

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0069.5
(595 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
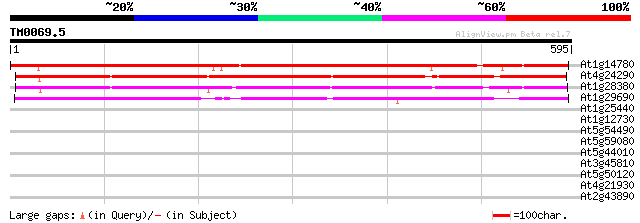
Score E
Sequences producing significant alignments: (bits) Value
At1g14780 unknown protein 684 0.0
At4g24290 unknown protein 496 e-140
At1g28380 unknown protein 462 e-130
At1g29690 unknown protein 401 e-112
At1g25440 unknown protein 35 0.15
At1g12730 unknown protein 32 0.77
At5g54490 unknown protein 31 2.2
At5g59080 putative protein 30 3.8
At5g44010 unknown protein 30 5.0
At3g45810 respiratory burst oxidase - like protein 30 5.0
At5g50120 putative protein 29 8.5
At4g21930 unknown protein 29 8.5
At2g43890 putative polygalacturonase 29 8.5
>At1g14780 unknown protein
Length = 627
Score = 684 bits (1765), Expect = 0.0
Identities = 356/622 (57%), Positives = 446/622 (71%), Gaps = 38/622 (6%)
Query: 2 GEGIVEKALNSLGKGFDLTSDFRLKFCK------GEERLVILNETERRELTVPGFGPVTD 55
G ++E A+ SLGKGFDLT+DFRLK+CK G++RLV+L++T+ REL +PGFG +
Sbjct: 5 GGDVIETAVKSLGKGFDLTADFRLKYCKDGDGSAGDDRLVVLDQTQNRELHIPGFGVFQN 64
Query: 56 VSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAA 115
VS DI CDKG+ TR++SDIL F +MSE FNQ+SS+ GKIPSG FN FGF++GSWA+DAA
Sbjct: 65 VSADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAA 124
Query: 116 NTKFLGLDGYVIKLFNVHIDR-YPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLS 174
N K LGLD V+ LFN+HI L L+ +V AVPSSWDP LARFIE++GTH++ G+S
Sbjct: 125 NVKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVS 184
Query: 175 IGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNF----LPKTKDQKH---KVPP 227
+GG+D+V+V+QD SS+L+ L+ +L +LGDQLFTG+C L K H K P
Sbjct: 185 VGGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPE 244
Query: 228 AFDVFGP-QIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFI 286
AF+VF Q VAFNN + + +++GITVICAKRGGD + +HSEWL+TVP+KPDA++F+FI
Sbjct: 245 AFNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFI 303
Query: 287 PFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRT 346
P TSLLK PG G LSHA++LYLRYKPPL DL YFLD+ R WAP+HNDLP G N
Sbjct: 304 PITSLLKDVPGSGLLSHAMSLYLRYKPPLMDLQYFLDFSGPRAWAPVHNDLPFGAAPNMA 363
Query: 347 THSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNN 406
+ P+L +N MGPKLYVNT VT K PVTGMR FLEG KCNRLAIHLQ+L NT T +
Sbjct: 364 SAYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCNRLAIHLQHLDNTRTTVGE 423
Query: 407 KIEDTTTW--SEEIID-DRFLEAISGKKFSHVCTAPVKYNPSW-------SSDKDVAFIV 456
KI D W S++I D DR+ E ++GKKFSHVCT PVKY+P+W S DVAFIV
Sbjct: 424 KITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDPNWIKTTSNHKSQNDVAFIV 483
Query: 457 TGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTS 516
TG QL VKKH S+SVLHLRL ++KVS+ VV+++W G G+ S +FS++S
Sbjct: 484 TGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPI-----GTSQKSGIFSSMSMP 538
