNKSPYELILNKQPDYSLLKNFGCLCFVSTNAHERTKFTPRARACVFLGYPSGYKGYK 836
Query: 826 VYDLNTRDISISRNVIFHENIFPYK----LHRDDHECESSTFPQ------IPCLPSEPFD 875
V DL + +++SRNV+F E++FP+K L++ +S P + +P D
Sbjct: 837 VLDLESHSVTVSRNVVFKEHVFPFKTSELLNKAVDMFPNSILPLPAPLHFVETMPLIDED 896
Query: 876 YEYPIPPDNPVQIQPLSSG---------PSQNSEQ--------PIVHRVSQRPRKQPTYL 918
P D+ SS PS N+E PI S+R + P+YL
Sbjct: 897 SLIPTTTDSRTADNHASSSSSALPSIIPPSSNTETQDIDSNAVPITR--SKRTTRAPSYL 954
Query: 919 QDYHCTLAATSTVVP----------------ANSSSKGTSHPLSQVISYHKLDPSYHAFI 962
+YHC+L + + +P A+S K T +P+S V+SY K P ++I
Sbjct: 955 SEYHCSLVPSISTLPPTDSSIPIHPLPEIFTASSPKKTTPYPISTVVSYDKYTPLCQSYI 1014
Query: 963 MNITTTVEPTRYSEAVKHECW-RVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIK 1021
T EP +S+A+K E W RVA+ E++A+E N TW + PPDK +GCKWV+ IK
Sbjct: 1015 FAYNTETEPKTFSQAMKSEKWIRVAVE-ELQAMELNKTWSVESLPPDKNVVGCKWVFTIK 1073
Query: 1022 YKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVD 1081
Y DGT++RYKARLV +G+TQ EGIDF+DTFSPVAK+T+ +++L L + W L Q+DV
Sbjct: 1074 YNPDGTVERYKARLVAQGFTQQEGIDFLDTFSPVAKLTSAKMMLGLAAITGWTLTQMDVS 1133
Query: 1082 NAFLHAKLDEEIYMSLPQGMNSD-----KPNQVCLLQKSLYGLKQASRQWFSTLCQALQG 1136
+AFLH LDEEI+MSLPQG PN VC L KS+YGLKQASRQW+
Sbjct: 1134 DAFLHGDLDEEIFMSLPQGYTPPAGTILPPNPVCRLLKSIYGLKQASRQWYK-------- 1185
Query: 1137 LGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGE 1196
F A L+Y+DD+++A N+ E++ +K L +F+IKDLG
Sbjct: 1186 --------------------RFVAALVYIDDIMIASNNDAEVENLKALLRSEFKIKDLGP 1225
Query: 1197 AKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDI 1256
A+FFL GLLG K ++ PMD + L GTPL +
Sbjct: 1226 ARFFL--------------------------GLLGCKPSSIPMDPTLHLVRDMGTPLPNP 1259
Query: 1257 SSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYP 1316
++YR+LIGRLLYLT TRPDI Y V+QLSQF+SAP+D+H AAH+VLRYIK +PG GL Y
Sbjct: 1260 TAYRKLIGRLLYLTITRPDITYAVHQLSQFISAPSDIHLQAAHKVLRYIKANPGQGLMYS 1319
Query: 1317 AASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMA 1376
A L FSD+D A C DTR+SI+G+C++LG+SL+SW+SKKQ SRSS E+EYR+MA
Sbjct: 1320 ADYEICLNGFSDADWAACKDTRRSISGFCIYLGTSLISWKSKKQAVASRSSTESEYRSMA 1379
Query: 1377 ATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKI 1436
CE+ WL LL+DL + T P +FCDN+SA+H + NP +HERTKHIE+DCH VR++I
Sbjct: 1380 QATCEIIWLQQLLKDLHIPLTCPAKLFCDNKSALHSSLNPVFHERTKHIEIDCHTVRDQI 1439
Query: 1437 TQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
G + L V + Q ADI TK L P PF H+ ++ L + P
Sbjct: 1440 KAGNLKALHVPTENQHADILTKALHPGPFHHLLRQMSLSSLFLP 1483
>At4g17450 retrotransposon like protein
Length = 1433
Score = 872 bits (2252), Expect = 0.0
Identities = 489/1220 (40%), Positives = 688/1220 (56%), Gaps = 107/1220 (8%)
Query: 293 NSGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQEST 352
N + Y KCSY K+GH V+ CYKKHG+PPG K+ + S NL T + T
Sbjct: 288 NMAQGSYKKPKCSYCNKLGHLVDKCYKKHGYPPGSKWTKGQTIG-STNLASTQLQPVNET 346
Query: 353 SQGNTQEQEAARFGFTADQYHHLL------LLLPPSEQKNSTSQHTASINSCVQMLPGKN 406
T E F+ DQ ++ L + + +TS + S + V M+ +
Sbjct: 347 PNEKTDSYEE----FSTDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMISQIS 402
Query: 407 GNSISQISTKNGNPLDTT-----------WILDTGATDHICNTLSYFSSYKHVAPIPVSL 455
G +S S + L ++ W++D+GAT H+ + + +++ + V L
Sbjct: 403 GTFLSLFSNAYYDMLISSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTFVRL 462
Query: 456 PNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQD 515
PN G IQLS + L NVL++P F+FNLIS +L K L
Sbjct: 463 PNDCTVKIAGIGFIQLSDAISLHNVLYIPEFKFNLIS--ELTKEL--------------- 505
Query: 516 SNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHP 575
MIG V +LY+L+ + + S ++ + VC WH RLGHP
Sbjct: 506 -----MIGRGSQVGNLYVLDFNENNHTVSLKGTTSMCPEFSVCSSVVVDSVTWHKRLGHP 560
Query: 576 SFVKGQSIKDLFPYVQYTQD-------HVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVD 628
++ K + D+ + HVC VC ++KQK L F + F ++H+D
Sbjct: 561 AYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNMCSAAFDLVHID 620
Query: 629 IWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIV 688
WGP SV + + TW+YLLK+K +V + F V Q+ +K V
Sbjct: 621 TWGPFSVPTNDA--------------TWIYLLKNKSDVLHVFPAFINMVHTQYQTKLKSV 666
Query: 689 RSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFW 748
RSDN E + +A GI+ + SC ETP+QNS+VERKHQHILNVARALLFQ+++P FW
Sbjct: 667 RSDNAHELKFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVARALLFQSNIPLEFW 726
Query: 749 AHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHR 808
++ ++FLINRLPTP+L+N+ P++ L P LK FG LC++ST R KF+ R
Sbjct: 727 GDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKTFGCLCYSSTSPKQRHKFEPR 786
Query: 809 AKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHENIFPY--KLHRDDHECESSTFPQI 866
A+ CVFLG+ G KGY + D+ T +SISR+VIFHE+IFP+ +DD + FP +
Sbjct: 787 ARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPFISSTIKDDIK---DFFPLL 843
Query: 867 PCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLA 926
P+ D P+ + + P S + P +S+R +K P +LQD+HC
Sbjct: 844 Q-FPARTDDL--PLEQTSIIDTHPHQDVSSSKALVPF-DPLSKRQKKPPKHLQDFHC--- 896
Query: 927 ATSTVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVA 986
Y+ +HAFI NIT V P RYSEA + W A
Sbjct: 897 ------------------------YNNTTEPFHAFINNITNAVIPQRYSEAKDFKAWCDA 932
Query: 987 MN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGI 1046
M EI A+ R NTW +V PP+K IGCKWV+ IK+ DG+I+RYKARLV KGYTQ EG+
Sbjct: 933 MKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGL 992
Query: 1047 DFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGM----- 1101
D+ +TFSPVAK+T++R++L L + W +HQLD+ NAFL+ LDEEIYM +P G
Sbjct: 993 DYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVG 1052
Query: 1102 NSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTAL 1161
+ P+ +C L KS+YGLKQASRQW+ L L+G+GF +S ADHTL++K A G +
Sbjct: 1053 EALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKY-ANGVLMGV 1111
Query: 1162 LLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYAL 1221
L+YVDD+++ N D + L F+++DLG AK+FLG+ IARS+KGI + QRKY L
Sbjct: 1112 LVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYIL 1171
Query: 1222 ELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVN 1281
ELLS +G LG K ++ P+D S KL+ G PL+D +SYR+L+G+L+YL TRPDIAY VN
Sbjct: 1172 ELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVN 1231
Query: 1282 QLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSI 1341
L QF APT +H +A H+VLRY+KG+ G GLFY A L ++DSD C D+R+ +
Sbjct: 1232 TLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCV 1291
Query: 1342 TGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVS 1401
YCMF+G LVSW+SKKQ T S S+ EAE+RAM+ E+ WL L D +V P
Sbjct: 1292 AAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAY 1351
Query: 1402 MFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLS 1461
++CDN +A+HI +N +HERTK +ELDC+ RE + G + + V + Q+AD TK +
Sbjct: 1352 LYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTKAIH 1411
Query: 1462 PAPFRHIFSKLRLYDIHSPV 1481
PA F + K+ + +I +P+
Sbjct: 1412 PAQFHKLIGKMGVCNIFAPL 1431
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 852 bits (2202), Expect = 0.0
Identities = 495/1261 (39%), Positives = 707/1261 (55%), Gaps = 99/1261 (7%)
Query: 285 SGSGNYHSNSGRNRYSNK---KCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANL 341
S S + H+ R+ K CS G+ GH ++C++ GFP + +N R +N
Sbjct: 274 SSSRSEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERN---GGRGSN- 329
Query: 342 VQTTKEDQESTSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQM 401
++ G Q Q A +++ S T +H ++ V+
Sbjct: 330 --GRGRGGRGSNGGRGQGQVMAAHATSSNS----------SVFPEFTEEHMRVLSQLVKE 377
Query: 402 LPGKNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIE 461
S + +G ILD+GA+ H+ TLS ++ V P PV +G+
Sbjct: 378 KSNSGSTSNNNSDRLSGKTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCPVGFADGSKA 437
Query: 462 SATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKM 521
A G + LS + L NVLF+P+ LISV KL+K + TF+D C +QD ++ +
Sbjct: 438 FALSVGVLTLSNTVSLTNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDRSSKTL 497
