

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.1
(267 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
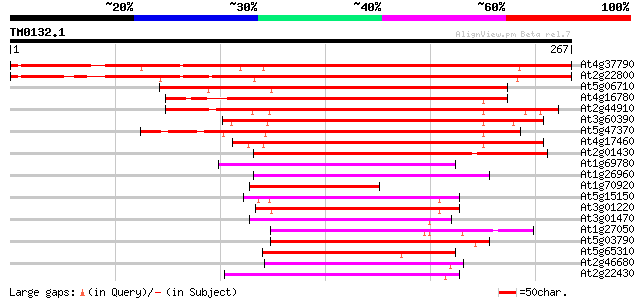
Score E
Sequences producing significant alignments: (bits) Value
At4g37790 homeobox protein (HAT22) 295 1e-80
At2g22800 homeobox protein 285 2e-77
At5g06710 homeobox protein 204 5e-53
At4g16780 DNA-binding homeotic protein Athb-2 198 2e-51
At2g44910 homeodomain transcription factor (ATHB-4) 194 4e-50
At3g60390 homeobox-leucine zipper protein HAT3 189 1e-48
At5g47370 homeobox-leucine zipper protein-like 184 3e-47
At4g17460 homeobox-leucine zipper protein HAT1 (hd-zip protein 1) 179 2e-45
At2g01430 putative homeodomain transcription factor 150 5e-37
At1g69780 homeobox gene 13 protein 85 5e-17
At1g26960 putative DNA-binding protein 84 1e-16
At1g70920 hypothetical protein 82 2e-16
At5g15150 homeobox protein 79 3e-15
At3g01220 putative homeobox-leucine zipper protein, HAT7 77 1e-14
At3g01470 homeobox protein (HAT5) 76 2e-14
At1g27050 unknown protein 75 5e-14
At5g03790 homeodomain -like protein 72 3e-13
At5g65310 homeobox-leucine zipper protein ATHB-5 (HD-zip protein... 71 5e-13
At2g46680 homeodomain transcription factor (ATHB-7) 71 5e-13
At2g22430 homeodomain transcription factor (ATHB-6) 70 2e-12
>At4g37790 homeobox protein (HAT22)
Length = 278
Score = 295 bits (756), Expect = 1e-80
Identities = 172/286 (60%), Positives = 199/286 (69%), Gaps = 27/286 (9%)
Query: 1 MGLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGN 60
MGLD D+ N GL L LGLS T N + +++T + + +PSLTL LSG
Sbjct: 1 MGLD-DSCNTGLVLGLGLSPTPNNYNHAIKKSSSTVDHR------FIRLDPSLTLSLSGE 53
Query: 61 N---KVYCEDPLELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEER----LSSRV 113
+ K ++ RQTS HS + SSFS+ RV + + E EAEE + SRV
Sbjct: 54 SYKIKTGAGAGDQICRQTSSHSGI-SSFSSGRVKREREISGGDGEEEAEETTERVVCSRV 112
Query: 114 SDEDDD--GTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQ 171
SD+ DD G +ARKKLRLTK+QSALLE++FK HSTLNPKQKQALARQLNLR RQVEVWFQ
Sbjct: 113 SDDHDDEEGVSARKKLRLTKQQSALLEDNFKLHSTLNPKQKQALARQLNLRPRQVEVWFQ 172
Query: 172 NRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMC 231
NRRARTKLKQTEVDCEFLKKCCETLTDENRRL+KELQ+LKALKL+QP YM M AATLTMC
Sbjct: 173 NRRARTKLKQTEVDCEFLKKCCETLTDENRRLQKELQDLKALKLSQPFYMHMPAATLTMC 232
Query: 232 PSCERLGDGG----------SNIKSPFTITPKPHFFNPFTHPSAAC 267
PSCERLG GG K F+I KP F+NPFT+PSAAC
Sbjct: 233 PSCERLGGGGVGGDTTAVDEETAKGAFSIVTKPRFYNPFTNPSAAC 278
>At2g22800 homeobox protein
Length = 274
Score = 285 bits (729), Expect = 2e-77
Identities = 174/289 (60%), Positives = 191/289 (65%), Gaps = 37/289 (12%)
