

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0070.9
(437 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
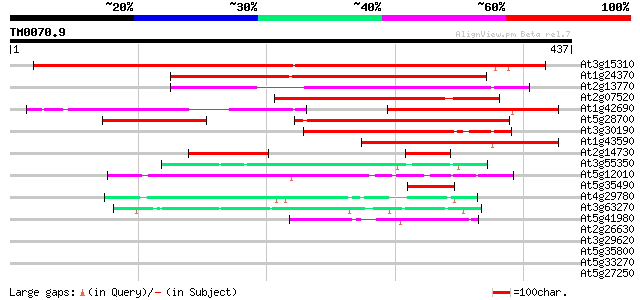
Score E
Sequences producing significant alignments: (bits) Value
At3g15310 unknown protein 412 e-115
At1g24370 hypothetical protein 326 2e-89
At2g13770 hypothetical protein 250 9e-67
At2g07520 pseudogene 199 2e-51
At1g42690 unknown protein 197 7e-51
At5g28700 putative protein 173 1e-43
At3g30190 hypothetical protein 153 2e-37
At1g43590 hypothetical protein 140 1e-33
At2g14730 hypothetical protein 76 4e-14
At3g55350 unknown protein 61 1e-09
At5g12010 putative protein 57 2e-08
At5g35490 putative protein 47 3e-05
At4g29780 unknown protein 45 8e-05
At3g63270 unknown protein 44 2e-04
At5g41980 unknown protein 43 3e-04
At2g26630 En/Spm-like transposon protein 41 0.001
At3g29620 hypothetical protein 39 0.004
At5g35800 putative protein 37 0.021
At5g33270 putative protein 35 0.11
At5g27250 putative protein 34 0.14
>At3g15310 unknown protein
Length = 415
Score = 412 bits (1060), Expect = e-115
Identities = 206/406 (50%), Positives = 278/406 (67%), Gaps = 8/406 (1%)
Query: 19 DFMNDTFVEDMMQQEMEFYQQQQHANTVRPKKTRRV-IKRDREAGNERLMKDYFSKNPVY 77
D M ++F + + QQ + + ++ V K +R R RE G+ L DYFS N ++
Sbjct: 11 DAMEESFDQLLDQQFENVFVELSNSEEVAKKNRKRAYFDRKREEGHVLLWNDYFSDNAIF 70
Query: 78 TEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAYGSP 137
+ FRRRFRM+K +FLRIV+ L S +F DA GR G SP+QKCTAAIR+LAYG
Sbjct: 71 PLQTFRRRFRMKKPLFLRIVDRLSSELMFFQHRRDATGRFGHSPIQKCTAAIRLLAYGYA 130
Query: 138 ADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGESRGFPGMLG 197
+D+VDEY+R+GE+TA+ CL+NF +G+ FG +YLR P ++ RLL G+ RGFPGM+G
Sbjct: 131 SDAVDEYLRMGETTAMSCLENFTKGIISFFGDEYLRAPTATNLRRLLNIGKIRGFPGMIG 190
Query: 198 SIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAGSNNDITVLN 257
S+DCMHWEWKNCP AWKGQYTRG G+PTI+LEA+ASQDLWIWH FFG G+ NDI +L+
Sbjct: 191 SLDCMHWEWKNCPTAWKGQYTRGS-GKPTIVLEAIASQDLWIWHVFFGPPGTLNDINILD 249
Query: 258 QSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQGEKKQKFAKR 317
+SP+F+++L+G AP VK++VN YH+ YYL DGIYP+W TF+++I +PQ K FA
Sbjct: 250 RSPIFDDILQGRAPNVKYKVNGREYHLAYYLTDGIYPKWATFIQSIRLPQNRKATLFATH 309
Query: 318 QEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDERNTYKGNF 377
QEA RKDVERAFGVLQ+RF I++ P+ W + +I+ ACIILHNMIVEDER+ Y F
Sbjct: 310 QEADRKDVERAFGVLQARFHIIKNPALVWDKEKIGNIMKACIILHNMIVEDERDGYNIQF 369
Query: 378 ---VYEQVNNDIS---DAEVLSGPIPAFRNMLERRAHQIEKSIHRQ 417
+ QV + + D +G NM+ R ++++H+Q
Sbjct: 370 DVSEFLQVEGNQTPQVDLSYSTGMPLNIENMMGMRNQLRDQNMHQQ 415
>At1g24370 hypothetical protein
Length = 413
Score = 326 bits (835), Expect = 2e-89
Identities = 155/246 (63%), Positives = 185/246 (75%), Gaps = 1/246 (0%)
Query: 126 TAAIRMLAYGSPADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQ 185