Query: 522 IGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQ 581
IG+ +Y L + + + + N S Q LWH RLGHPSF
Sbjct: 498 IGSGEERGGVYYLTDVTPAKIHTANVDSD--------------QALWHQRLGHPSFSVLS 543
Query: 582 SIKDLFPYVQYTQDHVCEVCPIAKQKRLKFPLS-NSTSDCIFQMIHVDIWGPVSVLSLNG 640
S+ H C+VC AKQ R FP S N T +C F +IH D+WGP V + G
Sbjct: 544 SLPLFSKTSSTVTSHSCDVCFRAKQTREVFPESINKTEEC-FSLIHCDVWGPYRVPASCG 602
Query: 641 FSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV-LS 699
YFLTIVDDYSR W YLL K EV+ ++ +F + QFG +VK+VRSDNG EF+ LS
Sbjct: 603 AVYFLTIVDDYSRAVWTYLLLEKSEVRQVLTNFLKYAEKQFGKTVKMVRSDNGTEFMCLS 662
Query: 700 QFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLI 759
++ E GIIH TSCV TPQQN VERKH+HILNVARALLFQA LP FW +I+ + +LI
Sbjct: 663 SYFRENGIIHQTSCVGTPQQNGRVERKHRHILNVARALLFQASLPIKFWGESILTAAYLI 722
Query: 760 NRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKS 819
NR P+ IL + P+++LH P + L+VFGS C+ + + KF R++ C+F+G+
Sbjct: 723 NRTPSSILSGRTPYEVLHGSKPVYSQLRVFGSACYVHRVTRDKDKFGQRSRSCIFVGYPF 782
Query: 820 GTKGYLVYDLNTRDISISRNVIFHENIFPYK--------------LHRDDH--------- 856
G KG+ VYD+ + +SR+VIF E +FPY + DD
Sbjct: 783 GKKGWKVYDIERNEFLVSRDVIFREEVFPYAGVNSSTLASTSLPTVSEDDDWAIPPLEVR 842
Query: 857 -ECESSTFPQIPCLPSEPF------DYEYP--------IPPDNPVQIQPLSSGPSQNSEQ 901
+S ++ C E D E P PP +P+ + P SG
Sbjct: 843 GSIDSVETERVVCTTDEVVLDTSVSDSEIPNQEFVPDDTPPSSPLSVSP--SGSPNTPTT 900
Query: 902 PIV-------------HRVSQRPRKQPTYLQDY--HCTLAATSTV--VPANSSSKGTS-- 942
PIV R S+R P L DY + + S++ +PA+ S T
Sbjct: 901 PIVVPVASPIPVSPPKQRKSKRATHPPPKLNDYVLYNAMYTPSSIHALPADPSQSSTVPG 960
Query: 943 ---HPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNT 999
PL+ +S S+ A++ IT VEP + EAV+ + W AM E++ALE N T
Sbjct: 961 KSLFPLTDYVSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKVWNDAMFTEVDALEINKT 1020
Query: 1000 WLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMT 1059
W +VD PP K IG +WV++ KY DGT++RYKARLVV+G Q+EG D+ +TF+PV +MT
Sbjct: 1021 WDIVDLPPGKVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKETFAPVVRMT 1080
Query: 1060 TLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGL 1119
T+R LL V+ W ++Q+DV NAFLH L+EE+YM LP G P++VC L+KSLYGL
Sbjct: 1081 TVRTLLRNVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHSHPDKVCRLRKSLYGL 1140
Query: 1120 KQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIK 1179
KQA R WF L +L GF QS+ D++L+ + +L+YVDD+L+ GND ++
Sbjct: 1141 KQAPRCWFKKLSDSLLRFGFVQSYEDYSLF-SYTRNNIELRVLIYVDDLLICGNDGYMLQ 1199
Query: 1180 LVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPM 1239
K L + F +KDLG+ K+FLG+ ++R +GI L+QRKYAL++++DSG LG + A TP+
Sbjct: 1200 KFKDYLSRCFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADSGNLGSRPAHTPL 1259
Query: 1240 DCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAH 1299
+ + L++ G LSD YRRL+GRLLYL TRP+++Y V+ L+QF+ P + H AA
Sbjct: 1260 EQNHHLASDDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNPREAHFDAAL 1319
Query: 1300 RVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKK 1359
RV+RY+KGSPG G+ A L + DSD C TR+SI+ Y + LG S +SW++KK
Sbjct: 1320 RVVRYLKGSPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVLLGGSPISWKTKK 1379
Query: 1360 QTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYH 1419
Q T S SS EAEYRAM+ + E++WL LL++L + Q++P ++CD+++A+HIA NP +H
Sbjct: 1380 QDTVSHSSAEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAIHIAANPVFH 1439
Query: 1420 ERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHS 1479
ERTKHIE DCH VR+ + G++ V ++ QLAD+FTK L F ++ SKL + ++H+
Sbjct: 1440 ERTKHIESDCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLYLMSKLGVQNLHT 1499
Query: 1480 P 1480
P
Sbjct: 1500 P 1500
>At1g57640
Length = 1444
Score = 822 bits (2122), Expect = 0.0
Identities = 475/1252 (37%), Positives = 695/1252 (54%), Gaps = 144/1252 (11%)
Query: 298 RYSNKK-CSYYGKMGHTVEDCYKKHGFP-------------------------PGFKFKN 331
+ NKK C++ + GH+ E+C+ G+P PGF
Sbjct: 267 KLQNKKLCTHCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGRGRFGPGFNGGQ 326
Query: 332 PKYANRSANLVQTTK-EDQESTSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQ 390
P+ N+V T E ++ T A G T +Q+ ++ LL N ++
Sbjct: 327 PRPTY--VNVVMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWRGVVKLLNAGRSDNKSNA 384
Query: 391 HTASINSCVQMLPGKNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAP 450
H +C L T+WILDTGA+ H+ L S + ++P
Sbjct: 385 HETQSGTC---------------------SLFTSWILDTGASHHMTGNLELLSDMRSMSP 423
Query: 451 IPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDD 510
+ + L +GN A +G+++L IL +V ++ E +LISV +++
Sbjct: 424 VLIILADGNKRVAVSEGTVRLGSHLILKSVFYVKELESDLISVGQMM------------- 470
Query: 511 CLIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHY 570
D N C A AV + +PF +LWH
Sbjct: 471 ----DENHCV---NAAAVHT------SVKAPF-----------------------DLWHR 494
Query: 571 RLGHPSF-VKGQSIKDLFPYVQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDI 629
RLGH S + ++L + ++VC+ C AKQ R FPLS++ S FQ+IH D+
Sbjct: 495 RLGHASDKIVNLLPRELLSSGKEILENVCDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDV 554
Query: 630 WGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVR 689
WGP S +G YFLTIVDDYSR WVYL+ K E Q +KDF A V QF +KIVR
Sbjct: 555 WGPYRAPSYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVR 614
Query: 690 SDNGKEFV-LSQFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFW 748
SDNG EF+ + +++ KGI H TSCV TP QN VERKH+HILN+ARAL FQ++LP FW
Sbjct: 615 SDNGTEFLCMREYFLHKGIAHETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFW 674
Query: 749 AHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHR 808
I+ + +LINR P+ +L + P+++L+K P + L+VFGSLC+A KF R
Sbjct: 675 GECILSAAYLINRTPSMLLQGKSPYEMLYKTAPKYSHLRVFGSLCYAHNQNHKGDKFAAR 734
Query: 809 AKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHENIFPYK---LHRDDH----EC--- 858
++RCVF+G+ G KG+ ++DL + +SR+VIF E FPY + +D +C
Sbjct: 735 SRRCVFVGYPHGQKGWRLFDLEEQKFFVSRDVIFQETEFPYSKMSCNEEDERVLVDCVGP 794
Query: 859 ------------------ESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSE 900
E++ P + P P + P V + L + ++
Sbjct: 795 PFIEEAIGPRTIIGRNIGEATVGPNVATGPIIPEINQESSSPSEFVSLSSLDPFLASSTV 854
Query: 901 Q------------PIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQV 948
Q PI R S R ++P L+++ + ++ P SSS + +P+ +
Sbjct: 855 QTADLPLSSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVESISPEASSS--SLYPIEKY 912
Query: 949 ISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPD 1008
+ H+ S+ AF+ +T +EPT Y+EA+ + WR AM+ EIE+L N T+ +V+ PP
Sbjct: 913 VDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSIVNLPPG 972
Query: 1009 KTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLV 1068
K +G KWVY+IKY+ DG I+RYKARLVV G Q EG+D+ +TF+PVAKM+T+R+ L +
Sbjct: 973 KRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVRLFLGVA 1032
Query: 1069 STKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFS 1128
+ ++W +HQ+DV NAFLH L EE+YM LPQG D P++VC L KSLYGLKQA R WFS
Sbjct: 1033 AARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCDDPSKVCRLHKSLYGLKQAPRCWFS 1092
Query: 1129 TLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQK 1188
L AL+ GFTQS +D++L+ + G F +L+YVDD++++G+ D + K L
Sbjct: 1093 KLSSALKQYGFTQSLSDYSLFSYNN-DGIFVHVLVYVDDLIISGSCPDAVAQFKSYLESC 1151
Query: 1189 FRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSAS 1248
F +KDLG K+FLG+ ++R+ +G L+QRKY L+++S+ GLLG + + P++ + KLS S
Sbjct: 1152 FHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQNHKLSLS 1211
Query: 1249 SGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGS 1308
+ LSD S YRRL+GRL+YL TRP+++Y V+ L+QF+ P H AA RV+RY+K +
Sbjct: 1212 TSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVVRYLKSN 1271
Query: 1309 PGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSC 1368
PG G+ + S+ + + DSD A C TR+S+TGY + LG + +SW++KKQ T SRSS
Sbjct: 1272 PGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQPTVSRSSA 1331
Query: 1369 EAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELD 1428
