

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0060.4
(1482 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
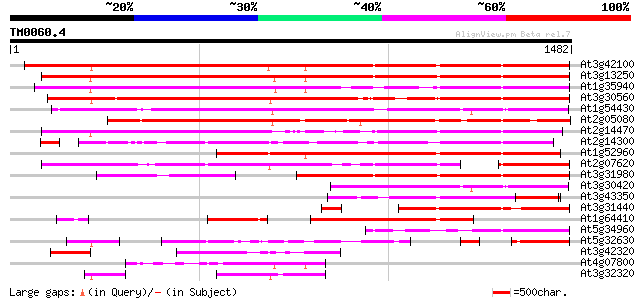
Score E
Sequences producing significant alignments: (bits) Value
At3g42100 putative protein 1274 0.0
At3g13250 hypothetical protein 1259 0.0
At1g35940 hypothetical protein 1142 0.0
At3g30560 hypothetical protein 1120 0.0
At1g54430 hypothetical protein 1041 0.0
At2g05080 putative helicase 1018 0.0
At2g14470 pseudogene 1009 0.0
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 855 0.0
At1g52960 hypothetical protein 843 0.0
At2g07620 putative helicase 820 0.0
At3g31980 hypothetical protein 619 e-177
At3g30420 hypothetical protein 471 e-132
At3g43350 putative protein 460 e-129
At3g31440 hypothetical protein 391 e-108
At1g64410 unknown protein 387 e-107
At5g34960 putative protein 334 3e-91
At5g32630 putative protein 321 2e-87
At3g42320 putative protein 301 1e-81
At4g07800 hypothetical protein 297 3e-80
At3g32320 hypothetical protein 234 4e-61
>At3g42100 putative protein
Length = 1752
Score = 1274 bits (3297), Expect = 0.0
Identities = 673/1471 (45%), Positives = 933/1471 (62%), Gaps = 37/1471 (2%)
Query: 39 LPQQRNVSDCTNNFNTNFAFVEAESQMYFDLGEMNMACQYCGAILWYHERAQKAKNAISP 98
LPQ + N +T + A Y D G+ C YCGA++W+ ER K + SP
Sbjct: 288 LPQPKKRGRPRLNKDTTNSAETATKSAYLDHGDATYKCNYCGALMWFAERINKKQQNKSP 347
Query: 99 DFSICCMKGKISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGS 158
F++CC KG + LP L+D+P L++NLLT D S +F +NIR YN +FA TS+GG+VD S
Sbjct: 348 TFTLCCGKGNVKLPLLKDSPALINNLLTGDDALSRNFRENIRIYNMIFAMTSLGGRVDNS 407
Query: 159 VNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEFENRLNHLSDSTG--- 215
+ G+GP F + G NYH IGSL P GD K++QL I DT NE +NR ++ G
Sbjct: 408 MPKGKGPNMFRLQGGNYHLIGSLKPNPGDYAKYSQLYIVDTENEVDNRATVINKGKGRRN 467
Query: 216 ---KCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSNQVALRLFRHRVN-DPKTY 271
K L +++ L+ M+++ N FR+ R+ +Q+ N +R+ R D +TY
Sbjct: 468 TPAKQKLKKEVIEALIEMLNKVNPYVDKFRQARERIQDDNDEPFHMRIVADRKGVDRRTY 527
Query: 272 NLPTVDEVAALIVEDFDTSDCGRDIILRT-SSGNLQRIYDTHSSFLPLQYPLIFPYGEEG 330
++PT EVAALI F S RDI+L ++G+L RI H S+L LQYPLI YGE+G
Sbjct: 528 SMPTSSEVAALIPGGFQPSMFDRDIVLEEKTTGHLTRISQIHISYLALQYPLILCYGEDG 587
Query: 331 FSDEIGFDGINYDSSIYKRTTISLREWVAFRLQERQFECKRITLSRRLLQQLVVDCYSMI 390
++ I + + K+ IS+R+W AFR+QER ECK +T S+RL QQ + D Y+ I
Sbjct: 588 YTPGIE-KCLPNSAKKKKKKCISMRQWFAFRIQERPNECKTLTRSKRLFQQFLCDAYTTI 646
Query: 391 ESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQD 450
ES RL Y++ Q +R + + +++A G T ++ G+++++PSS TGG RYM N D
Sbjct: 647 ESNRLSYIKFKQSKLRCENYNSLKKASEAGTTSMNEEGNQVLIPSSLTGGPRYMVQNYYD 706
Query: 451 AMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLK 510
AM IC+ YG+PDLF+T TCNPKWPEI RH AR LS DRPD+ R+F++KLD LMK+L
Sbjct: 707 AMAICKHYGFPDLFITFTCNPKWPEITRHCQARGLSVDDRPDIVARIFKIKLDSLMKDLT 766
Query: 511 KGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLL 570
GK G+ ++ M+T+EFQKRGLPHAHILL++ K KL + + ID +I AE+PD P L
Sbjct: 767 DGKMLGKTVASMHTVEFQKRGLPHAHILLFMDAKSKLPTADDIDKIISAEIPDKDKEPEL 826
Query: 571 YQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTN 630
Y+ + N M+HGPCG NSPCM G+CSK +PK + T DGYP+YRRR T
Sbjct: 827 YEVIKNSMIHGPCGAANMNSPCMVEGKCSKQYPKKHQDITKVGKDGYPIYRRRMTEDYIE 886
Query: 631 RRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVDRV--------- 681
+ G DNG+VVPYN KL ++YQAHIN+E+CN+S IKYLFKYINKG DRV
Sbjct: 887 KGGFKCDNGYVVPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINKGADRVVFIVEPVNQ 946
Query: 682 ---TMSMSVTQSSNEETTVVDEIKQYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHL 738
T + + + N DEIK + DCRY+S EAVWR + F + D+ V + S+H
Sbjct: 947 DKTTENATSGEPPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPLQDRSTAVQRLSFHD 1006
Query: 739 PKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLG------RGLTYAEFPSSFV 792
KQ + A I+DVLER + SMF+AW+ N +G R L Y++ P+ F
Sbjct: 1007 EGKQPVYAKPDADIEDVLERISNEDSMFMAWLTLNKNNDVGKNGKRARELLYSQIPAYFT 1066
Query: 793 YDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTF 852
+D K K+W R +G +GR+N+V YYLR+LLN+ +G S++D+++ N V+ +F
Sbjct: 1067 WDGKNKQWVKRIRGFSLGRINYVCRKMEVEYYLRVLLNIVKGPMSYDDIKTFNGVVYPSF 1126
Query: 853 RQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRETWSVLA 912
++AC A +L DD+ YIDG+ + + F G Y+R F LL +++A P HVW ETW +LA
Sbjct: 1127 KEACFARGILDDDQVYIDGLHEASQFCFGDYLRNFFAMLLLSDSLARPEHVWSETWHLLA 1186