TAA RMLAYG PADS DEY++IGESTA+E LK F + EVF +YLR P+ D+ RLL
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACRYLRSPDANDVARLLH 61
Query: 186 WGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFG 245
GESRGFP MLGS+DCMHW+WKNCP AW GQY G PTI+LEAVA DLWIWHA+FG
Sbjct: 62 IGESRGFPRMLGSLDCMHWKWKNCPTAWGGQYA-GRSRSPTIILEAVADYDLWIWHAYFG 120
Query: 246 IAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPM 305
+ GSNNDI VL S +F + G AP + +NE Y++ YYLADGIYP+W T V+TI
Sbjct: 121 LPGSNNDINVLEASHLFANLAEGTAPPASYVINEKPYNMSYYLADGIYPKWSTLVQTIHD 180
Query: 306 PQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMI 365
P+G KK+ FA +QEA RKDVERAFGVLQSRFAIV GPSR W+ + I+ +CII+HNMI
Sbjct: 181 PRGPKKKLFAMKQEACRKDVERAFGVLQSRFAIVAGPSRLWNKTVLHDIMTSCIIMHNMI 240
Query: 366 VEDERN 371
+EDER+
Sbjct: 241 IEDERD 246
>At2g13770 hypothetical protein
Length = 244
Score = 250 bits (639), Expect = 9e-67
Identities = 136/280 (48%), Positives = 167/280 (59%), Gaps = 38/280 (13%)
Query: 126 TAAIRMLAYGSPADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQ 185
TAA RMLAYG PADS DEY++IGESTA+E LK F + EVF YLR P+ D+ RLL
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACCYLRSPDANDVARLLH 61
Query: 186 WGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFG 245
GESRGFP EAVA DLWIWHA+FG
Sbjct: 62 IGESRGFP------------------------------------EAVADYDLWIWHAYFG 85
Query: 246 IAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPM 305
+ GSNN I VL S +F + G AP + +N Y++GYYLADGIYP+W T V+TI
Sbjct: 86 LPGSNNGINVLEASHLFANLAEGTAPPASYVINGKPYNMGYYLADGIYPKWSTLVQTIHD 145
Query: 306 PQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMI 365
P+G KK+ FA +QEA RKDVERAFGVLQ RFAIV GPSR W+ + I+ +CII+HNMI
Sbjct: 146 PRGPKKKLFAMKQEACRKDVERAFGVLQLRFAIVAGPSRLWNKTVLHDIMTSCIIMHNMI 205
Query: 366 VEDERNTYKGNFVYEQVNNDISDAEVLSGPIPAFRNMLER 405
+EDER+ + E+V I + E+ F+ L R
Sbjct: 206 IEDERDIDAP--IEERVEVPIEEVEMTGDDDTRFQEFLAR 243
>At2g07520 pseudogene
Length = 222
Score = 199 bits (506), Expect = 2e-51
Identities = 96/176 (54%), Positives = 126/176 (71%), Gaps = 7/176 (3%)
Query: 207 KNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVL 266
KNCP+AW+GQ+TRGD G TI EAVAS DLWIWH FFG A + NDI L++SPVF+++
Sbjct: 16 KNCPIAWEGQFTRGDKGTSTISFEAVASHDLWIWHTFFGCASTLNDINFLDRSPVFDDIE 75
Query: 267 RGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQGEKKQKFAKRQEAARKDVE 326
+G P V F V + Y++ YYLADGIYP + TFVK+I +PQ E + F + QE RKD+E
Sbjct: 76 QGNTPRVNFFVYQRQYNMAYYLADGIYPSYPTFVKSIRLPQSEPDKLFVQLQEGCRKDIE 135
Query: 327 RAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDERNTYKGNFV-YEQ 381
RAFGVL +RF I+ W+ D+ I+ +CIILHNMIVE+ER+TY ++ Y+Q
Sbjct: 136 RAFGVLHARFKII------WNIADLAIIMRSCIILHNMIVENERDTYAQHWTDYDQ 185
>At1g42690 unknown protein
Length = 333
Score = 197 bits (502), Expect = 7e-51
Identities = 106/218 (48%), Positives = 132/218 (59%), Gaps = 35/218 (16%)