EAEYRAMA E+ WL +L DL VS + +F D++SA+ ++ NP HERTKH+E+D
Sbjct: 1332 EAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHERTKHVEVD 1391
Query: 1429 CHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
CH +R+ I G++ V S QLADI TK L R+ KL + D+H+P
Sbjct: 1392 CHFIRDAILDGIIATSFVPSHKQLADILTKALGEKEVRYFLRKLGILDVHAP 1443
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 805 bits (2080), Expect = 0.0
Identities = 469/1255 (37%), Positives = 688/1255 (54%), Gaps = 105/1255 (8%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAA 363
C++ + GH V C+ HG+P + +NP+ S + S GN
Sbjct: 260 CTHCNRKGHEVTQCFLVHGYPDWWLEQNPQENQPSTRGRGSNGRGSSSGRGGNRSSAPTT 319
Query: 364 RFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQISTKNGNPLDT 423
R A+ P+ + Q I+ P + +S GN T
Sbjct: 320 RGRGRANNAQ----AAAPTVSGDGNDQIAQLISLLQAQRPSSSSERLS------GNTCLT 369
Query: 424 TWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFL 483
++DTGA+ H+ S + P PV+ P+G AT G++ L S+ L +VLF+
Sbjct: 370 DGVIDTGASHHMTGDCSILVDVFDITPSPVTKPDGKASQATKCGTLLLHDSYKLHDVLFV 429
Query: 484 PNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFP 543
P+F+ LISV KL+K F+D C +QD +IG +Y
Sbjct: 430 PDFDCTLISVSKLLKQTSSIAIFTDTFCFLQDRFLRTLIGAGEEREGVYYFTG------- 482
Query: 544 SCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQ-YTQDHVCEVCP 602
V + +F+ S +LWH RLGHPS S+ + Q + + C+ C
Sbjct: 483 ----VLAPRVHKASSDFAIS-GDLWHRRLGHPSTSVLLSLPECNRSSQGFDKIDSCDTCF 537
Query: 603 IAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKS 662
+KQ R FP+SN+ + F +IH D+WGP S G YFLT+VDDYSR W YL+ S
Sbjct: 538 RSKQTREVFPISNNKTMECFSLIHGDVWGPYRTPSTTGAVYFLTLVDDYSRSVWTYLMSS 597
Query: 663 KDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV-LSQFYAEKGIIHHTSCVETPQQNS 721
K EV L+K+FCA QFG VK R+DNG EF+ L+ ++ GI+H TSCV+TPQQN
Sbjct: 598 KTEVSQLIKNFCAMSERQFGKQVKAFRTDNGTEFMCLTPYFQTHGILHQTSCVDTPQQNG 657
Query: 722 IVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLP 781
VERKH+HILNVARA LFQ +LP FW +I+ + LINR P+ +L + P++LL + P
Sbjct: 658 RVERKHRHILNVARACLFQGNLPVKFWGESILTATHLINRTPSAVLKGKTPYELLFGERP 717
Query: 782 DITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNT------RDIS 835
L+ FG LC+A ++ KF R+++CVF+G+ G K + VYDL T RD+
Sbjct: 718 SYDMLRSFGCLCYAHIRPRNKDKFTSRSRKCVFIGYPHGKKAWRVYDLETGKIFASRDVR 777
Query: 836 ISRNV--------------------IFHENIFPYKLHRDDHECESST------------- 862
++ + + P D +SS+
Sbjct: 778 FHEDIYPYATATQSNVPLPPPTPPMVNDDWFLPISTQVDSTNVDSSSSSSPAQSGSIDQP 837
Query: 863 ---FPQIPCLPSEPFDYEYP--IPPDNPVQ--------------IQPLSSGPSQNSEQPI 903
Q P + P E +P +P + + LS+G S P
Sbjct: 838 PRSIDQSPSTSTNPVPEEIGSIVPSSSPSRSIDRSTSDLSASDTTELLSTGESSTPSSPG 897
Query: 904 VHRV---SQRPRKQPTYLQDYHCT-----------LAATSTVVP--------ANSSSKGT 941
+ + R +K+ L+D+ + + S V+P A+S S T
Sbjct: 898 LPELLGKGCREKKKSVLLKDFVTNTTSKKKTASHNIHSPSQVLPSGLPTSLSADSVSGKT 957
Query: 942 SHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWL 1001
+PLS ++ ++ AF+ I + EP + +A+ + W AM+ EI+ALE N+TW
Sbjct: 958 LYPLSDFLTNSGYSANHIAFMAAILDSNEPKHFKDAILIKEWCEAMSKEIDALEANHTWD 1017
Query: 1002 LVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTL 1061
+ D P K I KWVY++KY DGT++R+KARLVV G Q EG+DF +TF+PVAK+TT+
Sbjct: 1018 ITDLPHGKKAISSKWVYKLKYNSDGTLERHKARLVVMGNHQKEGVDFKETFAPVAKLTTV 1077
Query: 1062 RVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQ 1121
R +L++ + K+W +HQ+DV NAFLH L+EE+YM LP G P++VC L+KSLYGLKQ
Sbjct: 1078 RTILAVAAAKDWEVHQMDVHNAFLHGDLEEEVYMRLPPGFKCSDPSKVCRLRKSLYGLKQ 1137
Query: 1122 ASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLV 1181
A R WFS L AL+ +GFTQS+ D++L+ K+ + +L+YVDD+++AGN++D I
Sbjct: 1138 APRCWFSKLSTALRNIGFTQSYEDYSLFSLKNGD-TIIHVLVYVDDLIVAGNNLDAIDRF 1196
Query: 1182 KHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDC 1241
K LH+ F +KDLG+ K+FLGL ++R G L+QRKYAL+++ ++GLLG K + P+
Sbjct: 1197 KSQLHKCFHMKDLGKLKYFLGLEVSRGPDGFCLSQRKYALDIVKETGLLGCKPSAVPIAL 1256
Query: 1242 SQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRV 1301
+ KL++ +G ++ YRRL+GR +YLT TRPD++Y V+ LSQF+ AP H AA R+
Sbjct: 1257 NHKLASITGPVFTNPEQYRRLVGRFIYLTITRPDLSYAVHILSQFMQAPLVAHWEAALRL 1316
Query: 1302 LRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQT 1361
+RY+KGSP G+F + SS ++ A+ DSD C TR+S++ Y ++LG S +SW++KKQ
Sbjct: 1317 VRYLKGSPAQGIFLRSDSSLIINAYCDSDYNACPLTRRSLSAYVVYLGDSPISWKTKKQD 1376
Query: 1362 TTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHER 1421
T S SS EAEYRAMA T+ E++WL LL+DL V +SP+ + CD+++A+HIA NP +HER
Sbjct: 1377 TVSYSSAEAEYRAMAYTLKELKWLKALLKDLGVHHSSPMKLHCDSEAAIHIAANPVFHER 1436
Query: 1422 TKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYD 1476
TKHIE DCH VR+ + L+ + + Q+AD+ TK L F + S L + D
Sbjct: 1437 TKHIESDCHKVRDAVLDKLITTEHIYTEDQVADLLTKSLPRPTFERLLSTLGVTD 1491
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 740 bits (1911), Expect = 0.0
Identities = 454/1227 (37%), Positives = 682/1227 (55%), Gaps = 95/1227 (7%)
Query: 291 HSNSGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQE 350
+ G + + C + + GH E CY G+P ++ + +RS +QT
Sbjct: 164 YQGRGDEKDKSMVCKHCNRSGHASESCYAVIGYP---EWWGDRPRSRS---LQTRGRGGT 217
Query: 351 STSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSI 410
++S G G A Y + + + N + A+ + G NG +
Sbjct: 218 NSSGGR---------GRGAAAYANRVTV------PNHDTYEQANYALTDEDRDGVNGLTD 262
Query: 411 SQISTKNGNPLDTTWILDTG---ATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKG 467
SQ T IL++G AT+ + + + PI + L +G + +G
Sbjct: 263 SQWRTIKS-------ILNSGKDAATEKLTD----------MDPILIVLADGRERISVKEG 305
Query: 468 SIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARA 527
+++L + ++I+V ++ F+ +LIS+ +L+ R L SD ++QD + ++G R
Sbjct: 306 TVRLGSNLVMISVFYVEEFQSDLISIGQLMDENRCVLQMSDRFLVVQDRTSRMVMGAGRR 365
Query: 528 VRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLF 587
V + + ++ SV+ + +N LWH R+GHP+ I +
Sbjct: 366 VGGTFHFRSTEIAA-----SVTVKEEKNY---------ELWHSRMGHPAARVVSLIPESS 411
Query: 588 PYVQYTQ-DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLT 646
V T + C+VC AKQ R FPLS + + IF++I+ D+WGP S G YFLT
Sbjct: 412 VSVSSTHLNKACDVCHRAKQTRNSFPLSINKTLRIFELIYCDLWGPYRTPSHTGARYFLT 471
Query: 647 IVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV-LSQFYAEK 705
I+DDYSR W+YLL K E +K+F A QF V +K VRSDNG EF+ L++F+ E+
Sbjct: 472 IIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNVKIKTVRSDNGTEFLCLTKFFQEQ 531
Query: 706 GIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTP 765
G+IH SCV TP++N VERKH+H+LNVARAL FQA+LP FW ++ + +LINR P+
Sbjct: 532 GVIHERSCVATPERNDRVERKHRHLLNVARALRFQANLPIQFWGECVLTAAYLINRTPSS 591
Query: 766 ILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYL 825
+L++ P++ LHK+ P L+VFGSLC+A KF R++RCVF+G+ G KG+
Sbjct: 592 VLNDSTPYERLHKKQPRFDHLRVFGSLCYAHNRNRGGDKFAERSRRCVFVGYPHGQKGWR 651
Query: 826 VYDLNTRDISISRNVIFHENIFPYKLHRD----DHECESSTFPQIPCLPSEPFDYEY--- 878
++DL + +SR+V+F E FP+++ + + E E+ P + L E
Sbjct: 652 LFDLEQNEFFVSRDVVFSELEFPFRISHEQNVIEEEEEALWAPIVDGLIEEEVHLGQNAG 711
Query: 879 PIPP---DNPVQIQPLSSGPSQNSEQPIVHRVSQRPR---------KQPTYLQDYHCTLA 926
P PP +P+ SS ++ P+ V P PT LQ + A
Sbjct: 712 PTPPICVSSPISPSATSSRSEHSTSSPLDTEVVPTPATSTTSASSPSSPTNLQFLPLSRA 771
Query: 927 ATSTVVPANSSSKGTSHPLSQVISYHKLDP-SYHAFIMNITTTVE-PTR-----YSEAVK 979
+T A + + P + + +K P + F++N T E P++ Y +
Sbjct: 772 KPTT---AQAVAPPAVPPPRRQSTRNKAPPVTLKDFVVNTTVCQESPSKLNSILYQLQKR 828
Query: 980 HECWRVAMN*E----IEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARL 1035
+ R + + I+A E N+TW + D PP K IG +WVY++K+ DG+++RYKARL
Sbjct: 829 DDTRRFSASHTTYVAIDAQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARL 888
Query: 1036 VVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYM 1095
V G Q EG D+ +TF+PVAKM T+R+ L + +NW +HQ+DV NAFLH L EE+YM
Sbjct: 889 VALGNKQKEGEDYGETFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYM 948
Query: 1096 SLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLY--VKKS 1153
LP G + PN+VC L+K+LYGLKQA R WF L AL+ GF QS AD++L+ VK S