Query: 913 DGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYG 972
+ I + R N +L + ++ L EIEK+++ NG +LK+ + P S I
Sbjct: 1187 EDIENKKREDFKNPDLKLTLAEIRNYTLQEIEKIMLRNGATLKEIQDFP-QPSREGIDNS 1245
Query: 973 NILLFNELNFD-TVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGK 1031
N L+ +EL ++ + + H E LN Q IY+EI AV +D G F+YG+GGTGK
Sbjct: 1246 NRLVVDELRYNIDSNLKEKHDEWFQMLNTEQRGIYDEITGAVFNDLGGVFFIYGFGGTGK 1305
Query: 1032 TFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSP 1091
TF+W TL +RS +I+LNVASSGIASLLL GGRTAHS F IPLN DE S C I S
Sbjct: 1306 TFIWKTLAAAVRSRGQIVLNVASSGIASLLLEGGRTAHSRFAIPLNPDEFSVCKITPKSD 1365
Query: 1092 KAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQ 1151
A L+K +SLIIWDEAPM+S++ FE+LD++ DI+ N + NK FGGKVVV GGDFRQ
Sbjct: 1366 LANLIKEASLIIWDEAPMMSKFCFESLDKSFYDILN--NKD--NKVFGGKVVVFGGDFRQ 1421
Query: 1152 ILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETS-DVEKIKVFAEWVLD 1210
+LPVI R EIVM+++N+S LW CKVL LT+NMRL S +S + ++I+ F++W+L
Sbjct: 1422 VLPVINGAGRVEIVMSSLNASYLWDHCKVLKLTKNMRLLSGGLSSEEAKEIQQFSDWLLA 1481
Query: 1211 IGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY--PEIMENVGCAKYYEDKAIL 1268
+GDG + + NDG+A I +P++ L++++ NP+ I + IY P + + K+++ +AIL
Sbjct: 1482 VGDGRINEPNDGEALIDIPEELLIKEAGNPIEAISKEIYGDPSELHMINDPKFFQRRAIL 1541
Query: 1269 APTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEFLNDIRCSGIPD 1328
APT + V+ +NQY+L ER YLS+ S+ T+ + IT +FLN I+ +G+P
Sbjct: 1542 APTNEDVNTINQYMLEHLKSEERIYLSADSI-DPTDSDSLANPVITPDFLNSIQLTGMPH 1600
Query: 1329 HKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDL 1388
H L LK GAPVML+RNLD GLCNGTR+ +T L VV VI+ +G V I ++L
Sbjct: 1601 HALRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLAKQVVQAKVITRDRIGDIVLIPLINL 1660
Query: 1389 MPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKT 1448
P+D +P K +RRQFPL ++FAMTINKSQG++L VGLYLP PVFSHGQLYVA+SRV +
Sbjct: 1661 TPSDTKLPFKMRRRQFPLSVAFAMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTS 1720
Query: 1449 RAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
+ GLKILI ++D + + T N+VFKEVFQ I
Sbjct: 1721 KKGLKILILDKDGNMQKQTTNVVFKEVFQNI 1751
>At3g13250 hypothetical protein
Length = 1419
Score = 1259 bits (3258), Expect = 0.0
Identities = 654/1427 (45%), Positives = 929/1427 (64%), Gaps = 38/1427 (2%)
Query: 83 LWYHERAQKAKNAISPDFSICCMKGKISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSY 142
+W++ER K N+ +P FS+CC +G + LP+L+++P L+ LL D S H+ IR Y
Sbjct: 1 MWFNERINKKSNSENPKFSLCCGQGSVKLPFLKESPELIKKLLKGNDALSRHYRQFIRIY 60
Query: 143 NSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNE 202
N +FA TS+GGKVD S+ G+GP F + G NYH+IGSL P +GD K++QL I DT NE
Sbjct: 61 NMIFAMTSLGGKVDKSMPKGRGPAMFRLQGGNYHQIGSLKPKDGDYAKYSQLYIVDTENE 120
Query: 203 FENRLNHL------SDSTGKCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSNQV 256
ENR N + S + GK +LN L+ ++ M+++ N + FR R+ + N
Sbjct: 121 VENRANVIGKGNNGSSTKGKKNLNKQLIDAIIKMLNQVNPYVEKFRSARERIDSTNDEPF 180
Query: 257 ALRLFRHRVN-DPKTYNLPTVDEVAALIVEDFDTSDCGRDIIL-RTSSGNLQRIYDTHSS 314
+R+ R D + YN+PT EVAALI DF + RDIIL + S+G L+RI H S
Sbjct: 181 HMRIVSDRKGTDGRLYNMPTAGEVAALIPGDFVSQMPVRDIILEKKSTGRLKRISQIHIS 240
Query: 315 FLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYKRTTISLREWVAFRLQERQFECKRITL 374
+L LQYPLIF YGE+G++ G + K+ IS+R+W AFR+QER+ E +
Sbjct: 241 YLALQYPLIFCYGEDGYTP--GIEKCYKSGYTKKKKCISMRQWYAFRIQEREDESHTLLQ 298
Query: 375 SRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLP 434
S+RL QQ + D Y+ IES RL Y++ NQ +R + + I+E+ G T +S G+++++P
Sbjct: 299 SKRLFQQFLCDAYTTIESNRLAYIKFNQSKLRCENFNSIKESASSGSTTMSEEGNQVLIP 358
Query: 435 SSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLA 494
SSFTGG RYM DAM IC+ +G+PDLF+T TCNPKWPEI R+ R L+A DRPD+
Sbjct: 359 SSFTGGPRYMLQTYYDAMAICKHFGFPDLFITFTCNPKWPEITRYCEKRGLTADDRPDIV 418
Query: 495 CRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELID 554
R+F++KLD LMK+L + G+ ++ MYT+EFQKRGLPHAHILL+++ KL + + ID
Sbjct: 419 ARIFKIKLDSLMKDLTERHLLGKTVASMYTVEFQKRGLPHAHILLFMAANSKLPTADDID 478
Query: 555 SVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDS 614
+I AE+P+ P LY+ + N M+HGPCG T+SPCM +G+CSK +PK E T +
Sbjct: 479 KIISAEIPNKDKEPELYEVIKNSMMHGPCGSANTSSPCMVDGQCSKLYPKKHQEITKVGA 538
Query: 615 DGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYI 674
DGYP+YRRR T + GV DN +VVPYN KL ++YQAHIN+E+CN++ IKYLFKYI