Query: 1 MGLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGN 60
MG D D N GL L LG S N + +T SS EPSLTL LSG+
Sbjct: 1 MGFD-DTCNTGLVLGLGPSPIPNNYN----STIRQSS--------VYKLEPSLTLCLSGD 47
Query: 61 NKVYCEDPLE-LSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSD--ED 117
V + L RQTS HS V SSFS+ RVVKRER D E E EE +SD ED
Sbjct: 48 PSVTVVTGADQLCRQTSSHSGV-SSFSSGRVVKRER-DGGEESPEEEEMTERVISDYHED 105
Query: 118 DDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRART 177
++G +ARKKLRLTK+QSALLEESFK HSTLNPKQKQ LARQLNLR RQVEVWFQNRRART
Sbjct: 106 EEGISARKKLRLTKQQSALLEESFKDHSTLNPKQKQVLARQLNLRPRQVEVWFQNRRART 165
Query: 178 KLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERL 237
KLKQTEVDCEFLKKCCETL DEN RL+KE+QELK LKL QP YM M A+TLT CPSCER+
Sbjct: 166 KLKQTEVDCEFLKKCCETLADENIRLQKEIQELKTLKLTQPFYMHMPASTLTKCPSCERI 225
Query: 238 GDG-------------------GSNIKSPFTITPKPHFFNPFTHPSAAC 267
G G GS K F+I+ KPHFFNPFT+PSAAC
Sbjct: 226 GGGGGGNGGGGGGSGATAVIVDGSTAKGAFSISSKPHFFNPFTNPSAAC 274
>At5g06710 homeobox protein
Length = 336
Score = 204 bits (518), Expect = 5e-53
Identities = 107/171 (62%), Positives = 129/171 (74%), Gaps = 5/171 (2%)
Query: 72 SRQTSPHSDVVSSFSTARVVKR---ERVDLSCEEIEAEERLSSRVSDEDDDGTNA--RKK 126
S SP V SSF +K ER + + ER +SR S+ED+D N RKK
Sbjct: 132 SMSVSPPDSVTSSFQLDFGIKSYGYERRSNKRDIDDEVERSASRASNEDNDDENGSTRKK 191
Query: 127 LRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDC 186
LRL+K+QSA LE+SFK+HSTLNPKQK ALA+QLNLR RQVEVWFQNRRARTKLKQTEVDC
Sbjct: 192 LRLSKDQSAFLEDSFKEHSTLNPKQKIALAKQLNLRPRQVEVWFQNRRARTKLKQTEVDC 251
Query: 187 EFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERL 237
E+LK+CCE+LT+ENRRL+KE++EL+ LK + P YM + A TLTMCPSCER+
Sbjct: 252 EYLKRCCESLTEENRRLQKEVKELRTLKTSTPFYMQLPATTLTMCPSCERV 302
>At4g16780 DNA-binding homeotic protein Athb-2
Length = 284
Score = 198 bits (504), Expect = 2e-51
Identities = 106/164 (64%), Positives = 127/164 (76%), Gaps = 12/164 (7%)
Query: 75 TSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQS 134
+SP+S V SS T + +RE E + SR +D+DG N+RKKLRL+K+QS
Sbjct: 90 SSPNSTVSSS--TGKRSERE---------EDTDPQGSRGISDDEDGDNSRKKLRLSKDQS 138
Query: 135 ALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCE 194
A+LEE+FK HSTLNPKQKQALA+QL LRARQVEVWFQNRRARTKLKQTEVDCEFL++CCE
Sbjct: 139 AILEETFKDHSTLNPKQKQALAKQLGLRARQVEVWFQNRRARTKLKQTEVDCEFLRRCCE 198
Query: 195 TLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERL 237
LT+ENRRL+KE+ EL+ALKL+ YM MS TLTMCPSCE +
Sbjct: 199 NLTEENRRLQKEVTELRALKLSPQFYMHMSPPTTLTMCPSCEHV 242
>At2g44910 homeodomain transcription factor (ATHB-4)
Length = 318
Score = 194 bits (493), Expect = 4e-50