Query: 517 IQGKD------QKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLV 570
+ QK VV+DS VFP GPPVP K++KFVD SQLC+GPQ SPGHWLV
Sbjct: 539 LTSGSVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFVDLSQLCRGPQHSPGHWLV 597
Query: 571 TGARLMLDKGKICLWAKFSLLN 592
TG RL LDKGK+CL KF+LL+
Sbjct: 598 TGVRLYLDKGKLCLHVKFALLH 619
>At4g24290 unknown protein
Length = 606
Score = 496 bits (1277), Expect = e-140
Identities = 274/594 (46%), Positives = 384/594 (64%), Gaps = 25/594 (4%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKG---EERLVILNETERR-ELTVPGFGPVTDVSVDIKC 62
E A+ S+G G+DL D RLK+CKG + RL+ + E + E+ +PG + +VS IKC
Sbjct: 12 EVAIGSIGCGYDLAIDLRLKYCKGGSKDSRLLDIKEGDDNCEIVLPGGISIPNVSKSIKC 71
Query: 63 DKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGL 122
DKG+ R++SDIL F QM+E FNQ+ S+ GKIPSG FN +F F + W DAA TK L
Sbjct: 72 DKGERMRFRSDILPFQQMAEQFNQELSLAGKIPSGLFNAMFEFSS-CWQKDAAYTKNLAF 130
Query: 123 DGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVL 182
DG I L++V +D+ ++L + V +AVPS+WDP ALARFI+ +GTHI+V + +GGKD++
Sbjct: 131 DGVFISLYSVALDKSQVLLREHVKQAVPSTWDPAALARFIDIYGTHIIVSVKMGGKDVIY 190
Query: 183 VKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNS 242
KQ SS L+P +L+K L E+ D+ F + + T ++ + + ++ + S
Sbjct: 191 AKQQHSSKLQPEDLQKRLKEVADKRFV-EASVVHNTGSERVQASSKVETKEQRLRFADTS 249
Query: 243 TC--VCAKDGITVICAKRGG-DTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRG 299
+ K+ +C +RGG D + H+EWL TV +PD + SFIP TSLL PG G
Sbjct: 250 SLGSYANKEDYVFMCKRRGGNDNRNLMHNEWLQTVQMEPDVISMSFIPITSLLNGVPGSG 309
Query: 300 FLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGP 359
FLSHAINLYLRYKPP+ +L FL++Q R WAP+ ++LPLGP + SL + GP
Sbjct: 310 FLSHAINLYLRYKPPIEELHQFLEFQLPRQWAPVFSELPLGP-QRKQQSCASLQFSFFGP 368
Query: 360 KLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEII 419
KLYVNT V VGKRP+TGMRL+LEG + NRLAIHLQ+L + P + + + + +E
Sbjct: 369 KLYVNTTPVDVGKRPITGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLNRSIRQESH 428
Query: 420 DDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFS 479
D R+ E ++ K +SHVCT PV+ SD D++ +VTG QLHV+ H ++VL LRL FS
Sbjct: 429 DRRYYEKVNWKNYSHVCTEPVE------SDDDLS-VVTGAQLHVESHGFKNVLFLRLCFS 481
Query: 480 KVSNAIVVK-SNWTQ--GSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFP 536
+V A +VK S W + G +SG S S F+A K +PA V ++S+++P
Sbjct: 482 RVVGATLVKNSEWDEAVGFAPKSGLISTLISHHFTAAQ-----KPPPRPADVNINSAIYP 536
Query: 537 TGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSL 590
GPPVP QA KLLKFVDTS++ +GPQ+SPG+W+V+GARL+++KGKI L K+SL
Sbjct: 537 GGPPVPTQAPKLLKFVDTSEMTRGPQESPGYWVVSGARLLVEKGKISLKVKYSL 590
>At1g28380 unknown protein
Length = 612
Score = 462 bits (1188), Expect = e-130
Identities = 270/602 (44%), Positives = 359/602 (58%), Gaps = 31/602 (5%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGE---ERLVILNETERRELTVPGFGPVTDVSVDIKCD 63