Sbjct: 539 DGYPIYRRRLTDDYIEKGGVKCDNRYVVPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYI 598
Query: 675 NKGVDRVTMSMS-VTQSSNEETTV-----------VDEIKQYHDCRYLSPCEAVWRTFSF 722
NKG DRV + + ++++ +TT DEIK + DCRY+S EA+WR F F
Sbjct: 599 NKGPDRVVFIVEPIKEATSSDTTAPVVESDTTEKKKDEIKDWFDCRYVSASEAIWRIFKF 658
Query: 723 YIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLG--- 779
I + PV K S+H KQ F+ +A + DVLER + S FLAW+ N K +G
Sbjct: 659 PIQHRSTPVQKLSFHDKGKQPAYFDAKAKMADVLERVSNEDSQFLAWLTLNRKNAVGKNG 718
Query: 780 ---RGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCT 836
R YAE P+ F +D + K++ R +G +GR+N+V + YYLR+LLN+ RG
Sbjct: 719 KRARDCLYAEIPAYFTWDGENKQFKKRTRGFSLGRINYVSRKMEDEYYLRVLLNIVRGPQ 778
Query: 837 SFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNA 896
S++D+++VN V+ +++ AC A +L DD+ YI+G+++ + F G Y+R F LL ++
Sbjct: 779 SYDDIKTVNGVVYPSYKLACFARGILDDDQVYINGLIEASQFCFGDYLRNFFSMMLLSDS 838
Query: 897 MADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKD 956
+A P HVW ETW +L++ I+ R N EL + ++ L EIEK+++ NG +L+D
Sbjct: 839 LARPEHVWSETWHLLSEDILIKKRDEFKNQELTLTEAQIQNYTLQEIEKIMLFNGATLED 898
Query: 957 FDNMPCVDSNILIQYGNILLFNELNFDT-VEMSKLHVECLNKLNGGQAKIYEEIISAVNS 1015
++ P S I N L+ +EL ++ ++ K H + + KL Q IY++I +AV +
Sbjct: 899 IEHFP-KPSREGIDNSNRLIIDELRYNNQSDLKKKHSDWIQKLTPEQRGIYDQITNAVFN 957
Query: 1016 DGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIP 1075
D G FVYG+GGTGKTF+W TL +RS+ +I LNVASSGIASLLL GGRTAHS F IP
Sbjct: 958 DLGGVFFVYGFGGTGKTFIWKTLAAAVRSKGQICLNVASSGIASLLLEGGRTAHSRFSIP 1017
Query: 1076 LNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFN 1135
LN DE S C I S A+L+K +SLIIWDEAPM+S++ FEALD++ DI++ + N
Sbjct: 1018 LNPDEFSVCKIKPKSDLADLIKEASLIIWDEAPMMSKFCFEALDKSFSDIIKRVD----N 1073
Query: 1136 KPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNS-E 1194
K FGGKV+V GGDFRQ+LPVI RAEIVM+++N+S LW CKVL LT+NMRL +N
Sbjct: 1074 KVFGGKVMVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWDHCKVLRLTKNMRLLNNDLS 1133
Query: 1195 TSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY--PEI 1252
+ ++I+ F++W+L +GDG + + NDG+ I +P++ L+Q++ NP+ I R IY P
Sbjct: 1134 VDEAKEIQEFSDWLLAVGDGRVNEPNDGEVIIDIPEELLIQEADNPIEAISREIYGDPTK 1193
Query: 1253 MENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADW 1312
+ + K+++ +AILAP + V+ +NQY+L ER YLS+ S+ ++ ++
Sbjct: 1194 LHEISDPKFFQRRAILAPKNEDVNTINQYMLEHLDSEERIYLSADSI-DPSDSDSLKNPV 1252
Query: 1313 ITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVI 1372
IT +FLN I+ SG+P H L LK GAPVML+RNLD GLCNGTR+ +T L ++V VI
Sbjct: 1253 ITPDFLNSIKVSGMPHHSLRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCSHIVEAKVI 1312
Query: 1373 SGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNP 1432
+G +G+ V+I +++ P+D +P K +RRQFPL ++F MTINKSQG++L VGLYLP P
Sbjct: 1313 TGDRIGQIVYIPLINITPSDTKLPFKMRRRQFPLSVAFVMTINKSQGQSLEQVGLYLPKP 1372
Query: 1433 VFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
VFSHGQLYVA+SRV ++ GLKILI +++ + T N+VFKEVFQ I
Sbjct: 1373 VFSHGQLYVALSRVTSKTGLKILILDKEGKIQKQTTNVVFKEVFQNI 1419
>At1g35940 hypothetical protein
Length = 1678
Score = 1142 bits (2953), Expect = 0.0
Identities = 616/1444 (42%), Positives = 872/1444 (59%), Gaps = 121/1444 (8%)
Query: 66 YFDLGEMNMACQYCGAILWYHERAQKAKNAISPDFSICCMKGKISLPYLQDAPTLLSNLL 125
Y D G+ C+YCGA++WY ER +K + F++CC +G + LP+L+++P LL NLL
Sbjct: 325 YLDHGDPTYKCKYCGAMMWYDERIRKKETNKESVFTLCCGEGSVKLPFLKESPHLLKNLL 384
Query: 126 TNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAE 185
+ P S H+ DN R +N +FA TS GGKVD S+ G+GP F + G NYH IGSL
Sbjct: 385 SGNHPLSKHYRDNARIFNMVFAMTSFGGKVDKSMPKGRGPAMFRLQGGNYHLIGSLKLTP 444
Query: 186 GDNPKFAQLNIYDTRNEFENRLNHLSD------STGKCSLNSDLVVELMAMVDEFNVLAK 239
GD K++QL I DT NE ENR ++ ++GK +L+ +L+ ++ M++ N +
Sbjct: 445 GDYAKYSQLYIIDTENEVENRATVINKGKNAKPASGKPNLDKNLIEAIIKMLNRCNPYVR 504
Query: 240 SFRRVRDHVQEANSNQVALRLFRHRVN-DPKTYNLPTVDEVAALIVEDFDTSDCGRDIIL 298
FR R+ +Q + +R+ R + +TY++PT EVAALI DF RDI++
Sbjct: 505 RFRTARERIQTNDEEPFHMRIIADRQGVEGRTYSMPTTSEVAALIPGDFRHGMPDRDIVI 564
Query: 299 -RTSSGNLQRIYDTHSSFLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYK-RTTISLRE 356
+ S+G+L+RI H S+L LQYPLIF YGE+GF G + + S K + IS+R+
Sbjct: 565 GKKSNGHLKRINQIHISYLALQYPLIFCYGEDGFRP--GIEKCSKSKSKKKNKKCISMRQ 622
Query: 357 WVAFRLQERQFECKRITLSRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEA 416
W AFR+QER+ EC+ + S+RL QQ + D Y+ IES RL Y++ NQ +R + + ++EA