Query: 14 NECVEDFMNDTFVEDMMQQEMEFYQQQQHANTVRPKKTRRVIKRDREAGNERLMKDYFSK 73
N ++D +D F + Q +E Y +Q V+P+K + I+R+RE G+ +L+ DYF++
Sbjct: 7 NYNLDDMFDDKF-DQCFDQALESYGNRQR---VKPRKKKLYIERNREEGHIQLVNDYFTE 62
Query: 74 NPVYTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLA 133
NP Y +FRRRFRM K +F+RIVE + PYF DA R G S LQK TAAIRMLA
Sbjct: 63 NPTYPPHIFRRRFRMNKSLFMRIVERFSNEVPYFKQRRDATRRLGFSALQKSTAAIRMLA 122
Query: 134 YGSPADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGESRGFP 193
YG AD+ YLRRP +D+ RLL GE RGFP
Sbjct: 123 YGIAADA------------------------------YLRRPTRDDLIRLLHIGEQRGFP 152
Query: 194 GMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEA 231
GM+GSIDCMHWEWKNCP AWKGQYTRG G+PTI+LEA
Sbjct: 153 GMIGSIDCMHWEWKNCPTAWKGQYTRGS-GKPTIVLEA 189
Score = 103 bits (256), Expect = 2e-22
Identities = 60/138 (43%), Positives = 87/138 (62%), Gaps = 5/138 (3%)
Query: 295 EWGTFVKTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSI 354
E TF+++I + QG K FA QEA RKDVERAFGVLQ+RFAI++ P+ + + +I
Sbjct: 188 EAATFIQSISILQGNKASLFATTQEACRKDVERAFGVLQARFAIIKHPALFHDKVKIGNI 247
Query: 355 IYACIILHNMIVEDERNTYKGNFVYEQVNNDISDAE----VLSGPIPA-FRNMLERRAHQ 409
+ ACIILHNMIVED+R+ Y V E V+ + + + + +P+ NM+ A
Sbjct: 248 MRACIILHNMIVEDKRDGYTQFDVSEFVHPESASSSQVDFTYATVMPSNLGNMMATGARV 307
Query: 410 IEKSIHRQLQADLVEHIW 427
++ H +L+ADLVEH+W
Sbjct: 308 RDRIKHEELKADLVEHVW 325
>At5g28700 putative protein
Length = 292
Score = 173 bits (439), Expect = 1e-43
Identities = 87/167 (52%), Positives = 119/167 (71%), Gaps = 2/167 (1%)
Query: 223 GRPTIMLEAVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMY 282
G T+++ VASQDLWIWHA FG G+ NDI VL++SPVF+++L+G A VK+ VN Y
Sbjct: 105 GESTLLV--VASQDLWIWHAIFGPPGTLNDINVLDRSPVFDDILQGRASKVKYVVNGKDY 162
Query: 283 HIGYYLADGIYPEWGTFVKTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGP 342
++ YYL DGIYP+W TF+++I PQG+K F Q+A RKDVERAFGV Q+RF+IV+ P
Sbjct: 163 NLAYYLTDGIYPKWATFIQSISNPQGDKASLFVTTQKACRKDVERAFGVFQARFSIVKHP 222
Query: 343 SRWWHPNDMKSIIYACIILHNMIVEDERNTYKGNFVYEQVNNDISDA 389
+ + + +I+ ACIILHNMIVEDER+ Y V E V+ + + +
Sbjct: 223 ALFHDKVKIGNIMRACIILHNMIVEDERDGYTQCDVSEFVHPEFASS 269
Score = 92.0 bits (227), Expect = 6e-19
Identities = 47/81 (58%), Positives = 55/81 (67%)
Query: 73 KNPVYTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRML 132
+N Y +FR RFRM K +F+RIVE + PYF DA GR S LQK TAAIRML
Sbjct: 30 ENATYPPHIFRHRFRMNKPLFMRIVERFSNEVPYFKQRRDATGRLDFSALQKSTAAIRML 89
Query: 133 AYGSPADSVDEYVRIGESTAI 153
AYG AD+VDEY+RIGEST +
Sbjct: 90 AYGIAADAVDEYLRIGESTLL 110
>At3g30190 hypothetical protein
Length = 263
Score = 153 bits (386), Expect = 2e-37
Identities = 83/162 (51%), Positives = 103/162 (63%), Gaps = 8/162 (4%)
Query: 230 EAVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLA 289
+AVA D+WIWHA+FG+ GSNNDI VL +F + AP + N Y++GYYLA