Identities = 112/203 (55%), Positives = 140/203 (68%), Gaps = 19/203 (9%)
Query: 75 TSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVS----DEDDDGTN---ARKKL 127
+SP+S V S R + R +E EAE SR +D+DG N +RKKL
Sbjct: 109 SSPNSAVSSLSGNKRDLAVAR---GGDENEAERASCSRGGGSGGSDDEDGGNGDGSRKKL 165
Query: 128 RLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCE 187
RL+K+Q+ +LEE+FK+HSTLNPKQK ALA+QLNLRARQVEVWFQNRRARTKLKQTEVDCE
Sbjct: 166 RLSKDQALVLEETFKEHSTLNPKQKLALAKQLNLRARQVEVWFQNRRARTKLKQTEVDCE 225
Query: 188 FLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNI-K 245
+LK+CC+ LT+ENRRL+KE+ EL+ALKL+ LYM M+ TLTMCPSCER+ + +
Sbjct: 226 YLKRCCDNLTEENRRLQKEVSELRALKLSPHLYMHMTPPTTLTMCPSCERVSSSAATVTA 285
Query: 246 SPFTIT-------PKPHFFNPFT 261
+P T T P P P+T
Sbjct: 286 APSTTTTPTVVGRPSPQRLTPWT 308
>At3g60390 homeobox-leucine zipper protein HAT3
Length = 315
Score = 189 bits (481), Expect = 1e-48
Identities = 102/160 (63%), Positives = 120/160 (74%), Gaps = 7/160 (4%)
Query: 102 EIE-AEERLSSRVSDEDDDGT---NARKKLRLTKEQSALLEESFKQHSTLNPKQKQALAR 157
EIE A L DED G ++RKKLRL+KEQ+ +LEE+FK+HSTLNPKQK ALA+
Sbjct: 135 EIERASCSLGGGSDDEDGSGNGDDSSRKKLRLSKEQALVLEETFKEHSTLNPKQKMALAK 194
Query: 158 QLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQ 217
QLNLR RQVEVWFQNRRARTKLKQTEVDCE+LK+CCE LTDENRRL+KE+ EL+ALKL+
Sbjct: 195 QLNLRTRQVEVWFQNRRARTKLKQTEVDCEYLKRCCENLTDENRRLQKEVSELRALKLSP 254
Query: 218 PLYMPMS-AATLTMCPSCERLG--DGGSNIKSPFTITPKP 254
LYM M TLTMCPSCER+ S++ P + P
Sbjct: 255 HLYMHMKPPTTLTMCPSCERVAVTSSSSSVAPPVMNSSSP 294
>At5g47370 homeobox-leucine zipper protein-like
Length = 283
Score = 184 bits (468), Expect = 3e-47
Identities = 106/192 (55%), Positives = 133/192 (69%), Gaps = 17/192 (8%)
Query: 63 VYCEDPLELSRQTSPHSDVVSSFSTARVVKRERVDLSC---------EEIEAEERLSSRV 113
V CE+ +S SP+S + S+ S R ER +S +EI + S
Sbjct: 64 VNCEEDTGVS---SPNSTISSTISGKR---SEREGISGTGVGSGDDHDEITPDRGYSRGT 117
Query: 114 SDEDDDG-TNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQN 172
SDE++DG +RKKLRL+K+QSA LEE+FK+H+TLNPKQK ALA++LNL ARQVEVWFQN
Sbjct: 118 SDEEEDGGETSRKKLRLSKDQSAFLEETFKEHNTLNPKQKLALAKKLNLTARQVEVWFQN 177
Query: 173 RRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMC 231
RRARTKLKQTEVDCE+LK+C E LT+ENRRL+KE EL+ LKL+ Y M+ TL MC
Sbjct: 178 RRARTKLKQTEVDCEYLKRCVEKLTEENRRLQKEAMELRTLKLSPQFYGQMTPPTTLIMC 237
Query: 232 PSCERLGDGGSN 243
PSCER+G S+
Sbjct: 238 PSCERVGGPSSS 249
>At4g17460 homeobox-leucine zipper protein HAT1 (hd-zip protein 1)
Length = 282
Score = 179 bits (453), Expect = 2e-45
Identities = 95/152 (62%), Positives = 119/152 (77%), Gaps = 4/152 (2%)
Query: 107 ERLSSR-VSDEDDD--GTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRA 163