Sbjct: 623 WFAFRIQEREVECQTLLRSKRLFQQCLCDAYTTIESNRLNYIKFNQSKLRCENYTSVKEA 682
Query: 417 IHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEI 476
G T + G+++++P+SFTGG RYM + DAM IC+ YG+PDLF+T TCNPKWPEI
Sbjct: 683 AAAGATTMEEEGNQLLIPASFTGGPRYMVQSYYDAMAICKHYGFPDLFITFTCNPKWPEI 742
Query: 477 ERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAH 536
R+ R L+ DRP++ R+F++KLD LM +L K G+ ++ MYT+EFQKRGLPHAH
Sbjct: 743 TRYYDKRGLNPEDRPNIIARIFKIKLDSLMNDLTVKKMLGKTVASMYTVEFQKRGLPHAH 802
Query: 537 ILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNG 596
ILL++ K KL + + ID +I AE+PD + P LY+ + N M+HGPCG NSPCM +G
Sbjct: 803 ILLFMHAKSKLPTADDIDKLISAEIPDKEKEPDLYEVIKNSMIHGPCGSANVNSPCMVDG 862
Query: 597 RCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHI 656
CSK +PK + T SDGYP+YRRR T + G+ DN +VVPYN KL ++Y AHI
Sbjct: 863 ECSKLYPKKHQDITKIGSDGYPIYRRRKTDDYVEKGGIKCDNRYVVPYNKKLSLRYNAHI 922
Query: 657 NIEYCNKSNCIKYLFKYINKGVDRVTMSMSVTQSS---NEETTVVD---------EIKQY 704
N+E+CN++ IKYLFKYINKG D+V + TQ + + ET + EIK +
Sbjct: 923 NVEWCNQNGSIKYLFKYINKGPDKVVFIVEPTQQTTAGDSETPQQEQGSAEKKKNEIKDW 982
Query: 705 HDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKS 764
DCRY+S EAVWR F + I PV K S+H+ KQ F+ ++ I+DVLER S
Sbjct: 983 FDCRYVSASEAVWRIFKYPIQHISTPVQKLSFHVEGKQPAYFDPKSNIEDVLERVANVDS 1042
Query: 765 MFLAWMEANCKYPLG------RGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPG 818
F+AW+ N + +G R YAE P+ F +D + K + R +G IGR+++V
Sbjct: 1043 QFMAWLTLNRRNAVGKNGKRARECLYAEIPAYFTWDGENKSFKKRTRGFSIGRIHYVSRK 1102
Query: 819 SGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATF 878
+ Y+LR+LLN+ RG TS+ ++++ + V+ TF++AC A +L DD+ +IDG+++
Sbjct: 1103 MEDEYFLRVLLNIVRGPTSYAEIKTYDGVVYKTFKEACFARGILDDDQVFIDGLVEA--- 1159
Query: 879 ASGSYIRELFVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQL 938
+HV +TW +LA+ I+ R N +L + +K
Sbjct: 1160 ----------------------THVRSQTWHLLAEDILKTKRDEFKNPDLTLTETEIKNY 1197
Query: 939 CLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKL 998
L EIEK+++ NG +L+D D P E L
Sbjct: 1198 TLQEIEKIMLSNGATLEDIDEFP--------------------------KPTRDEWKQML 1231
Query: 999 NGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIA 1058
Q +Y I AV ++ G FVYG+GGTGKTF+W TL+ +R +I+LNVASSGIA
Sbjct: 1232 TPEQRGVYNAITEAVFNNLGGVFFVYGFGGTGKTFIWKTLSAAIRCRGQIVLNVASSGIA 1291
Query: 1059 SLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEAL 1118
SLLL GGRTAHS F IPLN DE S +L
Sbjct: 1292 SLLLEGGRTAHSRFGIPLNHDEFSV---------------------------------SL 1318
Query: 1119 DRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFC 1178
D++ DI++ N NK FGGKVVV GGDFRQ+LPVI RAEIVM+++N+S LW C
Sbjct: 1319 DKSFSDIIKNTN----NKVFGGKVVVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWDHC 1374
Query: 1179 KVLTLTENMRLFSNS-ETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQS 1237
KVL LT+NMRL +N+ ++ ++I+ F++W+L + DG + + NDG A I +P+D L+ +
Sbjct: 1375 KVLKLTKNMRLLANNLSATEAKEIQEFSDWLLAVSDGRINEPNDGVATIDIPEDLLITNA 1434
Query: 1238 SNPVADIVRAIY--PEIMENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLS 1295
P+ I IY P+I+ + K+++ +AILAP + V+ +N+Y+L ER YLS
Sbjct: 1435 DKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIYLS 1494
Query: 1296 SYSVWSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGT 1355
+ S+ T+ + IT +FLN I+ G+P+H L LK GAPVML+RNLD GLCNGT
Sbjct: 1495 ADSI-DPTDSDSLNNPVITPDFLNSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCNGT 1553
Query: 1356 RIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTIN 1415
R+ +T L +V VI+G +G V I ++L PTD +P K +RRQFPL ++FAMTIN
Sbjct: 1554 RLQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTIN 1613
Query: 1416 KSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEV 1475
KSQG++L H+GLYLP PVFSHGQLYVA+SRV ++ GLKILI ++D + T N+VFKEV
Sbjct: 1614 KSQGQSLEHIGLYLPKPVFSHGQLYVALSRVTSKKGLKILILDKDGKLQKQTTNVVFKEV 1673
Query: 1476 FQRI 1479
FQ I
Sbjct: 1674 FQNI 1677
>At3g30560 hypothetical protein
Length = 1473
Score = 1120 bits (2896), Expect = 0.0
Identities = 601/1399 (42%), Positives = 860/1399 (60%), Gaps = 71/1399 (5%)
Query: 101 SICCMKGKISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVN 160
S CCM+G+I LP L+++P + L T+ P + +F N+R YN +F+FTS+GGKVD SV
Sbjct: 126 SRCCMQGQIVLPMLKESPEYMWWLFTSDHPDAKNFRANVRPYNMLFSFTSLGGKVDRSVK 185
Query: 161 NGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEFENRLNHLSDSTG----- 215
G+GP F + G+NYH I +L P GD KF QL + DT NE +NR+ +S
Sbjct: 186 KGRGPSMFALQGENYHLIDALKPKPGDYAKFQQLYVMDTDNEVDNRIEVMSKGNNGKTEG 245
Query: 216 -KCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQ-EANSNQVALRLFRHRVNDPKTYNL 273
K + V L+ M+DE N +FR RD E +R+ +R D + YNL
Sbjct: 246 VKQKFKKETVSALLKMLDEINPHVANFRIARDRFNIEKEDANFHMRIISNRDTDGRVYNL 305
Query: 274 PTVDEVAALIVEDFDTSDCGRDIILRTSSGNLQRIYDTHSSFLPLQYPLIFPYGEEGFSD 333
P+V EVA LI DFD + RDI+L+ S L+RI++ H S+L LQYPL+FP GE+G+
Sbjct: 306 PSVGEVA-LIPGDFDDNLDKRDIVLQIKSEKLRRIHECHVSYLSLQYPLLFPKGEDGYRL 364
Query: 334 EIGFDGINYDSSIYKRTTISLREWVAFRLQERQFECKRITLSRRLLQQLVVDCYSMIESQ 393
I N K+ +S+R+W +RLQER+ E + S+RLLQQ +VD ++MIES
Sbjct: 365 GIKKTETNTSKRKKKQKDVSMRQWFDYRLQERKDEKHILLRSKRLLQQFIVDAFTMIESN 424
Query: 394 RLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMG 453
RL +++ NQ +R +++A GD DLS G +++P SFTGG YM N DAMG
Sbjct: 425 RLRFIKKNQTKLRSTNKQAVQDASDAGDNDLSNKGKSVIIPPSFTGGPAYMQQNYLDAMG 484
Query: 454 ICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGK 513
C+ +G+PDLF+T TCN KWP+I R V R L+A D+PD+ CR+++MKL+ LM +L K
Sbjct: 485 TCKHFGFPDLFITFTCNSKWPKITRFVKERKLNAEDKPDIICRIYKMKLENLMDDLTKNH 544
Query: 514 FFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQC 573
FG+ YTIEFQKRGLPHAHI++W+ + K + +D +I AE+PD + P LYQ
Sbjct: 545 IFGKTKLATYTIEFQKRGLPHAHIIVWMDPRYKFPIADHVDKIIFAEIPDKENDPKLYQA 604
Query: 574 VSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRG 633
VS M+HGPC + NSPCM+NG+CSK++PKN VE T D++GYP+YRRR+TG +
Sbjct: 605 VSECMIHGPCRLVNPNSPCMENGKCSKYYPKNHVENTFLDNEGYPIYRRRDTGRFIKKNK 664
Query: 634 VDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVDRVTMSMS------V 687
DN +VVPYN LL KY+AHIN+E+CN+S +KYLFKY+NKG DRVT+S+ V
Sbjct: 665 YPCDNRYVVPYNDFLLRKYRAHINVEWCNQSVSVKYLFKYVNKGPDRVTVSLEPHRKEVV 724
Query: 688 TQSSNEETTVVD-----EIKQYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQ 742
++ +N T D +++ Y DCRY+S CEA+WR + IH + V K ++H KQ
Sbjct: 725 SEENNVGETNNDPQEQNQVEDYFDCRYVSACEAMWRIKGYPIHYRQTLVTKLTFHEKGKQ 784
Query: 743 VILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLGRGLTYAEFPSSFVYDKKKKEWHP 802
I E + VL R ++ F AW E N + P L Y + P+ + ++ K K +
Sbjct: 785 PIYVKEGETAESVLYRVNDDETQFTAWFELNKRDPEAAKLLYEQIPNFYTWNGKDKNFRR 844
Query: 803 RK-KGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSL 861
RK G +GR+N VPP + Y+LR+L+N RG F+D++++ VH T+R AC AL L
Sbjct: 845 RKMPGFVVGRINHVPPKIDDAYHLRILINNIRGPKGFDDIKTIEGVVHKTYRDACYALGL 904
Query: 862 LKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRETWSVLADGIVHALRR 921
L DD++YI+GI + + Y+R+LFV L+ +++ P+ VW TW +L++ +R
Sbjct: 905 LDDDKDYINGIEEANFWCFHKYVRKLFVIMLIFESLSSPAVVWEHTWKILSEDFQRKVR- 963
Query: 922 THNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELN 981
+ LK IE RSL++ +P + + N L+ +E N
Sbjct: 964 -----------DKLKCPAAKNIETTW---NRSLEE-KMLPQLKPGDEPAF-NQLILDERN 1007
Query: 982 FDTVEMSKLHVECLNKLNGGQAKIYEEIISAV-NSDGGEFLFVYGYGGTGKTFLWTTLTY 1040
++ + +H + L L Q K+Y++I+ AV N+ GG+
Sbjct: 1008 YNRETLKTIHDDWLKMLTTEQKKVYDKIMDAVLNNKGGD--------------------- 1046
Query: 1041 KLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSS 1100
I LNVASSGIASLLL GGRTAHS F IPL E S C + +GS AEL+ +
Sbjct: 1047 -------ICLNVASSGIASLLLEGGRTAHSRFGIPLTPHETSTCNMERGSDLAELVTAAK 1099
Query: 1101 LIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGS 1160
LIIWDEAPM+S+Y FE+LD++L+DI + PE + PFGGK+++ GGDFRQILPVI
Sbjct: 1100 LIIWDEAPMMSKYCFESLDKSLKDI--LSTPE--DMPFGGKLIIFGGDFRQILPVILAAG 1155
Query: 1161 RAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSDVEKIKVFAEWVLDIGDGVLGDYN 1220
R IV +++NSS LW++CKV LT+NMRL + + ++ +I+ F++W+L +G+G L N
Sbjct: 1156 RELIVKSSLNSSHLWQYCKVFKLTKNMRLLQDIDINEAREIEDFSKWILAVGEGKLNQPN 1215
Query: 1221 DGDADIKVPKDTLVQQSSNPVADIVRAIYPEIMENVGCAKYYEDKAILAPTLDAVDLVNQ 1280
DG I++ D L+ + NP+ I++A+Y + K+++D+AIL PT D V+ +N
Sbjct: 1216 DGVTQIQIRDDILIPEGDNPIESIIKAVYGTSFDEERDPKFFQDRAILCPTNDDVNSIND 1275
Query: 1281 YVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVM 1340
++LS G E+ Y SS S+ ++ + T +FLN I+ SG+P+H L LK G PVM
Sbjct: 1276 HMLSKLTGEEKIYRSSDSI-DPSDTRADKNPVYTPDFLNKIKISGLPNHLLWLKVGCPVM 1334
Query: 1341 LMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQ 1400
L+RNLD GL NGTR+ + L +V G +++GT VG+ V I RM L P+D +P K +
Sbjct: 1335 LLRNLDSHGGLMNGTRLQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFKMK 1394
Query: 1401 RRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNED 1460