Sbjct: 949 KLPPGFEASHPNKVCRLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGS 1008
Query: 1154 ATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGII 1213
+L+YVDD+++ GN + K L F +KDLG K+FLG+ +ARS GI
Sbjct: 1009 VR---IKILIYVDDLIITGNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIY 1065
Query: 1214 LNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTR 1273
+ QRKYAL+++S++GLLG K A P++ + KL S+ L+D YRRL+GRL+YL TR
Sbjct: 1066 ICQRKYALDIISETGLLGVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTR 1125
Query: 1274 PDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAG 1333
D+A+ V+ L++F+ P + H AAA RV+RY+K PG G+F + +T + DSD AG
Sbjct: 1126 LDLAFSVHILARFMQEPREDHWAAALRVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAG 1185
Query: 1334 CVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQ 1393
+R+S+TGY + G S +SW++KKQ T S+SS EAEYRAM+ E+ WL LL L
Sbjct: 1186 DPMSRRSVTGYFVQFGDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLG 1245
Query: 1394 VSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLA 1453
VS P+ M CD++SA++IA NP +HERTKHIE+D H VR++ +G++ V ++ QLA
Sbjct: 1246 VSHVQPMIMCCDSKSAIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLA 1305
Query: 1454 DIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
DIFTKPL F KL + ++++P
Sbjct: 1306 DIFTKPLGRDCFSAFRIKLGIRNLYAP 1332
>At1g53810
Length = 1522
Score = 615 bits (1587), Expect = e-176
Identities = 387/1121 (34%), Positives = 571/1121 (50%), Gaps = 93/1121 (8%)
Query: 425 WILDTGATDHICNTLSYFS-SYKHVAPIPVSLPNGNIESATIKGSIQLSPS---FILINV 480
WI D+ A+ H+ N S + + + +GN T GS ++ S L V
Sbjct: 326 WIPDSAASAHVTNNRHVLQQSQPYHGSDSIMVADGNFLPITHTGSGSIASSSGKIPLKEV 385
Query: 481 LFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLS 540
L P+ +L+SV KL + F D I D K++ R LY L L
Sbjct: 386 LVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKATKKLLVMGRNRDGLYSLEEPKLQ 445
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPS------FVKGQSIKDLFPYVQYTQ 594
S S +WH RLGH + +SI + V+
Sbjct: 446 VLYSTRQNSASSE-------------VWHRRLGHANAEVLHQLASSKSIIIINKVVKT-- 490
Query: 595 DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + K RL F LS + + IH D+WGP S+ GF Y++ +D YSR+
Sbjct: 491 --VCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPSPTSSVQGFRYYVVFIDHYSRF 548
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYA---EKGIIHHT 711
TW Y LK K + + F V NQ G +KI + D G EF+ SQF + GI +
Sbjct: 549 TWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHGIQQNM 608
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN-Q 770
SC TPQQN + ERKH+HI+ + +++FQ+ LP +W + + F+IN LPT LDN +
Sbjct: 609 SCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSSLDNNE 668
Query: 771 CPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLN 830
P+Q L+ + P+ + L+VFG C+ + TKFD R+ +CVFLG+ KGY
Sbjct: 669 SPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGYRCLYPP 728
Query: 831 TRDISISRNVIFHENIFPYK-----LHRDDH----ECESSTFPQIPCLPSEPFDYEYPIP 881
T I ISR+V+F EN P++ LH D E +F + P++P YP+
Sbjct: 729 TGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVT--PTQPDQSRYPVS 786
Query: 882 PDNPVQIQPLSSGPSQ--------------NSEQPIVHRVSQRPRK-----QPTYLQDYH 922
+ LS+ P+ + + + VS P + + YH
Sbjct: 787 SIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIGDSYH 846
Query: 923 CTLAATSTVVPANSS--SKGTSHPLSQVISYHKLDP--SYHAFIMN-------------- 964
A +S PA SS S P+ + P + HA +
Sbjct: 847 SPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKRYVL 906
Query: 965 ITTTV---EPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIK 1021
+T V EP +EA+KH W AM E+ + TW LV P+ +G WV+R K
Sbjct: 907 LTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGSMWVFRTK 966
Query: 1022 YKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVD 1081
DG++D+ KARLV KG+ Q EGID+++T+SPV + T+R++L + + W L Q+DV
Sbjct: 967 LHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLKWELKQMDVK 1026
Query: 1082 NAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFT 1140
NAFLH L E +YM P G ++ KP+ VCLL KSLYGLKQ+ R WF L GF
Sbjct: 1027 NAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFGFI 1086
Query: 1141 QSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFF 1200
S D +L+V S+ LLLYVDD+++ GN+ + + +L+++FR+KD+G+ +F
Sbjct: 1087 CSLFDPSLFVY-SSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMKDMGQVHYF 1145
Query: 1201 LGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYR 1260
LG+ I G+ ++Q+KYA +LL + + TP+ ++ SD + +R
Sbjct: 1146 LGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEVFSDPTYFR 1205
Query: 1261 RLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASS 1320
L G+L YLT TRPDI + VN + Q + P+ R+LRYIKG+ G+ Y + SS
Sbjct: 1206 SLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMGIQYNSNSS 1265
Query: 1321 TV---------LTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAE 1371
+V L+A+SDSD A C +TR+S+ GYC F+G +++SW SKKQ T SRSS EAE
Sbjct: 1266 SVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPTVSRSSTEAE 1325
Query: 1372 YRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHI 1431
YR+++ T E++W+ +L+++ VS +FCDN SA+++ NP++H RTKH ++D H
Sbjct: 1326 YRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHARTKHFDVDHHY 1385
Query: 1432 VREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
+RE++ + + + LQLADIFTK L F + KL
Sbjct: 1386 IRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKL 1426
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 614 bits (1583), Expect = e-175
Identities = 401/1080 (37%), Positives = 561/1080 (51%), Gaps = 163/1080 (15%)
Query: 425 WILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLP 484
WI+D+GA+ H+C+ L+ F HV+ + V+LPNG + T G+I ++ + IL NVL +P
Sbjct: 99 WIIDSGASSHVCSDLTMFRELIHVSGVTVTLPNGTRVAITHTGTICITSTLILHNVLLVP 158
Query: 485 NFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPS 544
+F+FNLISV LVK+L + F D C IQ+ MIG + +LYIL S PS
Sbjct: 159 DFKFNLISVCCLVKTLSYSAHFFADCCYIQELTRGLMIGRGKTYNNLYILETQRTSFSPS 218
Query: 545 CNSVSTCKSQNLVCEFSPSVQN---LWHYRLGHPSFVKGQSIKDLFPYVQYTQDHVCEVC 601
+ S+ F+ +VQ+ LWH RLG + Y+ Y + +
Sbjct: 219 LPAASS---------FTGTVQDDCLLWHQRLG------------IRHYLHYRNLYFLTLV 257
Query: 602 PIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLK 661
+ + + N + VS N F F+ ++ +++Y
Sbjct: 258 DDCTRTTWVYMMKNKSE--------------VS----NIFPVFVKLI--FTQYNAKIKAI 297
Query: 662 SKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNS 721
D V+ L F FV Q G+IH SC TPQQNS
Sbjct: 298 RSDNVKELA--FTKFVKEQ-------------------------GMIHQFSCAYTPQQNS 330
Query: 722 IVER------------KHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN 769
+VER H +++ R ++F + + S ++ P IL
Sbjct: 331 VVERYPSGYKGYKVLDLESHSISITRNVVFHETKFPFKTSKFLKES---VDMFPNSILPL 387
Query: 770 QCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDL 829
P + + +P L+ + S AS + L T+ D+
Sbjct: 388 PAPLHFV-ESMPLDDDLRADDNNASTSNSASSASSIPP-------LPSTVNTQNTDALDI 439
Query: 830 NTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYP---IPPDNPV 886
+T + I+R + + ++ C S F P+ E P IPP
Sbjct: 440 DTNSVPIAR----PKRNAKAPAYLSEYHCNSVPFLS-SLSPTTSTSIETPSSSIPPKKIT 494
Query: 887 QIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLS 946
P+S+ S + P+ H Y C
Sbjct: 495 TPYPMSTAISYDKLTPLFH--------------SYICA---------------------- 518
Query: 947 QVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKP 1006
+ ++ AF T ++ +++ A E + ALE+N TW++
Sbjct: 519 -----YNVETEPKAF----TQAMKSEKWTRAANEE---------LHALEQNKTWIVESLT 560
Query: 1007 PDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLS 1066
K +GCKWV+ IKY DG+I+RYKARLV +G+TQ EGID+M+TFSPVAK ++++LL
Sbjct: 561 EGKNVVGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLG 620
Query: 1067 LVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMN-----SDKPNQVCLLQKSLYGLKQ 1121
L + W L Q+DV NAFLH +LDEEIYMSLPQG S VC L KSLYGLKQ
Sbjct: 621 LAAATGWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQ 680
Query: 1122 ASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLV 1181
ASRQW+ L G F QS AD+T++VK S T S +L+YVDD+++A ND ++ +