EKA++ +G G+DL SD R CK RLV ++ T R+L PG V +VS IKCD
Sbjct: 16 EKAVSVIGLGYDLCSDVRFSACKTTPDGSRLVEIDPTRNRDLIFPGGIVVNNVSSSIKCD 75
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLD 123
KG+ TR +SDILSF QMSE FNQ + GKIPSG FN +F F W DA++ K L D
Sbjct: 76 KGERTRLRSDILSFNQMSEKFNQDMCLSGKIPSGMFNNMFAFSK-CWPKDASSVKTLAYD 134
Query: 124 GYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLV 183
G+ I L++V I R L L +V VPSSWD ALA FIEK+GTH++VG+++GGKD++ V
Sbjct: 135 GWFISLYSVEIVRKQLTLRDEVKREVPSSWDSAALAGFIEKYGTHVVVGVTMGGKDVIHV 194
Query: 184 KQDVSSNLEPSELKKNLDELGDQLFT-------GTCNFLPKTKDQKHKVPPAFDVFGPQI 236
KQ SN EP E++K L GD+ F + +++ + FG +
Sbjct: 195 KQMRKSNHEPEEIQKMLKHWGDERFCVDPVESKSPASVYSGKPKEENLLQWGLQPFGTSV 254
Query: 237 VAFNNSTCVCAK-DGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAA 295
+++ + K + I +C +RGG +H WL TV P+ + F+P TSLL
Sbjct: 255 ---SSAVVMHTKNEEIMRVCIRRGGVDLGQSHERWLSTVSQAPNVISMCFVPITSLLSGL 311
Query: 296 PGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLN 355
PG GFLSHA+NLYLRYKPP+ +L FL++Q R WAP++ DLPLG + SPSL +
Sbjct: 312 PGTGFLSHAVNLYLRYKPPIEELHQFLEFQLPRQWAPVYGDLPLG-LRRSKQSSPSLQFS 370
Query: 356 LMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWS 415
LMGPKLYVNT+KV G+RPVTG+R FLEG K N LAIHLQ+L P L+ +DT
Sbjct: 371 LMGPKLYVNTSKVDSGERPVTGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTYEPI 430
Query: 416 EEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLR 475
EE ++ + + FSHVCT PV+YN + S D A IVT L VK R VL LR
Sbjct: 431 EEPVEKGYYVPVKWGIFSHVCTYPVQYNGARSD--DTASIVTKAWLEVKGMGMRKVLFLR 488
Query: 476 LLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPAL------VV 529
L FS ++A+ KS W ST SG VFS IST + PA +
Sbjct: 489 LGFSLDASAVTRKSCWDNLSTNSRKSG------VFSMISTRLSTGLSPNPATTKPQSKID 542
Query: 530 VDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFS 589
++S+V+P GP PV+ KLL VDT ++ +GP++ PG+W+VTGA+L ++ GKI + AK+S
Sbjct: 543 INSAVYPRGPSPPVK-PKLLSLVDTKEVMRGPEEQPGYWVVTGAKLCVEAGKISIKAKYS 601
Query: 590 LL 591
LL
Sbjct: 602 LL 603
>At1g29690 unknown protein
Length = 561
Score = 401 bits (1031), Expect = e-112
Identities = 240/596 (40%), Positives = 335/596 (55%), Gaps = 65/596 (10%)
Query: 6 VEKALNSLGKGFDLTSDFRLKFCKGE--ERLVILNETERRELTVPGFGPVTDVSVDIKCD 63
+ A+ +LG+GFD+TSD RL +CKG RLV + E + R+L + + +V DI C
Sbjct: 21 LRNAIQALGRGFDVTSDVRLLYCKGAPGSRLVRIEEGQNRDLELSHGFLLPNVPADIDCS 80
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLD 123
+G+ + + SF +M+E FN +S + G IP G FN +F + GSW DAA+TK L L
Sbjct: 81 RGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNY-TGSWQVDAASTKSLALV 139
Query: 124 GYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLV 183
GY I L++V + + LVL ++ AVPSSWDP +LA FIE +GTHI+ ++IGG+D+V +
Sbjct: 140 GYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGGRDVVYI 199