Sbjct: 104 QAVADYDMWIWHAYFGLPGSNNDINVLEAYHLFANLAEDTAPPASYVSNGKPYNMGYYLA 163
Query: 290 DGIYPEWGTFVKTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPN 349
DGIY +W T V+TI P+G KK+ FA + E RKDVE AF VL SRFAIV PSR W N
Sbjct: 164 DGIYSKWSTLVQTIHDPRGPKKKLFAMKLETCRKDVEPAFEVLHSRFAIVAEPSRLW--N 221
Query: 350 DMKSIIYACIILHNMIVEDERNTYKGNFVYEQVNNDISDAEV 391
M S CII+HNMI+EDER+ + EQV I + E+
Sbjct: 222 KMTS----CIIMHNMIIEDERDIDAP--IEEQVKVPIEEVEM 257
>At1g43590 hypothetical protein
Length = 168
Score = 140 bits (354), Expect = 1e-33
Identities = 75/158 (47%), Positives = 100/158 (62%), Gaps = 5/158 (3%)
Query: 275 FRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQS 334
F VN Y++ YYL D IYP+W TF+++I +PQ EK FA QEA RKDVERAFGVLQ+
Sbjct: 3 FVVNGNEYNLAYYLTDEIYPKWATFIQSISLPQDEKASLFATNQEACRKDVERAFGVLQA 62
Query: 335 RFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDERNTYK----GNFVYEQVNNDISDAE 390
RFAIV+ P+ W + +I+ ACIILHNMIVEDER+ Y +F + + +
Sbjct: 63 RFAIVKHPALIWDKIKIGNIMRACIILHNMIVEDERDGYTQYDVSDFAHPESASSSQVDF 122
Query: 391 VLSGPIPA-FRNMLERRAHQIEKSIHRQLQADLVEHIW 427
S +P+ NM+ R +++ H L+ADLVEHIW
Sbjct: 123 TYSTDMPSNLGNMMATRTRLRDRTKHEHLKADLVEHIW 160
>At2g14730 hypothetical protein
Length = 117
Score = 75.9 bits (185), Expect = 4e-14
Identities = 34/62 (54%), Positives = 47/62 (74%)
Query: 140 SVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGESRGFPGMLGSI 199
+VD+Y+RIGE+T + C+ + VE + +FG +YLRRP +D+ RLL+ GE RGF GM GSI
Sbjct: 19 AVDKYLRIGENTLMSCMIHSVEAIIYLFGKEYLRRPTRQDLKRLLRIGELRGFLGMTGSI 78
Query: 200 DC 201
DC
Sbjct: 79 DC 80
Score = 48.1 bits (113), Expect = 9e-06
Identities = 23/35 (65%), Positives = 26/35 (73%)
Query: 309 EKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPS 343
+K FA QE RKDVERAFGVLQ+RFAIV P+
Sbjct: 81 DKDSLFATNQEVCRKDVERAFGVLQARFAIVTNPT 115
>At3g55350 unknown protein
Length = 406
Score = 61.2 bits (147), Expect = 1e-09
Identities = 68/262 (25%), Positives = 106/262 (39%), Gaps = 16/262 (6%)
Query: 119 LSPLQKCTAAIRMLAYGSPADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEE 178
LS + A+R L G + E + +ST + FVE + E +L P++
Sbjct: 109 LSLNDRVAVALRRLGSGESLSVIGETFGMNQSTVSQITWRFVESM-EERAIHHLSWPSKL 167
Query: 179 DMTRLLQWGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLW 238
D + ++ + G P G+ID H V + ++ L+AV D+
Sbjct: 168 DEIKS-KFEKISGLPNCCGAIDITHIVMNLPAVEPSNKVWLDGEKNFSMTLQAVVDPDMR 226
Query: 239 IWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPM-VKFRVNETMYHIGYYLADGIYPEWG 297
G GS ND VL S + V +G K ++E Y + D +P
Sbjct: 227 FLDVIAGWPGSLNDDVVLKNSGFYKLVEKGKRLNGEKLPLSERTELREYIVGDSGFPLLP 286
Query: 298 TFV-----KTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHP--ND 350
+ K +PQ E F KR A K + A L+ R+ I+ G W P N
Sbjct: 287 WLLTPYQGKPTSLPQTE----FNKRHSEATKAAQMALSKLKDRWRIINGVM--WMPDRNR 340
Query: 351 MKSIIYACIILHNMIVEDERNT 372
+ II+ C +LHN+I++ E T