+R SSR SDE++D G RKKLRL+K+QSA+LE++FK+H+TLNPKQK ALA++L L A
Sbjct: 114 DRSSSRGTSDEEEDYGGETCRKKLRLSKDQSAVLEDTFKEHNTLNPKQKLALAKKLGLTA 173
Query: 164 RQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPM 223
RQVEVWFQNRRARTKLKQTEVDCE+LK+C E LT+ENRRL+KE EL+ALKL+ LY M
Sbjct: 174 RQVEVWFQNRRARTKLKQTEVDCEYLKRCVEKLTEENRRLEKEAAELRALKLSPRLYGQM 233
Query: 224 S-AATLTMCPSCERLGDGGSNIKSPFTITPKP 254
S TL MCPSCER+ S+ + +++ P
Sbjct: 234 SPPTTLLMCPSCERVAGPSSSNHNQRSVSLSP 265
>At2g01430 putative homeodomain transcription factor
Length = 162
Score = 150 bits (380), Expect = 5e-37
Identities = 76/140 (54%), Positives = 102/140 (72%), Gaps = 2/140 (1%)
Query: 117 DDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRAR 176
DD RKKLRLT+EQS LLE+SF+Q+ TLNPKQK+ LA+ L LR RQ+EVWFQNRRAR
Sbjct: 18 DDGSAPPRKKLRLTREQSRLLEDSFRQNHTLNPKQKEVLAKHLMLRPRQIEVWFQNRRAR 77
Query: 177 TKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCER 236
+KLKQTE++CE+LK+ +LT+EN RL +E++EL+A+K+ SA++LTMCP CER
Sbjct: 78 SKLKQTEMECEYLKRWFGSLTEENHRLHREVEELRAMKVGPTTV--NSASSLTMCPRCER 135
Query: 237 LGDGGSNIKSPFTITPKPHF 256
+ S ++ + K F
Sbjct: 136 VTPAASPSRAVVPVPAKKTF 155
>At1g69780 homeobox gene 13 protein
Length = 294
Score = 84.7 bits (208), Expect = 5e-17
Identities = 46/113 (40%), Positives = 65/113 (56%)
Query: 100 CEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQL 159
C ++E ++ DD KK RL EQ LE++F+ + L P++K LAR L
Sbjct: 60 CCDLETGNNMNGEEDYSDDGSQMGEKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARAL 119
Query: 160 NLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKA 212
L+ RQ+ +WFQNRRAR K KQ E D + LK+ +TL EN L+ Q+L+A
Sbjct: 120 GLQPRQIAIWFQNRRARWKTKQLEKDYDTLKRQFDTLKAENDLLQTHNQKLQA 172
>At1g26960 putative DNA-binding protein
Length = 255
Score = 83.6 bits (205), Expect = 1e-16
Identities = 47/112 (41%), Positives = 65/112 (57%)
Query: 117 DDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRAR 176
DD KK RL EQ LE+ F+ + L +K LAR L L+ RQ+ +WFQNRRAR
Sbjct: 63 DDGSKMGEKKRRLNMEQLKALEKDFELGNKLESDRKLELARALGLQPRQIAIWFQNRRAR 122
Query: 177 TKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATL 228
+K KQ E D + LK+ E+L DEN L+ + Q+L+A +A P+ + L
Sbjct: 123 SKTKQLEKDYDMLKRQFESLRDENEVLQTQNQKLQAQVMALKSREPIESINL 174
>At1g70920 hypothetical protein
Length = 104
Score = 82.4 bits (202), Expect = 2e-16
Identities = 43/62 (69%), Positives = 47/62 (75%)
Query: 115 DEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
D+ + G RKKLRLTKEQS LLEESF Q+ TL PKQK+ LA L L RQVEVWFQNRR
Sbjct: 29 DDSNSGGRRRKKLRLTKEQSHLLEESFIQNHTLTPKQKKDLATFLKLSQRQVEVWFQNRR 88