Query: 184 KQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNST 243
+Q SS L SE++ ++++ K + H+ + + GP
Sbjct: 200 RQHQSSPLPVSEIENYVNDM--------------IKHRFHEA-ESQSITGP--------- 235
Query: 244 CVCAKD-GITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLS 302
+ KD ITVI +RGGD +H+ W TVP PD ++ +F P SLL+ PG L+
Sbjct: 236 -LKYKDKDITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLEGVPGLRHLT 294
Query: 303 HAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLY 362
AI LYL YKPP+ DL YFLDYQ R WAP ++L + SL +LMGPKL+
Sbjct: 295 RAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQFSLMGPKLF 349
Query: 363 VNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIED-----TTTW-SE 416
++ +VTVG++PVTG+RL LEG K NRL+IHLQ+L++ P +L + W
Sbjct: 350 ISADQVTVGRKPVTGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVPIGAPKWQGP 409
Query: 417 EIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRL 476
E D R+ E I K FSHV T+P+++ + D IVTG QL V S++VLHL+L
Sbjct: 410 EEQDSRWFEPIKWKNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGVWDFGSKNVLHLKL 469
Query: 477 LFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFP 536
LFSKV + +S W SG G S S+
Sbjct: 470 LFSKVPGCTIRRSVWDHTPVASSGRLEPGGPSTSSSTE---------------------- 507
Query: 537 TGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
V Q+ KL K VD+S++ KGPQD PGHWLVTGA+L ++KGKI L K+SLLN
Sbjct: 508 ---EVSGQSGKLAKIVDSSEMLKGPQDLPGHWLVTGAKLGVEKGKIVLRVKYSLLN 560
>At1g25440 unknown protein
Length = 417
Score = 34.7 bits (78), Expect = 0.15
Identities = 29/110 (26%), Positives = 42/110 (37%), Gaps = 16/110 (14%)
Query: 429 GKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVK 488
G K + C + VK W D AF+ V + H R+ S A+V
Sbjct: 10 GAKTARACDSCVKRRARWYCAADDAFLCQSCDSLVHSANPLARRHERVRLKTASPAVVKH 69
Query: 489 SN------------WTQGSTQRS----GSGSGSGSSVFSAISTSIQGKDQ 522
SN W G T+++ GSG + SS+F + I +DQ
Sbjct: 70 SNHSSASPPHEVATWHHGFTRKARTPRGSGKKNNSSIFHDLVPDISIEDQ 119
>At1g12730 unknown protein
Length = 474
Score = 32.3 bits (72), Expect = 0.77
Identities = 19/70 (27%), Positives = 38/70 (54%), Gaps = 5/70 (7%)
Query: 271 LLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPY----FLDYQA 326
+ + ++P + F+++ FTS+LK+ P G + ++L+ + L+D+ Y F Y
Sbjct: 353 MFRLKHRPCFLAFAYLAFTSILKSYPSVGDAALYLSLWALFVNELTDMEYSFFIFCGYIG 412
Query: 327 HRLWAPI-HN 335
L +P+ HN
Sbjct: 413 FSLLSPVMHN 422
>At5g54490 unknown protein
Length = 127
Score = 30.8 bits (68), Expect = 2.2
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 13/69 (18%)
Query: 2 GEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERREL-TVPGFGPVTDVSVDI 60
GEG++E+ + KGF+L D +++ VI E+ RR TV G G +TD V
Sbjct: 32 GEGLIEE----ICKGFELLMD--------KDKGVITFESLRRNASTVLGLGDLTDDDVRY 79
Query: 61 KCDKGDLTR 69
++GD R
Sbjct: 80 MINEGDFDR 88
>At5g59080 putative protein