RRQFPL ++FAMTINKSQG++L +VG+YLP PVFSHGQLYVA+SRVK++ GLK+LI +
Sbjct: 1395 RRQFPLSVAFAMTINKSQGQSLGNVGIYLPKPVFSHGQLYVAMSRVKSKGGLKVLITDSK 1454
Query: 1461 ISQRDVTKNIVFKEVFQRI 1479
Q++ T N+VFKE+F+ +
Sbjct: 1455 GKQKNETTNVVFKEIFRNL 1473
>At1g54430 hypothetical protein
Length = 1639
Score = 1041 bits (2693), Expect = 0.0
Identities = 590/1384 (42%), Positives = 822/1384 (58%), Gaps = 93/1384 (6%)
Query: 111 LPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQFVI 170
LP P +L LLT S H+ NIR+YNS+ AFTS+G ++D +V + GP F I
Sbjct: 320 LPPEPQPPQMLKKLLTE----SPHYQRNIRTYNSILAFTSMGAQIDKNVMHKGGPFTFRI 375
Query: 171 SGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEFENRLNHLSDSTGKCSLNSDLVVELMAM 230
GQN+H++GSL+P EG PK QL I+DT NE NR++ + +T LN +V +L+
Sbjct: 376 HGQNHHKLGSLVPEEGKPPKILQLYIFDTANEVHNRISAVKRTTKVGELNEKIVKDLITT 435
Query: 231 VDEFNVLAKSFRRVRDHVQEANSNQVALRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTS 290
VD FN LAK FR+ RD + + + +++L + K Y++PT DE+A LIV DF +
Sbjct: 436 VDTFNCLAKVFRKARDRYEAGDCPEFSIKLIGQKKKG-KQYDMPTTDEIAGLIVGDFSKN 494
Query: 291 DCGRDIILRTSSGNLQRIYDTHSSFLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYKRT 350
RD+I+ S LQ+I D H F+ LQYPL+FPYGE GF + I + K
Sbjct: 495 IGERDVIVHHKSSGLQQISDLHPLFMTLQYPLLFPYGEIGFHEGIPV--------VEKGM 546
Query: 351 TISLREWVAFRLQERQFECKRITLSRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFL 410
TI S+RLL Q +VD Y+ IE +RL + R NQ+ +R D
Sbjct: 547 TI--------------------VRSKRLLHQYIVDAYTSIEQERLRWYRLNQKKLRADQY 586
Query: 411 SGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCN 470
+ +++A+ RGDTD +G R++LP+S+TG RYM DAM ICR YG PDLF+TMT N
Sbjct: 587 NNVKDAVARGDTDAKSIGKRVILPASYTGSPRYMVEKYHDAMAICRWYGNPDLFITMTTN 646
Query: 471 PKWPEIERHVSARCLSAYD-RPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQK 529
PKW EI H+ A + RPD+ CRVF++KLD+L+ + KG FF + I+ +YTIEFQK
Sbjct: 647 PKWEEISEHLKTYGNDAANVRPDIECRVFKIKLDELLADFNKGLFFPKPIAIVYTIEFQK 706
Query: 530 RGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTN 589
RGLPHAHILLWL K + ID I AE+P P + V +M+HGPCG +R +
Sbjct: 707 RGLPHAHILLWLQGDLKKPTPNDIDKYISAEIPVKDKDPEGHTLVEQHMMHGPCGKDRPS 766
Query: 590 SPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLL 649
SPCM+ G CSK FP+ FV T + G+ +YRRRN + LDN FVVP+N ++L
Sbjct: 767 SPCMEKGICSKKFPREFVNHTKMNESGFILYRRRNDQRYVLKGQTRLDNRFVVPHNLEIL 826
Query: 650 MKYQAHINIEYCNKSNCIKYLFKYINKGVDRVTMSMSVTQSSNE-------ETTVVDEIK 702
KY+AHIN+E+CNKS+ IKYLFKYI KGVD+ T + S N E +EI
Sbjct: 827 KKYKAHINVEWCNKSSAIKYLFKYITKGVDKATFIIQKGNSVNGQGSGNGFEEKPRNEIN 886
Query: 703 QYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVK 762
+Y DCRYLS CEA+WR F F IH PPV + HLP +Q +F E +++V R +
Sbjct: 887 EYLDCRYLSACEAMWRIFMFNIHHHNPPVQRLPLHLPGEQSTIFEEEENLENVEYRYGHE 946
Query: 763 KSMFLAWMEANCKYPLGRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGEL 822
++M + E N R L Y + P+ FV+D K + RK+ IGR+ + P +G+L
Sbjct: 947 RTMLTEYFELNKICEDARKLKYVQVPTMFVWDSTNKMYTRRKQRENIGRIVNILPTAGDL 1006
Query: 823 YYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGS 882
YYLR+LLN +G TSF+ L++V VH++F+ AC LL D+E+ D + + A +++
Sbjct: 1007 YYLRILLNKVKGATSFDYLKTVGGVVHESFKAACHTRGLLDGDKEWHDAMDEAAQWSTSY 1066
Query: 883 YIRELFVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLME 942
+R LFV L+ +++P +W W +AD + ++ N +L + E L++ L+E
Sbjct: 1067 LLRSLFVLILIYCEVSEPLKLWSHCWESMADDVFRKQQKVLNFPQLELKAEELEKYTLIE 1126
Query: 943 IEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQ 1002
IE LL + +SL D+ MP + NKLN Q
Sbjct: 1127 IETLLRQHEKSLSDYPEMPQPE-------------------------------NKLNEQQ 1155
Query: 1003 AKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLL 1062
IY++++ +V + G+ F+YG GGTGKTFL+ T+ LRS K ++ VASS IA+LLL
Sbjct: 1156 RIIYDDVLKSVINKEGKLFFLYGAGGTGKTFLYKTIISALRSNGKNVMPVASSAIAALLL 1215
Query: 1063 HGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTL 1122
GGRTAHS F IP+N EDS C I GS A +L LIIWDEAPM R+ FEA+DRTL
Sbjct: 1216 PGGRTAHSRFKIPINVHEDSICDIKIGSMLANVLSKVDLIIWDEAPMAHRHTFEAVDRTL 1275
Query: 1123 RDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLT 1182
RDI+ + + + K FGGK V+LGGDFRQILPVIP+G+R E V A IN S LW C
Sbjct: 1276 RDILSVGDEKALTKTFGGKTVLLGGDFRQILPVIPQGTRQETVSAAINRSYLWESCHKYL 1335
Query: 1183 LTENMRLFSNSETSDVEKIKVFAEWVLDIGDGVLG------DYNDGDADIKVPKDTLVQQ 1236
L++NMR+ E+IK FAEW+L +GDG D + + +I + K+ L+ +
Sbjct: 1336 LSQNMRV-------QPEEIK-FAEWILQVGDGEAPRKTHGIDDDQEEDNIIIDKNLLLPE 1387