Sbjct: 341 LPRIIFVCCLLHNIIIDMEDQT 362
>At5g12010 putative protein
Length = 502
Score = 57.4 bits (137), Expect = 2e-08
Identities = 72/322 (22%), Positives = 133/322 (40%), Gaps = 22/322 (6%)
Query: 77 YTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAYGS 136
Y EE F++ FRM K F I + L S + D R + Q+ I LA G
Sbjct: 170 YPEEDFKKAFRMSKSTFELICDELNSA----VAKEDTALRNAIPVRQRVAVCIWRLATGE 225
Query: 137 PADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGES-RGFPGM 195
P V + +G ST + + + + +V +YL+ P++E + + + ES G P +
Sbjct: 226 PLRLVSKKFGLGISTCHKLVLEVCKAIKDVLMPKYLQWPDDESLRNIRERFESVSGIPNV 285
Query: 196 LGSIDCMHWEWKNCPVAWKGQYT-----RGDHGRPTIMLEAVASQDLWIWHAFFGIAGSN 250
+GS+ H ++ + R +I ++AV + G GS
Sbjct: 286 VGSMYTTHIPIIAPKISVASYFNKRHTERNQKTSYSITIQAVVNPKGVFTDLCIGWPGSM 345
Query: 251 NDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQGEK 310
D VL +S ++ G + G+ L D + + + + Q
Sbjct: 346 PDDKVLEKSLLYQRANNGGLLKGMWVAGGP----GHPLLDWVLVPYTQ--QNLTWTQHAF 399
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDER 370
+K ++ Q A++ AFG L+ R+A ++ + D+ +++ AC +LHN I E
Sbjct: 400 NEKMSEVQGVAKE----AFGRLKGRWACLQKRTE-VKLQDLPTVLGACCVLHN-ICEMRE 453
Query: 371 NTYKGNFVYEQVNNDISDAEVL 392
+ + E +++++ VL
Sbjct: 454 EKMEPELMVEVIDDEVLPENVL 475
>At5g35490 putative protein
Length = 106
Score = 46.6 bits (109), Expect = 3e-05
Identities = 22/36 (61%), Positives = 26/36 (72%)
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWW 346
K+ FA +QEA RKDVER F VLQS+FAI+ S W
Sbjct: 69 KKMFAAKQEACRKDVERVFRVLQSKFAIIARSSNCW 104
>At4g29780 unknown protein
Length = 540
Score = 45.1 bits (105), Expect = 8e-05
Identities = 68/304 (22%), Positives = 119/304 (38%), Gaps = 37/304 (12%)
Query: 75 PVYTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAY 134
P + E+ FRR FRM K F I E L + + + + R + ++ + LA
Sbjct: 206 PDFPEDEFRREFRMSKSTFNLICEELDT----TVTKKNTMLRDAIPAPKRVGVCVWRLAT 261
Query: 135 GSPADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGES-RGFP 193
G+P V E +G ST + + + +V +YL P++ ++ ES P
Sbjct: 262 GAPLRHVSERFGLGISTCHKLVIEVCRAIYDVLMPKYLLWPSDSEINSTKAKFESVHKIP 321
Query: 194 GMLGSIDCMHWEW---KNCPVAW--KGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAG 248
++GSI H K A+ K R +I ++ V + D G G
Sbjct: 322 NVVGSIYTTHIPIIAPKVHVAAYFNKRHTERNQKTSYSITVQGVVNADGIFTDVCIGNPG 381
Query: 249 SNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQG 308
S D +L +S + + R A M+ R + + + G+ L D + +
Sbjct: 382 SLTDDQILEKSSLSRQ--RAARGML--RDSWIVGNSGFPLTDYLLVPY------------ 425
Query: 309 EKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRW--------WHPNDMKSIIYACII 360
+Q Q A + + G+ + F ++G RW D+ ++ AC +
Sbjct: 426 -TRQNLTWTQHAFNESIGEIQGIATAAFERLKG--RWACLQKRTEVKLQDLPYVLGACCV 482
Query: 361 LHNM 364
LHN+
Sbjct: 483 LHNI 486
>At3g63270 unknown protein
Length = 396
Score = 43.9 bits (102), Expect = 2e-04