Sbjct: 681 ASRQWYKRLSSVFLGANFIQSPADNTMFVKVSCT-SIIVVLVYVDDLMIASNDSSAVENL 739
Query: 1182 KHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDC 1241
K L +F+IKDLG A+FFLGL IARS +GI + QRKYA LL D GL G K ++ PMD
Sbjct: 740 KELLRSEFKIKDLGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDP 799
Query: 1242 SQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRV 1301
+ L+ GT L + +SYR L+GRLLYL TRPDI + V+ LSQFLSAPTD+H AAH+V
Sbjct: 800 NLHLTKEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKV 859
Query: 1302 LRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQT 1361
LRY+KG+PG GL Y A+S L FSD+D C D+R+S+TG+C++LG+SL++W+SKKQ+
Sbjct: 860 LRYLKGNPGQGLMYSASSELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQS 919
Query: 1362 TTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHER 1421
SRSS E+EYR++A CE+ WL LL+DL V+ T P +FCDN+SA+H+A NP +HER
Sbjct: 920 VVSRSSTESEYRSLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHER 979
Query: 1422 TKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSPV 1481
TKHIE+DCH VR++I G + L V + QLADI TKPL P IFS L + SPV
Sbjct: 980 TKHIEIDCHTVRDQIKAGKLKTLHVPTGNQLADILTKPLHPVQ-SPIFSLLFRFTGTSPV 1038
>At3g61330 copia-type polyprotein
Length = 1352
Score = 585 bits (1509), Expect = e-167
Identities = 355/1064 (33%), Positives = 575/1064 (53%), Gaps = 77/1064 (7%)
Query: 425 WILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQL----SPSFILINV 480
W LD+GA++H+C S F+ V+L + + KG+I + + NV
Sbjct: 335 WYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNV 394
Query: 481 LFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLS 540
++P+ + N++S+ +L++ + + D++ I+D + + + +++LN
Sbjct: 395 YYIPSMKTNILSLGQLLEK-GYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR--- 450
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF--VKGQSIKDL---FPYVQYTQD 595
N ++ C +C S LWH R GH +F ++ S K++ P + + +
Sbjct: 451 -----NDIAQCLK---MCYKEESW--LWHLRFGHLNFGGLELLSRKEMVRGLPCINHP-N 499
Query: 596 HVCEVCPIAKQKRLKFPL-SNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + KQ ++ FP S+S + ++IH D+ GP+ SL +YFL +DD+SR
Sbjct: 500 QVCEGCLLGKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRK 559
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQF--YAE-KGIIHHT 711
TWVY LK K EV + K F A V + G+ +K +RSD G EF +F Y E GI
Sbjct: 560 TWVYFLKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQL 619
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQC 771
+ +PQQN +VERK++ IL +AR++L LPK WA A+ +++L+NR PT + +
Sbjct: 620 TVPRSPQQNGVVERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKT 679
Query: 772 PFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNT 831
P + + P ++ L+VFGS+ A R+K D ++++ +F+G+ + +KGY +Y+ +T
Sbjct: 680 PQEAWSGRKPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDT 739
Query: 832 RDISISRNVIF-HENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQP 890
+ ISRN++F E + + + +D+ + FP F+ + P P
Sbjct: 740 KKTIISRNIVFDEEGEWDWNSNEEDY----NFFPH--------FEEDEPEP--------- 778
Query: 891 LSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVIS 950
E+P S+ P PT + TS+ + +SS + Q +
Sbjct: 779 -------TREEP----PSEEPTTPPT---------SPTSSQIEESSSERTPRFRSIQEL- 817
Query: 951 YHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKT 1010
Y + + + + EP + +A++ + WR AM+ EI+++++N+TW L P
Sbjct: 818 YEVTENQENLTLFCLFAECEPMDFQKAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHK 877
Query: 1011 PIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVST 1070
IG KWVY+ K G ++RYKARLV KGY+Q GID+ + F+PVA++ T+R+++SL +
Sbjct: 878 AIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRVGIDYDEVFAPVARLETVRLIISLAAQ 937
Query: 1071 KNWFLHQLDVDNAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFST 1129
W +HQ+DV +AFL+ L+EE+Y+ PQG + + ++V L+K LYGLKQA R W +
Sbjct: 938 NKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTR 997
Query: 1130 LCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKF 1189
+ + + F + +H LY+K A L YVDD++ GN+ + K + ++F
Sbjct: 998 IDKYFKEKDFIKCPYEHALYIKIQKEDILIACL-YVDDLIFTGNNPSIFEEFKKEMTKEF 1056
Query: 1190 RIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASS 1249
+ D+G ++LG+ + + GI + Q YA E+L + TPM+C KLS
Sbjct: 1057 EMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKIDDSNPVCTPMECGIKLSKKE 1116
Query: 1250 GTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSP 1309
D ++++ L+G L YLT TRPDI Y V +S+++ PT H AA R+LRYIKG+
Sbjct: 1117 EGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTV 1176
Query: 1310 GCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCE 1369
GL Y S L +SDSD G VD RKS +G+ ++G + +W SKKQ + S+CE
Sbjct: 1177 NFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCE 1236
Query: 1370 AEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDC 1429
AEY A + VC WL LL++L + Q P +F DN+SA+ +A NP +H+R+KHI+
Sbjct: 1237 AEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRY 1296
Query: 1430 HIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLR 1473
H +RE +++ V L V + Q+AD FTKPL R F K+R
Sbjct: 1297 HYIRECVSKKDVQLEYVKTHDQVADFFTKPLK----RENFIKMR 1336
>At1g48710 hypothetical protein
Length = 1352
Score = 584 bits (1506), Expect = e-166
Identities = 355/1064 (33%), Positives = 573/1064 (53%), Gaps = 77/1064 (7%)
Query: 425 WILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQL----SPSFILINV 480
W LD+GA++H+C S F+ V+L + + KG+I + + NV
Sbjct: 335 WYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNV 394
Query: 481 LFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLS 540
++P+ + N++S+ +L++ + + D++ I+D + + + +++LN
Sbjct: 395 YYIPSMKTNILSLGQLLEK-GYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR--- 450
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF--VKGQSIKDL---FPYVQYTQD 595
N ++ C +C S LWH R GH +F ++ S K++ P + + +
Sbjct: 451 -----NDIAQCLK---MCYKEESW--LWHLRFGHLNFGGLELLSRKEMVRGLPCINHP-N 499
Query: 596 HVCEVCPIAKQKRLKFPL-SNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + KQ ++ FP S+S + ++IH D+ GP+ SL +YFL +DD+SR
Sbjct: 500 QVCEGCLLGKQFKMSFPKESSSRAQKSLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRK 559
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQF--YAE-KGIIHHT 711
TWVY LK K EV + K F A V + G+ +K +RSD G EF +F Y E GI
Sbjct: 560 TWVYFLKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQL 619
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQC 771
+ +PQQN + ERK++ IL +AR++L LPK WA A+ +++L+NR PT + +
Sbjct: 620 TVPRSPQQNGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKT 679
Query: 772 PFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNT 831
P + + ++ L+VFGS+ A R+K D ++++ +F+G+ + +KGY +Y+ +T
Sbjct: 680 PQEAWSGRKSGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDT 739
Query: 832 RDISISRNVIF-HENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQP 890
+ ISRN++F E + + + +D+ + FP F+ + P P
Sbjct: 740 KKTIISRNIVFDEEGEWDWNSNEEDY----NFFPH--------FEEDEPEP--------- 778
Query: 891 LSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVIS 950
E+P S+ P PT + TS+ + +SS + Q +
Sbjct: 779 -------TREEP----PSEEPTTPPT---------SPTSSQIEESSSERTPRFRSIQEL- 817
Query: 951 YHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKT 1010
Y + + + + EP + EA++ + WR AM+ EI+++++N+TW L P
Sbjct: 818 YEVTENQENLTLFCLFAECEPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHK 877
Query: 1011 PIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVST 1070
IG KWVY+ K G ++RYKARLV KGY Q GID+ + F+PVA++ T+R+++SL +
Sbjct: 878 TIGVKWVYKAKKNSKGEVERYKARLVAKGYIQRAGIDYDEVFAPVARLETVRLIISLAAQ 937
Query: 1071 KNWFLHQLDVDNAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFST 1129
W +HQ+DV +AFL+ L+EE+Y+ PQG + + ++V L+K+LYGLKQA R W +
Sbjct: 938 NKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKALYGLKQAPRAWNTR 997
Query: 1130 LCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKF 1189
+ + + F + +H LY+K A L YVDD++ GN+ + K + ++F
Sbjct: 998 IDKYFKEKDFIKCPYEHALYIKIQKEDILIACL-YVDDLIFTGNNPSMFEEFKKEMTKEF 1056