Query: 1237 SSNPVADIVRAIYPEIMENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSS 1296
+ NP+ + R+++P+ + + A+L P + VD +N Y+LS PG + Y S+
Sbjct: 1388 TENPLEVLCRSVFPDFTNTFQDLENLKGTAVLTPRNETVDEINDYLLSKVPGLAKEYFSA 1447
Query: 1297 YSV---WSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCN 1353
S+ ++TE+ G E + E+LN + G+P H+L LK G P+ML+RNL+ GLCN
Sbjct: 1448 DSIDRDEALTEE-GFEMSY-PMEYLNSLEFPGLPAHRLCLKVGVPIMLLRNLNQKEGLCN 1505
Query: 1354 GTRIVVTHLRPNVVGGIVISGTHVGR-QVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAM 1412
GTR++VTHL V+ ++S T R +V I R+ L P D P +RRQFP+ + +AM
Sbjct: 1506 GTRLIVTHLGDKVLKAEILSDTTKERKKVLIPRIILSPQDSKHPFTLRRRQFPVRMCYAM 1565
Query: 1413 TINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVF 1472
TINKSQG+TL+ V LYLP PVFSHGQLYVA+SRV + GL +L ++ + VT NIV+
Sbjct: 1566 TINKSQGQTLNRVALYLPKPVFSHGQLYVALSRVTSPKGLTVLDTSKKKEGKYVT-NIVY 1624
Query: 1473 KEVF 1476
+EVF
Sbjct: 1625 REVF 1628
>At2g05080 putative helicase
Length = 1219
Score = 1018 bits (2632), Expect = 0.0
Identities = 555/1250 (44%), Positives = 772/1250 (61%), Gaps = 78/1250 (6%)
Query: 258 LRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTSDCGRDIILRTSSGNLQRIYDTHSSFLP 317
+R+ R D + YN+PT EVA LI DF RDII+ SG LQRI + +LP
Sbjct: 1 MRIVSKRETDGRVYNVPTTSEVAMLIPGDFTIDIPCRDIIVEEKSGKLQRISEILPCYLP 60
Query: 318 LQYPLIFPYGEEGFSDEIGFD--GINYDSSIYKRTTISLREWVAFRLQERQFECKRITLS 375
LQYPL+FPYGE+GF I G D K IS+R+W AFR+ ER+ E + S
Sbjct: 61 LQYPLLFPYGEDGFRTGIEKHQTGAGKDK---KNKFISIRQWFAFRIHERKHEKHILLRS 117
Query: 376 RRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPS 435
+RL QQ +VD Y IES RL Y++ NQ ++R D + +++A G DL G LP+
Sbjct: 118 KRLWQQFLVDSYIAIESNRLGYIKLNQSSLRADNYNSVQKASEEGKCDLKYQGLACYLPA 177
Query: 436 SFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLAC 495
+FTGG RYM N DAM +C+ +G+PD F+T TCNPKWPE+ R R L DRP++ C
Sbjct: 178 TFTGGPRYMRNMYLDAMAVCKHFGFPDYFITFTCNPKWPELIRFCGERNLRVDDRPEIIC 237
Query: 496 RVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDS 555
++F+MKLD LM +L K G+ + MYTIEFQKRGLPHAHIL+WL +K KL E ID
Sbjct: 238 KIFKMKLDSLMLDLTKRNILGKTSTSMYTIEFQKRGLPHAHILIWLDSKCKLTRAEHIDK 297
Query: 556 VICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSD 615
I AE+PD P L++ + MVHGPCGV PCM+NG+CSKF+PK+ V KT D +
Sbjct: 298 AISAEIPDKLKDPELFEVIKEMMVHGPCGVVNPKCPCMENGKCSKFYPKDHVPKTIIDKE 357
Query: 616 GYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYIN 675
G+P+YRRR ++ DN +V+PYN L ++Y+AHIN+E+CN+S +KY+FKYI+
Sbjct: 358 GFPIYRRRRIDDFVQKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIH 417
Query: 676 KGVDRVTMSMSVTQSS---------------NEETTVVDEIKQYHDCRYLSPCEAVWRTF 720
KG DRVT+ + + +S E +E++ + +CRY+S CEA WR
Sbjct: 418 KGPDRVTVVVGSSLNSKNKEKGKQKVNADTDGSEPKKKNEVEDFFNCRYVSACEAAWRIL 477
Query: 721 SFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLGR 780
+ IH + V+K S+HLP +Q I F ++ VL + + S+ +A R
Sbjct: 478 KYPIHYRSTSVMKLSFHLPGEQYIYFKGDEEVETVLNKADLDGSIQIA-----------R 526
Query: 781 GLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFED 840
LTY P+ F YD K+K+++ RKKG IGR+N+VP + YYLR+LLNV G SFE+
Sbjct: 527 KLTYPNIPTRFTYDPKEKKFNLRKKGFAIGRINYVPRDIEDGYYLRILLNVVPGPRSFEE 586
Query: 841 LRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADP 900
L++VN ++ ++ AC AL LL +D+EYID + + ++SG Y+R+LFV L +A+ P
Sbjct: 587 LKTVNGVLYKEWKDACEALGLLDNDQEYIDDLKRTSFWSSGWYLRQLFVIML--DALISP 644
Query: 901 SHVWRETWSVLADGIVHALRRTHNN------LELVIDNEHLKQLCLMEIEKLLMINGRSL 954
+VW TW L++ I + ++ N +L++ +E K L EI+ +L NG SL
Sbjct: 645 ENVWAATWQHLSEDIQNEKKKYFNRPVTCLFTDLILSDEEKKVYALQEIDHILRRNGTSL 704
Query: 955 KDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVN 1014
+ MP V + N+L+ +E +D +K H + + KL Q +Y+ II AVN
Sbjct: 705 TYYKTMPQVPRDPRFD-TNVLILDEKGYDRESETKKHADSIKKLTLEQKSVYDNIIGAVN 763
Query: 1015 SDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCI 1074
+ G FVYG+GGTGKTFLW TL+ LRS+ I+LNVASSGIASLLL GGRTAHS I
Sbjct: 764 ENVGGVFFVYGFGGTGKTFLWKTLSAALRSKGDIVLNVASSGIASLLLEGGRTAHSRSGI 823
Query: 1075 PLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECF 1134
PLN +E + C + GS +A L+K +SLIIWDEAPM+SR+ FE+LDR+L D IC C
Sbjct: 824 PLNPNEFTTCNMKAGSDRANLVKEASLIIWDEAPMMSRHCFESLDRSLSD---ICG-NCD 879
Query: 1135 NKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSE 1194
NKPFGGKVVV GGDFRQ+LPVIP A+IVMA +NSS LW CKVLTLT+NM LFS
Sbjct: 880 NKPFGGKVVVFGGDFRQVLPVIPGADTADIVMAALNSSYLWSHCKVLTLTKNMCLFSE-- 937