Identities = 75/301 (24%), Positives = 117/301 (37%), Gaps = 28/301 (9%)
Query: 82 FRRRFRMRKHVFLRIV----EALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAYGSP 137
F+ FR K F I E L S P L++++ GR LS ++ A+R LA G
Sbjct: 65 FKHFFRASKTTFSYICSLVREDLISRPPSGLINIE--GRL-LSVEKQVAIALRRLASGDS 121
Query: 138 ADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEEDMTRL-LQWGESRGFPGML 196
SV +G+ST + F+E + E +LR P+ + + + ++ E G P
Sbjct: 122 QVSVGAAFGVGQSTVSQVTWRFIEAL-EERAKHHLRWPDSDRIEEIKSKFEEMYGLPNCC 180
Query: 197 GSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAGSNNDITVL 256
G+ID H P ++ L+ V ++ + G G +L
Sbjct: 181 GAIDTTH-IIMTLPAVQASDDWCDQEKNYSMFLQGVFDHEMRFLNMVTGWPGGMTVSKLL 239
Query: 257 NQSPVFN-----EVLRGAAPMVKFRVNETMYHIGYYLADGI-YP--EWGTFVKTIPMPQG 308
S F ++L G + I Y+ GI YP W P
Sbjct: 240 KFSGFFKLCENAQILDGNPKTLSQGA-----QIREYVVGGISYPLLPWLITPHDSDHP-S 293
Query: 309 EKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMK--SIIYACIILHNMIV 366
+ F +R E R AF L+ + I+ W P+ K SII C +LHN+I+
Sbjct: 294 DSMVAFNERHEKVRSVAATAFQQLKGSWRIL--SKVMWRPDRRKLPSIILVCCLLHNIII 351
Query: 367 E 367
+
Sbjct: 352 D 352
>At5g41980 unknown protein
Length = 374
Score = 43.1 bits (100), Expect = 3e-04
Identities = 41/151 (27%), Positives = 62/151 (40%), Gaps = 18/151 (11%)
Query: 219 RGDHGRPTIMLEAVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVN 278
R +G T + A +S DL + G GS +D VLN + L+ P K
Sbjct: 165 RNGNGLLTQNVLAASSFDLRFNYVLAGWEGSASDQQVLNAALTRRNKLQ--VPQGK---- 218
Query: 279 ETMYHIGYYLADGIYPEWGTFVKTI----PMPQGEKKQKFAKRQEAARKDVERAFGVLQS 334
YY+ D YP F+ + E K+ F +R + + + R FG L+
Sbjct: 219 -------YYIVDNKYPNLPGFIAPYHGVSTNSREEAKEMFNERHKLLHRAIHRTFGALKE 271
Query: 335 RFAIVRGPSRWWHPNDMKSIIYACIILHNMI 365
RF I+ + +K +I AC LHN +
Sbjct: 272 RFPILLSAPPYPLQTQVKLVIAAC-ALHNYV 301
>At2g26630 En/Spm-like transposon protein
Length = 292
Score = 40.8 bits (94), Expect = 0.001
Identities = 48/195 (24%), Positives = 76/195 (38%), Gaps = 36/195 (18%)
Query: 196 LGSIDCMHWEWKNCPVAWKGQYTRGDHGR---PTIMLEAVASQDLWIWHAFFGIAGSNND 252
+G++D H PV Q + GR PT+ + A+ + D+ +A+ G+ G +D
Sbjct: 45 VGALDGTH-----VPVRPPSQTAKKYKGRKLEPTMNVLAICNFDMKFIYAYVGVPGRAHD 99
Query: 253 ITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYP--------------EWGT 298
VLN NE P K YYL D YP G
Sbjct: 100 TKVLNYCAT-NEPYFSHPPNGK-----------YYLVDSGYPTRTGYLGPHRRMRYHLGQ 147
Query: 299 FVKTIPMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYAC 358
F + P ++ F ++ R +ER FGV ++++ IV + I+ A
Sbjct: 148 FGR--GGPPVTARELFNRKHSGLRSVIERTFGVWKAKWRIVDRKHPKYGLAKWIKIVTAT 205
Query: 359 IILHNMIVEDERNTY 373
+ LHN I + R +
Sbjct: 206 MALHNFIRDSHREDH 220
>At3g29620 hypothetical protein
Length = 222
Score = 39.3 bits (90), Expect = 0.004
Identities = 39/148 (26%), Positives = 64/148 (42%), Gaps = 27/148 (18%)
Query: 231 AVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIG-YYLA 289