Query: 1190 RIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASS 1249
+ D+G ++LG+ + + GI + Q YA E+L + TPM+C KLS
Sbjct: 1057 EMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKE 1116
Query: 1250 GTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSP 1309
D ++++ L+G L YLT TRPDI Y V +S+++ PT H AA R+LRYIKG+
Sbjct: 1117 EGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTV 1176
Query: 1310 GCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCE 1369
GL Y S L +SDSD G VD RKS +G+ ++G + +W SKKQ S+CE
Sbjct: 1177 NFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVVLSTCE 1236
Query: 1370 AEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDC 1429
AEY A + VC WL LL++L + Q P +F DN+SA+ +A NP +H+R+KHI+
Sbjct: 1237 AEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRY 1296
Query: 1430 HIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLR 1473
H +RE +++ V L V + Q+ADIFTKPL R F K+R
Sbjct: 1297 HYIRECVSKKDVQLEYVKTHDQVADIFTKPLK----REDFIKMR 1336
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 566 bits (1458), Expect = e-161
Identities = 317/816 (38%), Positives = 464/816 (56%), Gaps = 72/816 (8%)
Query: 661 KSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVL-SQFYAEKGIIHHTSCVETPQQ 719
++K + +L+++FCA QF V+ +RSDNG EF+ + ++ E GI+H TSCV+TPQQ
Sbjct: 346 RAKQTLPSLIRNFCAMADRQFRKPVRSIRSDNGTEFMCHTSYFQEHGILHETSCVDTPQQ 405
Query: 720 NSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQ 779
N+ VERKH+HILNVAR LFQ + P P+P+L + P+++L +
Sbjct: 406 NARVERKHRHILNVARTCLFQGNFPT-----------------PSPVLKGKTPYEVLFGK 448
Query: 780 LPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRN 839
P L+ FG LC+A + KF R+++C+F+G+ T NT
Sbjct: 449 QPSYDMLRTFGCLCYAHIRPRDKDKFASRSRKCIFIGYPHETA-----TPNT-------- 495
Query: 840 VIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNS 899
H++I P D++ P P P N + +S
Sbjct: 496 ---HDSIDPTSTSSDENNTPPEPVTPQAEQPHSPSSISSPHIVHNKGSVHSRHLNEDHDS 552
Query: 900 EQPIVHRV---SQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDP 956
P + + RP+ P YL+DY +S P SS + +S +S + +
Sbjct: 553 SSPGLPELLGKGHRPKHPPVYLKDYVAHKVHSS---PHTSSPGLSDSNVSPTVSANHI-- 607
Query: 957 SYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKW 1016
AF+ I + E + + V + W AM EIEALE N+TW + D P K I KW
Sbjct: 608 ---AFMAAILDSNEQNHFKDDVLIKEWCDAMQKEIEALEANHTWDVTDLPHGKKAISSKW 664
Query: 1017 VYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLH 1076
VY++K+ DGT++R+KARLVV G Q EGIDF +TF+PVAKMTT+R+LL++ + K+W +
Sbjct: 665 VYKLKFNSDGTLERHKARLVVMGNHQKEGIDFKETFAPVAKMTTVRLLLAVAAAKDWDVF 724
Query: 1077 QLDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQG 1136
Q+DV NAFLH L++E SLYGLKQA R WF+ L AL+
Sbjct: 725 QMDVHNAFLHGDLEQE----------------------SLYGLKQAPRCWFAKLSTALRK 762
Query: 1137 LGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGE 1196
LGFTQS+ D++L+ + G+ L+YVDD ++ GN++ I K LH+ F +KDLG+
Sbjct: 763 LGFTQSYEDYSLF-SLNRDGTVIHFLVYVDDFIIVGNNLKAIDHFKEHLHKCFHMKDLGK 821
Query: 1197 AKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDI 1256
K+FLGL ++R G L+Q+KYAL++++++GLLG K + PM+ KL + S +
Sbjct: 822 LKYFLGLEVSRGADGFCLSQQKYALDIINEAGLLGYKPSAVPMELHHKLGSISSPVFDNP 881
Query: 1257 SSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYP 1316
+ YRRL+ R +YLT TRPD++Y V+ LSQF+ P + H A R++RY+KGSP G+
Sbjct: 882 AQYRRLVDRFIYLTITRPDLSYAVHILSQFMQTPLEAHWHATLRLVRYLKGSPDQGILLR 941
Query: 1317 AASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMA 1376
+ + LTA+ DSD C TR+S++ Y ++LG + +SW++KKQ T S SS EAEYRAMA
Sbjct: 942 SDRALSLTAYCDSDYNPCPRTRRSLSAYVLYLGDTPISWKTKKQDTVSSSSAEAEYRAMA 1001
Query: 1377 ATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKI 1436
T+ E++WL L+ L V T P+ +FCD+Q+A+HIA NP +HERTKHIE DCH VR+ +
Sbjct: 1002 YTLKEIKWLKALMTTLGVDHTQPILLFCDSQAAIHIAANPVFHERTKHIEKDCHQVRDAV 1061
Query: 1437 TQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
T ++ T + D+ TK L F + S L
Sbjct: 1062 TDKVIS----TPHISTTDLLTKALPRPTFERLLSTL 1093
Score = 82.4 bits (202), Expect = 2e-15
Identities = 87/356 (24%), Positives = 139/356 (38%), Gaps = 51/356 (14%)
Query: 258 LIAARENSNDGRSSQSNSGYQSNSGYQSGSGNYHSNSGRNRYSNKKCSYYGKMGHTVEDC 317
+I ++N + RS ++ + N ++ S + + ++ C++ + GH V DC
Sbjct: 38 VIRGQQNLDVARSKETTNSEAINFSVKTPSAPQVAAVYAPKPRDRSCTHCHRQGHDVTDC 97
Query: 318 YKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAARFGFTADQYHHLLL 377
+ HGFP + Y + + V + + S + ++E +
Sbjct: 98 FLVHGFPEWY------YEQKGGSRVSSDNREVVSRLENKPAKREGRSSKGNGRGRGRVNS 151
Query: 378 LLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQISTKNGNPLDTTWILDTGATDHICN 437
P N + Q T I+ P +S GN T I+D+GA+ H+
Sbjct: 152 ARAPLSSSNGSDQITQLISLLQAQRPKSTSERLS------GNTCLTDVIIDSGASHHMTG 205
Query: 438 TLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLV 497
S + P V+ P+G AT ++ LS S+ L +VLF+P+F+ LISV KL+
Sbjct: 206 DCSILVDVFDIIPSAVTKPDGKASCATKCVTLLLSSSYKLQDVLFVPDFDCTLISVSKLL 265
Query: 498 KSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLV 557
K FS +IG +Y ++ +S ST
Sbjct: 266 KQTG---PFSR-----------TLIGAGEVRERVYYFTGVLVASAHKTSSDST------- 304
Query: 558 CEFSPSVQNLWHYRLGHP--SFVKG-----QSIKDLFPYVQYTQDHVCEVCPIAKQ 606
S LWH RLGHP SF+ QS KDL + C+ C AKQ
Sbjct: 305 -----SSGALWHRRLGHPSTSFLLSLPECHQSSKDL------GKIDSCDTCSRAKQ 349
>At3g60170 putative protein
Length = 1339
Score = 559 bits (1441), Expect = e-159
Identities = 346/1085 (31%), Positives = 564/1085 (51%), Gaps = 75/1085 (6%)
Query: 422 DTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFI---LI 478
D W LD+G ++H+ + +FS + V L N S KGS+++ + + +
Sbjct: 297 DEVWFLDSGCSNHMTGSKEWFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQVIP 356
Query: 479 NVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDS 538
V ++P NL+S+ +L + + D C + + ++ T + ++ L
Sbjct: 357 EVYYVPELRNNLLSLGQL-QERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFL---- 411
Query: 539 LSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDL--------FPYV 590
L+ P NS+ C V + +LWH R GH + + +K L P +
Sbjct: 412 LASKPQKNSL--CLQTEEVMD---KENHLWHCRFGH---LNQEGLKLLAHKKMVIGLPIL 463
Query: 591 QYTQDHVCEVCPIAKQKRLKFPLSNS-TSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVD 649
+ T++ +C +C KQ R S S Q++H DI GP++ +S +G Y L+ +D
Sbjct: 464 KATKE-ICAICLTGKQHRESMSKKTSWKSSTQLQLVHSDICGPITPISHSGKRYILSFID 522
Query: 650 DYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV---LSQFYAEKG 706
D++R TWVY L K E K F A V + G + +R+D G EF +F G
Sbjct: 523 DFTRKTWVYFLHEKSEAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEFGEFCRSHG 582
Query: 707 IIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPI 766
I + TPQQN + ERK++ I+N R++L + +PK+FW+ A S+ + NR PT
Sbjct: 583 ISRQLTAAFTPQQNGVAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHIQNRSPTAA 642
Query: 767 LDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLV 826
++ P + + P + + +VFG + + R+K D ++K+CVFLG +K + +
Sbjct: 643 VEGMTPEEAWSGRKPVVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVSEESKAWRL 702
Query: 827 YDLNTRDISISRNVIFHEN-----------IFPYKLHRDDHECESSTFPQIPCLPSEPFD 875
YD + I IS++V+F E+ L D + E ++ P + P
Sbjct: 703 YDPVMKKIVISKDVVFDEDKSWDWDQADVEAKEVTLECGDEDDEKNSEVVEPIAVASP-- 760
Query: 876 YEYPIPPDNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPAN 935
+ DN V P+ + PS + P+ +V+ R R+ P ++ DY
Sbjct: 761 --NHVGSDNNVSSSPILA-PSSPAPSPVAAKVT-RERRPPGWMADY-------------- 802
Query: 936 SSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALE 995
+ +G +++ + ++ + T +P ++ +AVK + WR AM EIE++
Sbjct: 803 ETGEG-----------EEIEENLSVMLLMMMTEADPIQFDDAVKDKIWREAMEHEIESIV 851
Query: 996 RNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPV 1055
+NNTW L P TPIG KWVY+ K +DG +D+YKARLV KGY Q GID+ + F+PV
Sbjct: 852 KNNTWELTTLPKGFTPIGVKWVYKTKLNEDGEVDKYKARLVAKGYAQCYGIDYTEVFAPV 911
Query: 1056 AKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQK 1114
A++ T+R +L++ S NW + QLDV +AFLH +L EE+Y+ P+G + + +V L+K
Sbjct: 912 ARLDTVRTILAISSQFNWEIFQLDVKSAFLHGELKEEVYVRQPEGFIREGEEEKVYKLRK 971