Length = 135
Score = 30.0 bits (66), Expect = 3.8
Identities = 19/55 (34%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTK 118
KG + S SFT +E+F K P SG F+T+F + A D +N+K
Sbjct: 4 KGRVGSSSSTSSSFT--AELFGSKDPSPPSSSSGIFSTMFPHPSKGSARDGSNSK 56
>At5g44010 unknown protein
Length = 360
Score = 29.6 bits (65), Expect = 5.0
Identities = 32/111 (28%), Positives = 49/111 (43%), Gaps = 21/111 (18%)
Query: 337 LPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQY 396
L LG V++R TH L+ + V+ VTV + +L L+ KC
Sbjct: 214 LLLGCVTSRWTH-------LIEGIMSVSYKSVTVSEEYEELCKLLLQRSKC--------- 257
Query: 397 LLNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWS 447
L LN+K+E+ + EI++ RF + K S + A + PSWS
Sbjct: 258 LKQNEIALNSKVEEILEYLTEILESRFHQL--WKLPSALTAAAI---PSWS 303
>At3g45810 respiratory burst oxidase - like protein
Length = 835
Score = 29.6 bits (65), Expect = 5.0
Identities = 13/37 (35%), Positives = 23/37 (62%)
Query: 157 ALARFIEKFGTHILVGLSIGGKDLVLVKQDVSSNLEP 193
A A+ +KF +L+GL IG + + +D+ +NL+P
Sbjct: 659 APAQSYQKFDILLLIGLGIGATPFISILKDMLNNLKP 695
>At5g50120 putative protein
Length = 388
Score = 28.9 bits (63), Expect = 8.5
Identities = 15/55 (27%), Positives = 28/55 (50%), Gaps = 6/55 (10%)
Query: 419 IDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLH 473
I+D E + GKK+ H+ T P SD+ ++ Q+ +++H+ S +H
Sbjct: 115 INDVVEEDVGGKKYMHLATMPT------ISDRFAKCLMPKNQVEIRRHKKASWVH 163
>At4g21930 unknown protein
Length = 183
Score = 28.9 bits (63), Expect = 8.5
Identities = 28/104 (26%), Positives = 45/104 (42%), Gaps = 26/104 (25%)
Query: 429 GKKFSHVCT-APVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVV 487
G++ HV T APVK P WS V + + +H +++ NA V
Sbjct: 85 GQRKRHVATSAPVKV-PDWSKILKVESVKS---MHNNNNDN-------------DNADVA 127
Query: 488 KSNWTQGSTQ-------RSGSGSGSGSSVFSAISTSIQGKDQKK 524
+W RS +G G GSSVF + +++G+D ++
Sbjct: 128 DCDWESAMVPPHEYVAARSRNGDG-GSSVFLGVGRTLKGRDMRR 170
>At2g43890 putative polygalacturonase
Length = 392
Score = 28.9 bits (63), Expect = 8.5
Identities = 17/59 (28%), Positives = 28/59 (46%), Gaps = 2/59 (3%)
Query: 469 RSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPAL 527
R+VL L+ V N I+V N+ + + GSG + + +IQG + + AL
Sbjct: 288 RNVLFQNLIMKNVQNPIIVDQNYC--PSNQGCPKQGSGVKISQVVYRNIQGTSRTQQAL 344
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,169,193
Number of Sequences: 26719
Number of extensions: 639136
Number of successful extensions: 1653
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1621
Number of HSP's gapped (non-prelim): 14
length of query: 595
length of database: 11,318,596
effective HSP length: 105
effective length of query: 490
effective length of database: 8,513,101
effective search space: 4171419490
effective search space used: 4171419490
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0069.5