AV + D+ + + G+ GS +D VL+ + ++ N HIG YYL
Sbjct: 31 AVCNFDMLFTYIYVGVFGSAHDTKVLSLA-------------MEGDPNFPHPHIGKYYLV 77
Query: 290 DGIYP----EWGTFVKTI--------PMPQGEKKQKFAKRQEAARKDVERAFGVLQSRFA 337
D Y G F +T P K+KF R R +ER FGV + ++
Sbjct: 78 DSGYALRRGYLGPFRQTWYHHNQFQNQAPPNNHKEKFNWRHSLLRCVIERTFGVWKGKWR 137
Query: 338 IVRGPSRWWHPNDMKSIIYACIILHNMI 365
I++ + W++ + I+ A + LHN +
Sbjct: 138 IMQDRA-WYNIVTTRKIMVATMALHNFV 164
>At5g35800 putative protein
Length = 73
Score = 37.0 bits (84), Expect = 0.021
Identities = 25/72 (34%), Positives = 38/72 (52%), Gaps = 13/72 (18%)
Query: 364 MIVEDERNTYKGNFV-YEQVNNDISDAEVLSGP-------IPAFRNMLERRAHQIEKSIH 415
MIV DER TY ++ Y Q S+A S P +P F + + R+ + ++H
Sbjct: 1 MIVNDERETYAQHWSDYNQ-----SEASGSSTPQPFSIEVLPEFASHVRARSELRDSNLH 55
Query: 416 RQLQADLVEHIW 427
+LQA LV++IW
Sbjct: 56 HELQAGLVKYIW 67
>At5g33270 putative protein
Length = 343
Score = 34.7 bits (78), Expect = 0.11
Identities = 38/187 (20%), Positives = 73/187 (38%), Gaps = 26/187 (13%)
Query: 196 LGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAGSNNDITV 255
+G++D H + P + + RG T+ + A+ + + +A+ G+ G +D V
Sbjct: 117 IGALDGTHVSVR--PPSGDVERYRGRKSEATMNILALCNFSMKFTYAYVGVPGRAHDTKV 174
Query: 256 LNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPM---------- 305
L +E P K YYL D YP ++
Sbjct: 175 LTYCAT-HEASFPHPPAGK-----------YYLVDSGYPTRSGYLGPHRRTRYHLELFNR 222
Query: 306 --PQGEKKQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHN 363
P ++ F +R + R +ER FGV ++++ I+ + I+ + + LHN
Sbjct: 223 GGPPTNSRELFNRRHSSLRSVIERTFGVWKAKWRILDRKHLKYEVKKWIKIVTSTMALHN 282
Query: 364 MIVEDER 370
I + ++
Sbjct: 283 YIRDSQQ 289
>At5g27250 putative protein
Length = 348
Score = 34.3 bits (77), Expect = 0.14
Identities = 38/148 (25%), Positives = 58/148 (38%), Gaps = 26/148 (17%)
Query: 231 AVASQDLWIWHAFFGIAGSNNDITVLNQSPVFNEVLRGAAPMVKFRVNETMYHIGYYLAD 290
A + DL + G GS +D VL Q + R P K YYLAD
Sbjct: 147 AACNFDLEFIYVLSGWEGSAHDSKVL-QDALTRRTNRLQVPEGK-----------YYLAD 194
Query: 291 GIYPEWGTFVKTIPMPQ-------GE------KKQKFAKRQEAARKDVERAFGVLQSRFA 337
+P F+ + + GE + + F R + R +ER FG+ +SRF
Sbjct: 195 CGFPNRRNFLAPLRSTRYHLQDFRGEGRDPTNQNELFNLRHASLRNVIERIFGIFKSRFL 254
Query: 338 IVRGPSRWWHPNDMKSIIYACIILHNMI 365
I + + + I+ +C LHN +
Sbjct: 255 IFKSAPPFSFKTQAE-IVLSCAALHNFL 281
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,326,810
Number of Sequences: 26719
Number of extensions: 450358
Number of successful extensions: 1047
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 1010
Number of HSP's gapped (non-prelim): 39
length of query: 437
length of database: 11,318,596
effective HSP length: 102
effective length of query: 335
effective length of database: 8,593,258
effective search space: 2878741430
effective search space used: 2878741430
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0070.9