Query: 1115 SLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGND 1174
+LYGLKQA R W+S + F + ++HTL+ K+ G+ + LYVDD++ G+D
Sbjct: 972 ALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFT-KTRVGNILIVSLYVDDLIFTGSD 1030
Query: 1175 MDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKS 1234
K S+ +F + DLG+ K FLG+ + +S GI + QR+YA E+L+ G+ +
Sbjct: 1031 KAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQRRYAREVLARFGMDESNA 1090
Query: 1235 ATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMH 1294
P+ KL+ D + +++L+G L+YLT TRPD+ Y V +S+F+S P H
Sbjct: 1091 VKNPIVPGTKLTKDENGEKVDETMFKQLVGSLMYLTVTRPDLMYGVCLISRFMSNPRMSH 1150
Query: 1295 EAAAHRVLRYIKGSPGCGLFYPAAS--STVLTAFSDSDGAGCVDTRKSITGYCMFLGSSL 1352
AA R+LRY+KG+ G+FY S L AF+DSD AG ++ R+S +G+ + S
Sbjct: 1151 WLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYAGDLNDRRSTSGFVFLMASGA 1210
Query: 1353 VSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHI 1412
+ W SKKQ + S+ EAEY A A C+ WL +L+ L + S + CDN S + +
Sbjct: 1211 ICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLGAEEKSATVINCDNSSTIQL 1270
Query: 1413 AHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
+ +P H ++KHIE+ H +R+ + +V L + Q+ADIFTKPL F + + L
Sbjct: 1271 SKHPVLHGKSKHIEVRFHYLRDLVNGDVVKLEYCPTEDQVADIFTKPLKLEQFEKLRALL 1330
Query: 1473 RLYDI 1477
+ ++
Sbjct: 1331 GMVNM 1335
>At4g23160 putative protein
Length = 1240
Score = 544 bits (1401), Expect = e-154
Identities = 288/585 (49%), Positives = 384/585 (65%), Gaps = 26/585 (4%)
Query: 886 VQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPL 945
+ I P S+ + +P VH +R RK P YLQDY+C A+ T+ H +
Sbjct: 14 IDIMP-SANIQNDVPEPSVHTSHRRTRK-PAYLQDYYCHSVASLTI-----------HDI 60
Query: 946 SQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDK 1005
SQ +SY K+ P YH+F++ I EP+ Y+EA + W AM+ EI A+E +TW +
Sbjct: 61 SQFLSYEKVSPLYHSFLVCIAKAKEPSTYNEAKEFLVWCGAMDDEIGAMETTHTWEICTL 120
Query: 1006 PPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLL 1065
PP+K PIGCKWVY+IKY DGTI+RYKARLV KGYTQ EGIDF++TFSPV K+T+++++L
Sbjct: 121 PPNKKPIGCKWVYKIKYNSDGTIERYKARLVAKGYTQQEGIDFIETFSPVCKLTSVKLIL 180
Query: 1066 SLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGM-----NSDKPNQVCLLQKSLYGLK 1120
++ + N+ LHQLD+ NAFL+ LDEEIYM LP G +S PN VC L+KS+YGLK
Sbjct: 181 AISAIYNFTLHQLDISNAFLNGDLDEEIYMKLPPGYAARQGDSLPPNAVCYLKKSIYGLK 240
Query: 1121 QASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKL 1180
QASRQWF L G GF QS +DHT ++K +AT F +L+YVDD+++ N+ +
Sbjct: 241 QASRQWFLKFSVTLIGFGFVQSHSDHTYFLKITAT-LFLCVLVYVDDIIICSNNDAAVDE 299
Query: 1181 VKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMD 1240
+K L F+++DLG K+FLGL IARS GI + QRKYAL+LL ++GLLG K ++ PMD
Sbjct: 300 LKSQLKSCFKLRDLGPLKYFLGLEIARSAAGINICQRKYALDLLDETGLLGCKPSSVPMD 359
Query: 1241 CSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHR 1300
S SA SG D +YRRLIGRL+YL TR DI++ VN+LSQF AP H+ A +
Sbjct: 360 PSVTFSAHSGGDFVDAKAYRRLIGRLMYLQITRLDISFAVNKLSQFSEAPRLAHQQAVMK 419
Query: 1301 VLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQ 1360
+L YIKG+ G GLFY + + L FSD+ C DTR+S GYCMFLG+SL+SW+SKKQ
Sbjct: 420 ILHYIKGTVGQGLFYSSQAEMQLQVFSDASFQSCKDTRRSTNGYCMFLGTSLISWKSKKQ 479
Query: 1361 TTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHE 1420
S+SS EAEYRA++ E+ WL ++LQ+ + P +FCDN +A+HIA N +HE
Sbjct: 480 QVVSKSSAEAEYRALSFATDEMMWLAQFFRELQLPLSKPTLLFCDNTAAIHIATNAVFHE 539
Query: 1421 RTKHIELDCHIVREKITQGLVHLLPVTSSLQL---ADIFTKPLSP 1462
RTKHIE DCH VRE+ V+ ++ S Q D FT+ LSP
Sbjct: 540 RTKHIESDCHSVRER----SVYQATLSYSFQAYDEQDGFTEYLSP 580
>At1g37110
Length = 1356
Score = 544 bits (1401), Expect = e-154
Identities = 357/1090 (32%), Positives = 561/1090 (50%), Gaps = 57/1090 (5%)
Query: 411 SQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQ 470
S+ + N + WILD+G T H+ + +F S++ + L + + + +G+I+
Sbjct: 294 SEALSVNEQMVKDLWILDSGCTSHMTSRRDWFISFQEKGNTTILLGDDHSVESQGQGTIR 353
Query: 471 LSPSF----ILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTAR 526
+ IL NV ++P+ NLIS L K L +R + +N + G+
Sbjct: 354 IDTHGGTIKILENVKYVPHLRRNLISTGTLDK-LGYRHEGGEGKVRYFKNNKTALRGSLS 412
Query: 527 AVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF--VKGQSIK 584
LY+L+ ST S+ E LWH RLGH S +K + K
Sbjct: 413 --NGLYVLDG------------STVMSELCNAETDKVKTALWHSRLGHMSMNNLKVLAGK 458
Query: 585 DLFPYVQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVL-SLNGFSY 643
L + + CE C + K K++ F + TS+ +H D+WG +V S++G Y
Sbjct: 459 GLIDRKEINELEFCEHCVMGKSKKVSFNVGKHTSEDALSYVHADLWGSPNVTPSISGKQY 518
Query: 644 FLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYA 703
FL+I+DD +R W+Y LKSKDE ++ + V NQ VK +R+DNG EF S+F +
Sbjct: 519 FLSIIDDKTRKVWLYFLKSKDETFDKFCEWKSLVENQVNKKVKCLRTDNGLEFCNSRFDS 578
Query: 704 ---EKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLIN 760
E GI H +C TPQQN + ER ++ I+ R LL ++ + ++FWA A + +LIN
Sbjct: 579 YCKEHGIERHRTCTYTPQQNGVAERMNRTIMEKVRCLLNKSGVEEVFWAEAAATAAYLIN 638
Query: 761 RLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSG 820
R P +++ P ++ + P L+ FGS+ + + K RA + FLG+ +G
Sbjct: 639 RSPASAINHNVPEEMWLNRKPGYKHLRKFGSIAYVH---QDQGKLKPRALKGFFLGYPAG 695
Query: 821 TKGYLVYDLNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPI 880
TKGY V+ L ISRNV+F E++ L + + ++ + E +
Sbjct: 696 TKGYKVWLLEEEKCVISRNVVFQESVVYRDLKVKEDDTDNLNQKETTSSEVEQNKFAEAS 755
Query: 881 PPDNPVQIQ----PLSSGP-SQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPAN 935
+Q+Q P++ G S +SE+ + + + + T L Y LA N
Sbjct: 756 GSGGVIQLQSDSEPITEGEQSSDSEEEVEYSEKTQETPKRTGLTTYK--LARDRVRRNIN 813
Query: 936 SSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKH---ECWRVAMN*EIE 992
++ T ++ ++ EP Y EA++ E W +A + E++
Sbjct: 814 PPTRFTEES----------SVTFALVVVENCIVQEPQSYQEAMESQDCEKWDMATHDEMD 863
Query: 993 ALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTID-RYKARLVVKGYTQIEGIDFMDT 1051
+L +N TW LVDKP D+ IGC+W++++K G R+KARLV KGYTQ EG+D+ +
Sbjct: 864 SLMKNGTWDLVDKPKDRKIIGCRWLFKLKSGIPGVEPTRFKARLVAKGYTQREGVDYQEI 923
Query: 1052 FSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKP-NQVC 1110
F+PV K ++R+L+SLV K+ L Q+DV FLH L+EE+YM P+G SD N+VC
Sbjct: 924 FAPVVKHVSIRILMSLVVDKDLELEQMDVKTTFLHGDLEEELYMEQPEGFVSDSSENKVC 983
Query: 1111 LLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLL 1170
L+KSLYGLKQ+ RQW + + F +S D +YVK + F LLLYVDD+L+
Sbjct: 984 RLKKSLYGLKQSPRQWNKRFDRFMSSQQFIRSEHDACVYVKHVSEHDFIYLLLYVDDMLI 1043
Query: 1171 AGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIIL--NQRKYALELLSDSG 1228
AG EI VK L +F +KD+G A LG+ I R +KG +L +Q Y ++L
Sbjct: 1044 AGASKAEINRVKEQLSTEFEMKDMGGASRILGIDIYRDRKGGVLKLSQEIYIRKVLDRFN 1103
Query: 1229 LLGGKSATTPMDCSQKLSA---SSGTPLSDISSYRRLIGRLLY-LTTTRPDIAYVVNQLS 1284
+ G K P+ KL+A +D+ Y +G ++Y + TRPD+AY + +S
Sbjct: 1104 MSGAKMTNAPVGAHFKLAAVREEDECVDTDVVPYSSAVGSIMYAMLGTRPDLAYAICLIS 1163
Query: 1285 QFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGY 1344
+++S P MH A V+RY+KG+ L + +T + DS+ A +D R+SI+GY
Sbjct: 1164 RYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTKEKDFTVTGYCDSNYAADLDRRRSISGY 1223
Query: 1345 CMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFC 1404
+G + VSW++ Q + S+ EAEY A+A E W+ LLQD+ + Q V ++C
Sbjct: 1224 VFTIGGNTVSWKASLQPVVAMSTTEAEYIALAEAAKEAMWIKGLLQDMGMQQ-DKVKIWC 1282
Query: 1405 DNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAP 1464
D+QSA+ ++ N YHERTKHI++ + +R+ + G V +L + +S D TK +
Sbjct: 1283 DSQSAICLSKNSVYHERTKHIDVRFNYIRDVVESGDVDVLKIHTSRNPVDALTKCIPVNK 1342
Query: 1465 FRHIFSKLRL 1474
F+ L+L
Sbjct: 1343 FKSALGVLKL 1352
>At1g36620 hypothetical protein
Length = 1152
Score = 537 bits (1383), Expect = e-152
Identities = 329/920 (35%), Positives = 486/920 (52%), Gaps = 84/920 (9%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAA 363
C++ G+ H+ + C+K HG P + KY + S+ + +G+ +A
Sbjct: 275 CTHCGRSNHSADTCFKLHGVPEWY---TEKYGDTSSGRGRGRSSTPRGRGRGHGNSYKA- 330
Query: 364 RFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQISTK------- 416