Query: 175 AR 176
AR
Sbjct: 89 AR 90
>At5g15150 homeobox protein
Length = 293
Score = 78.6 bits (192), Expect = 3e-15
Identities = 51/113 (45%), Positives = 67/113 (59%), Gaps = 10/113 (8%)
Query: 112 RVSDED---DDGTN---ARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQ 165
+V +ED DDG++ KK RL EQ LE+SF+ + L P++K LA+ L L+ RQ
Sbjct: 75 QVGEEDNLSDDGSHMMLGEKKKRLNLEQVRALEKSFELGNKLEPERKMQLAKALGLQPRQ 134
Query: 166 VEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRL----KKELQELKALK 214
+ +WFQNRRAR K KQ E D + LKK + L +N L KK EL ALK
Sbjct: 135 IAIWFQNRRARWKTKQLERDYDSLKKQFDVLKSDNDSLLAHNKKLHAELVALK 187
>At3g01220 putative homeobox-leucine zipper protein, HAT7
Length = 286
Score = 77.0 bits (188), Expect = 1e-14
Identities = 47/104 (45%), Positives = 63/104 (60%), Gaps = 7/104 (6%)
Query: 118 DDGTNA---RKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
DDG + KK RL EQ LE+SF+ + L P++K LA+ L ++ RQ+ +WFQNRR
Sbjct: 77 DDGAHTMLGEKKKRLQLEQVKALEKSFELGNKLEPERKIQLAKALGMQPRQIAIWFQNRR 136
Query: 175 ARTKLKQTEVDCEFLKKCCETLTDENRRL----KKELQELKALK 214
AR K +Q E D + LKK E+L +N L KK L E+ ALK
Sbjct: 137 ARWKTRQLERDYDSLKKQFESLKSDNASLLAYNKKLLAEVMALK 180
>At3g01470 homeobox protein (HAT5)
Length = 272
Score = 76.3 bits (186), Expect = 2e-14
Identities = 45/103 (43%), Positives = 60/103 (57%), Gaps = 7/103 (6%)
Query: 115 DEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
D+ D KK RLT EQ LLE+SF+ + L P++K LA++L L+ RQV VWFQNRR
Sbjct: 58 DDFYDDQLPEKKRRLTTEQVHLLEKSFETENKLEPERKTQLAKKLGLQPRQVAVWFQNRR 117
Query: 175 ARTKLKQTEVDCEFLKKCCETLTD-------ENRRLKKELQEL 210
AR K KQ E D + LK + L +N +L+ E+ L
Sbjct: 118 ARWKTKQLERDYDLLKSTYDQLLSNYDSIVMDNDKLRSEVTSL 160
>At1g27050 unknown protein
Length = 495
Score = 74.7 bits (182), Expect = 5e-14
Identities = 58/152 (38%), Positives = 77/152 (50%), Gaps = 29/152 (19%)
Query: 125 KKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEV 184
KK +LT Q LLEESF++ L P +K LA +L L+ QV VWFQNRRAR K KQ E
Sbjct: 68 KKRKLTPIQLRLLEESFEEEKRLEPDRKLWLAEKLGLQPSQVAVWFQNRRARYKTKQLEH 127
Query: 185 DCEFLKKCCETL-TD------ENRRLKKELQELKALK--------------------LAQ 217
DC+ LK L TD +N+ LK ++Q L L L +
Sbjct: 128 DCDSLKASYAKLKTDWDILFVQNQTLKSKVQFLNRLTSHYFQESVQNFDDTFKQVDLLKE 187
Query: 218 PLYMPMSAATLTMCPSCERLGDGGSNIKSPFT 249
L M + T ++ +RLG+ GS++KS T
Sbjct: 188 KLKMQENLETQSI--ERKRLGEEGSSVKSDNT 217
>At5g03790 homeodomain -like protein
Length = 236
Score = 72.0 bits (175), Expect = 3e-13
Identities = 44/105 (41%), Positives = 65/105 (61%), Gaps = 1/105 (0%)
Query: 125 KKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEV 184
KK RLT Q A LE SF++ L+ +K L+R+L L+ RQ+ VWFQNRRAR K KQ E