N+ + H +S S +PG + + S I
Sbjct: 331 ---------------------NNAQTSHPSSSASEFSDIPGVSKEAWSAIRNLLKQDTAT 369
Query: 417 -----NGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQL 471
+G +++D+GA+ H+ L + + V LPN AT KG++ L
Sbjct: 370 SSEKLSGKTNCVDFLIDSGASHHMTGFLDLLTEIYEIPHSVVVLPNAKHTIATKKGTLIL 429
Query: 472 SPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSL 531
+ L +VLF+P+ LISV +L++ L F+D C+IQD + +IG +
Sbjct: 430 GANMKLTHVLFVPDLSCTLISVARLLRELHCFAIFTDKVCVIQDRTSKMLIGVGTESNGV 489
Query: 532 YILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF-VKGQSIKDLFPYV 590
Y L + S N+V + LWH RLGHPS V + L +
Sbjct: 490 YHLQRAEV----------VATSANVVKWKTNKA--LWHMRLGHPSSKVLSSVLPSLEDFD 537
Query: 591 QYTQD--HVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIV 648
+ D +C+VC AKQ R F S + ++ F IH D+WGP S G YFLTIV
Sbjct: 538 SCSSDLKTICDVCVRAKQTRASFSESFNKAEECFSFIHYDVWGPYKHASSCGAHYFLTIV 597
Query: 649 DDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV-LSQFYAEKGI 707
DD+SR W++L+ +K EV +L++ F A + QF VK VRS+NG EF+ L ++AE+GI
Sbjct: 598 DDHSRAVWIHLMLAKSEVASLLQQFIAMASRQFNKQVKTVRSNNGTEFMSLKSYFAERGI 657
Query: 708 IHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPIL 767
+H SCV T QQN VERKH+HILNVAR+LLFQA LP FW +++ + +LINR PTPIL
Sbjct: 658 VHQISCVYTHQQNGRVERKHRHILNVARSLLFQAELPISFWEESVLTAAYLINRTPTPIL 717
Query: 768 DNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVY 827
D + P+++L+ Q P L+VFGSLCFA KF R ++C+F+G+ G KG+ +Y
Sbjct: 718 DGKTPYKILYSQPPSYASLRVFGSLCFARKHTGRLDKFQERGRKCIFVGYPHGQKGWRIY 777
Query: 828 DLNTRDISISRNVIFHENIFPY--KLHRDDHECESSTFPQIPCLPSEPFDYEY------- 878
D+ ++ +SR+V+F E+IFP+ K ++D ++ P P P+D E+
Sbjct: 778 DIESQIFFVSRDVVFQEDIFPFADKKNKDTFSSPAAVIPS----PILPYDDEFLDIYQIG 833
Query: 879 ------PIPP-----DNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAA 927
P+P D+P +++ P+ S P+ R R R++ L+DY T +A
Sbjct: 834 DVPATNPLPAIIDVNDSPPSSPIITATPAAASPPPL--RRGLRQRQENVRLKDYQ-TYSA 890
Query: 928 TSTVVPANSSSKGTS-HPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVA 986
S + GT +P++ +S PS F+ I+ P Y++A++ + WR A
Sbjct: 891 QCESTQTLSDNIGTCIYPMANYVSGEIFSPSNQHFLAAISMVDPPQTYNQAIREKEWRNA 950
Query: 987 MN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGI 1046
+ E++ALE TW + P IG KWV+RIKY +GT++RYKARLV G Q EGI
Sbjct: 951 VFFEVDALEDQGTWDITKLPQGVKAIGSKWVFRIKYNSNGTVERYKARLVALGNHQKEGI 1010
Query: 1047 DFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKP 1106
DF TF+PV KM T+R+LL + + K+W LHQ+DV NAFLH L E+IYM P G + P
Sbjct: 1011 DFTKTFAPVVKMQTVRLLLDVAAAKDWELHQMDVHNAFLHGDLKEDIYMKPPPGFKTTDP 1070
Query: 1107 NQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGS-FTALLLYV 1165
+ VC L+KS+YGLKQA R WF L +L GFTQS D++L+ S GS +++YV
Sbjct: 1071 SLVCKLKKSIYGLKQAPRCWFEKLSTSLLKFGFTQSKKDYSLFT--SIRGSKVLHVIVYV 1128
Query: 1166 DDVLLAGNDMDEIKLVKHSL 1185
DDV++ G + E K +L
Sbjct: 1129 DDVVICGKAVRENNTSKLAL 1148
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 534 bits (1376), Expect = e-151
Identities = 391/1259 (31%), Positives = 600/1259 (47%), Gaps = 136/1259 (10%)
Query: 258 LIAARENSNDGRSSQSNSGYQSNS---GYQSGSGNYHSNSGRNRYSNKK------CSYYG 308
++AAR+ + SQ+N G G N N G+NR +K C G
Sbjct: 168 MVAARDKERE--LSQNNRPVVEGHFARGRPDGKNNNQGNKGKNRSRSKSADGKRVCWICG 225
Query: 309 KMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAARFGFT 368
K GH + CYK ++ K Q+ + G
Sbjct: 226 KEGHFKKQCYK---------------------WIERNKSKQQGSDNG------------- 251
Query: 369 ADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQISTKNGNPLDTTWILD 428
++S ++ T + N + +L ++ + + W+LD
Sbjct: 252 ----------------ESSLAKSTEAFNPAMVLLATDETLVVT-------DSIANEWVLD 288
Query: 429 TGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPS----FILINVLFLP 484
TG + H+ +F +K ++ V + N GSI++ S IL +V ++P
Sbjct: 289 TGCSFHMTPRKDWFKDFKELSSGYVKMGNDTYSPVKGIGSIKIRNSDGSQVILTDVRYMP 348
Query: 485 NFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPS 544
N NLIS+ L F D +++ C I + +LYIL D ++
Sbjct: 349 NMTRNLISLGTLEDR---GCWFKSQDGILKIVKGCSTILKGQKRDTLYIL--DGVTEEGE 403
Query: 545 CNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSI---KDLFPYVQYTQDHVCEVC 601
+S + K + LWH RLGH S KG I K + CE C
Sbjct: 404 SHSSAEVKDETA----------LWHSRLGHMS-QKGMEILVKKGCLRREVIKELEFCEDC 452
Query: 602 PIAKQKRLKFPLSNSTSDCIFQMIHVDIWG-PVSVLSLNGFSYFLTIVDDYSRYTWVYLL 660
KQ R+ F + + +H D+WG P + SL YF++ VDDYSR W+Y L
Sbjct: 453 VYGKQHRVSFAPAQHVTKEKLAYVHSDLWGSPHNPASLGNSQYFISFVDDYSRKVWIYFL 512
Query: 661 KSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV---LSQFYAEKGIIHHTSCVETP 717
+ KDE ++ V NQ VK +R+DNG E+ +F E+GI+ H +C TP
Sbjct: 513 RKKDEAFEKFVEWKKMVENQSDRKVKKLRTDNGLEYCNHYFEKFCKEEGIVRHKTCAYTP 572
Query: 718 QQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLH 777
QQN I ER ++ I++ R++L ++ + K FWA A +++LINR P+ ++ P +
Sbjct: 573 QQNGIAERLNRTIMDKVRSMLSRSGMEKKFWAEAASTAVYLINRSPSTAINFDLPEEKWT 632
Query: 778 KQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISIS 837
LPD++ L+ FG L + + + + K + R+K+ +F + G KGY V+ L + IS
Sbjct: 633 GALPDLSSLRKFGCLAY---IHADQGKLNPRSKKGIFTSYPEGVKGYKVWVLEDKKCVIS 689
Query: 838 RNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQ 897
RNVIF E + L D S + + + + PD Q G +Q
Sbjct: 690 RNVIFREQVMFKDLKGDSQNTISESDLE-----------DLRVNPDMNDQEFTDQGGATQ 738
Query: 898 NSEQPIVHRVSQRP-RKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDP 956
++ P S P PT+ QD S V S+ + + + I K +P
Sbjct: 739 DNSNPSEATTSHNPVLNSPTHSQDEESE-EEDSDAVEDLSTYQLVRDRVRRTI---KANP 794
Query: 957 SYH-------AFIMNITTTVEPTRYSEAVKH---ECWRVAMN*EIEALERNNTWLLVDKP 1006
Y+ A+ EP Y EA+ E W AM E+ ++ +N+TW LV KP
Sbjct: 795 KYNESNMVGFAYYSEDDGKPEPKSYQEALLDPDWEKWNAAMKEEMVSMSKNHTWDLVTKP 854
Query: 1007 PDKTPIGCKWVYRIKYKQDGT-IDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLL 1065
IGC+WV+ K G R+ ARLV KG+TQ EG+D+ + FSPV K ++R LL
Sbjct: 855 EKVKLIGCRWVFTRKAGIPGVEAPRFIARLVAKGFTQKEGVDYNEIFSPVVKHVSIRYLL 914
Query: 1066 SLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-PNQVCLLQKSLYGLKQASR 1124
S+V N L Q+DV AFLH L+EEIYM+ P+G + N+VCLL++SLYGLKQ+ R
Sbjct: 915 SMVVHYNMELQQMDVKTAFLHGFLEEEIYMAQPEGFEIKRGSNKVCLLKRSLYGLKQSPR 974
Query: 1125 QWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHS 1184
QW + ++G+ +T+S D +Y KK ++ LLLYVDD+L+A + E+ +K
Sbjct: 975 QWNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYIYLLLYVDDMLIASANKSEVNELKQL 1034
Query: 1185 LHQKFRIKDLGEAKFFLGLAIARSQKG--IILNQRKYALELLSDSGLLGGKSATTPMDCS 1242
L ++F +KDLG+AK LG+ I+R + + L+Q Y ++L + K +TP+
Sbjct: 1035 LSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYVKKVLRSFQMDNAKPVSTPLGIH 1094
Query: 1243 QKLSASSGTPLSD------ISSYRRLIGRLLY-LTTTRPDIAYVVNQLSQFLSAPTDMHE 1295
KL A++ + I Y IG ++Y + TRPD+AY + +S+F+S P H
Sbjct: 1095 FKLKAATDKEYEEQFERMKIVPYANTIGSIMYSMIGTRPDLAYSLGVISRFMSKPLKDHW 1154
Query: 1296 AAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSW 1355
A VLRY++G+ L + +L + DSD DTR+SITGY +G + +SW
Sbjct: 1155 QAVKWVLRYMRGTEKKKLCFRKQEDFLLRGYCDSDYGSNFDTRRSITGYVFTVGGNTISW 1214
Query: 1356 RSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHN 1415
+SK Q + SS EAEY A+ V E WL +L SQ V + D+QSA+ +A N
Sbjct: 1215 KSKLQKVVAISSTEAEYMALTEAVKEALWLKGFAAELGHSQ-DYVEVHSDSQSAITLAKN 1273
Query: 1416 PSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRL 1474
+HERTKHI++ H +R+ I GL+ ++ + + A+IFTK + A F + LR+
Sbjct: 1274 SVHHERTKHIDIRLHFIRDIICAGLIKVVKIATECNPANIFTKTVPLAKFEGALNMLRV 1332
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,917,630
Number of Sequences: 26719
Number of extensions: 1415960
Number of successful extensions: 5613
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 31
Number of HSP's that attempted gapping in prelim test: 4762
Number of HSP's gapped (non-prelim): 356
length of query: 1481
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1369
effective length of database: 8,326,068
effective search space: 11398387092
effective search space used: 11398387092
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0133.5