Sbjct: 78 KKKRLTSGQLASLERSFQEEIKLDSDRKVKLSRELGLQPRQIAVWFQNRRARWKAKQLEQ 137
Query: 185 DCEFLKKCCETLTDENRRLKKELQELKALKLAQPLY-MPMSAATL 228
+ L++ + ++ E + L E+++L+AL Q L +SA T+
Sbjct: 138 LYDSLRQEYDVVSREKQMLHDEVKKLRALLRDQGLIKKQISAGTI 182
>At5g65310 homeobox-leucine zipper protein ATHB-5 (HD-zip protein
ATHB-5) (sp|P46667)
Length = 312
Score = 71.2 bits (173), Expect = 5e-13
Identities = 43/99 (43%), Positives = 61/99 (61%), Gaps = 7/99 (7%)
Query: 121 TNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLK 180
T A KK RL EQ LE++F+ + L P++K LA++L L+ RQV +WFQNRRAR K K
Sbjct: 68 TAAEKKRRLGVEQVKALEKNFEIDNKLEPERKVKLAQELGLQPRQVAIWFQNRRARWKTK 127
Query: 181 QTEVD-------CEFLKKCCETLTDENRRLKKELQELKA 212
Q E D + LK+ ++L +N L +++ELKA
Sbjct: 128 QLERDYGVLKSNFDALKRNRDSLQRDNDSLLGQIKELKA 166
>At2g46680 homeodomain transcription factor (ATHB-7)
Length = 258
Score = 71.2 bits (173), Expect = 5e-13
Identities = 41/99 (41%), Positives = 57/99 (57%), Gaps = 4/99 (4%)
Query: 122 NARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQ 181
N + R + EQ LE F+ + L P++K LAR+L L+ RQV +WFQN+RAR K KQ
Sbjct: 29 NKNNQRRFSDEQIKSLEMMFESETRLEPRKKVQLARELGLQPRQVAIWFQNKRARWKSKQ 88
Query: 182 TEVDCEFLKKCCETLTDENRRLKKELQ----ELKALKLA 216
E + L++ + L + LKKE Q EL+ LK A
Sbjct: 89 LETEYNILRQNYDNLASQFESLKKEKQALVSELQRLKEA 127
>At2g22430 homeodomain transcription factor (ATHB-6)
Length = 311
Score = 69.7 bits (169), Expect = 2e-12
Identities = 44/116 (37%), Positives = 64/116 (54%), Gaps = 4/116 (3%)
Query: 103 IEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLR 162
+E E + +E + KK RL+ Q LE++F+ + L P++K LA++L L+
Sbjct: 40 LEGYEEEEEAIVEERGHVGLSEKKRRLSINQVKALEKNFELENKLEPERKVKLAQELGLQ 99
Query: 163 ARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKE----LQELKALK 214
RQV VWFQNRRAR K KQ E D LK ++L L+++ LQE+ LK
Sbjct: 100 PRQVAVWFQNRRARWKTKQLEKDYGVLKTQYDSLRHNFDSLRRDNESLLQEISKLK 155
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,976,800
Number of Sequences: 26719
Number of extensions: 249274
Number of successful extensions: 1579
Number of sequences better than 10.0: 194
Number of HSP's better than 10.0 without gapping: 82
Number of HSP's successfully gapped in prelim test: 112
Number of HSP's that attempted gapping in prelim test: 1411
Number of HSP's gapped (non-prelim): 247
length of query: 267
length of database: 11,318,596
effective HSP length: 98
effective length of query: 169
effective length of database: 8,700,134
effective search space: 1470322646
effective search space used: 1470322646
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0132.1