Query: 1195 TSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIYPEI-- 1252
EW+L +GDG +G+ NDG+A I +P + L+ ++ +P+ I IY +I
Sbjct: 938 -----------EWILAVGDGRIGEPNDGEALIDIPSEFLITKAKDPIQAICTEIYGDITK 986
Query: 1253 MENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEAD- 1311
+ +++++AIL PT + V+ +N+ +L G E T+LSS S+ T D+G +
Sbjct: 987 IHEQKDPVFFQERAILCPTNEDVNQINETMLDNLQGEELTFLSSDSL--DTADIGSRNNP 1044
Query: 1312 WITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIV 1371
+T EFLN+++ G+ +HKL LK G+PVML+RN+D GL NGTR+ + + P ++ ++
Sbjct: 1045 VLTPEFLNNVKVLGLSNHKLRLKIGSPVMLLRNIDPIGGLMNGTRLQIMQMSPFILQAMI 1104
Query: 1372 ISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPN 1431
++G D +P + +R Q PL + FAMTINKSQG++L VG++LP
Sbjct: 1105 LTGDR--------------ADTKLPFRMRRTQLPLAVCFAMTINKSQGQSLKRVGIFLPR 1150
Query: 1432 PVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRIYS 1481
P FSH QLYVAISRV +++GLKILI N++ + TK F + F RI+S
Sbjct: 1151 PCFSHSQLYVAISRVTSKSGLKILIVNDEGKPQKQTKK--FTKKFLRIFS 1198
>At2g14470 pseudogene
Length = 1265
Score = 1009 bits (2608), Expect = 0.0
Identities = 571/1392 (41%), Positives = 805/1392 (57%), Gaps = 144/1392 (10%)
Query: 83 LWYHERAQKAKNAISPDFSICCMKGKISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSY 142
+WY ER +K + F++CC +G + LP+L+++P LL NLL+ P S H+ DN R++
Sbjct: 1 MWYDERIRKKETKKESGFTLCCGEGSVKLPFLKESPDLLKNLLSGNHPLSKHYRDNARTF 60
Query: 143 NSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNE 202
N +FA TS+GGKVD S+ G+GP F + G NYH IGSL P GD K++QL I DT NE
Sbjct: 61 NMVFAVTSLGGKVDKSMPKGRGPAMFRLQGGNYHLIGSLKPTPGDYAKYSQLYIVDTENE 120
Query: 203 FENRLNHLSD------STGKCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSNQV 256
ENR + ++GK +L+ +L+ ++ M++ N + FR R+ +Q +
Sbjct: 121 VENRAIVIGKGKIAKPASGKPNLDKNLIEVIIKMLNRCNPYVRKFRTARERIQTNDEEPF 180
Query: 257 ALRLFRHRVN-DPKTYNLPTVDEVAALIVEDFDTSDCGRDIIL-RTSSGNLQRIYDTHSS 314
+R+ R D +TY++ T EVAALI DF RDI++ + S+G+L+RI H S
Sbjct: 181 HMRIIADRQGVDGRTYSMHTTSEVAALIPGDFRHGMPDRDIVIEKKSNGHLKRINQIHIS 240
Query: 315 FLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYKRTTISLREWVAFRLQERQFECKRITL 374
+L LQYPLIF YGE+GF I S + IS+R+W AFR+QER+ EC+ +
Sbjct: 241 YLALQYPLIFCYGEDGFRPGIE-KCFKSKSKKKNKKCISMRQWFAFRIQEREVECQTLLR 299
Query: 375 SRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLP 434
S+RL QQ + D Y+ IES RL Y++ NQ +R + + ++EA G T + G+++++P
Sbjct: 300 SKRLFQQFLCDAYTTIESNRLNYIKFNQSKLRCENYTSVKEAAAAGATTMEEEGNQLLIP 359
Query: 435 SSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLA 494
+SF GG RYM + DAM IC+ YG+PDLF+T TCNPKWP I R+ R L+ DR D+
Sbjct: 360 ASFNGGPRYMVQSYYDAMAICKLYGFPDLFITFTCNPKWPHITRYCDKRGLNPKDRLDII 419
Query: 495 CRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELID 554
R+F++KLD LM +L K G+ ++ MYT+EFQKRGLPHAHILL++ K KL + + ID
Sbjct: 420 ARIFKIKLDSLMNDLTVKKMLGKTVASMYTVEFQKRGLPHAHILLFMHAKSKLPTSDDID 479
Query: 555 SVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDS 614
+I AE+PD + P LY+ + N M+HGPCG SPCM +G CSK +PK + T S
Sbjct: 480 KLISAEIPDKEKEPELYEVIKNSMIHGPCGSANVKSPCMVDGECSKLYPKKHQDITKVGS 539
Query: 615 DGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYI 674
DGYP+YRRR + G+ DN +V+PYN K ++Y AHIN+E+CN+++ IKYLFKYI
Sbjct: 540 DGYPIYRRRKIDDYVEKGGIKCDNRYVMPYNKKFSLRYNAHINVEWCNQNDSIKYLFKYI 599
Query: 675 NKGVDRVTMSMSVTQSSNEETTVVDEIKQYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQ 734
NKG D+V + TQ + D P +Q
Sbjct: 600 NKGPDKVIFIVEPTQQATAG-------------------------------DSETPQQEQ 628
Query: 735 SYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLGRGLTYAEFPSSFVYD 794
KK NE D RN V K+ A R YAE P+ F +D
Sbjct: 629 RSAEKKK-----NEIKDWFDC-RRNAVGKNGKRA-----------RECLYAEIPAYFTWD 671
Query: 795 KKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQ 854
+ K + R +G IGR+++V + Y+LR+LLN
Sbjct: 672 GENKAFKKRTRGFSIGRIHYVSRKMEDDYFLRVLLN------------------------ 707
Query: 855 ACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRETWSVLADG 914
+S+L +E F LL ++++ P+HVW +TW +LA+
Sbjct: 708 ----ISVL-----------------FWRLSQEFFAMLLLSDSLSRPAHVWSQTWHILAED 746
Query: 915 IVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMP--CVDSNILIQYG 972
I+ R N E D D P +D I
Sbjct: 747 ILKKKRDEFKNPE----------------------------DIDEFPKPTIDG---IDNS 775
Query: 973 NILLFNELNFDTVE-MSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGK 1031
N L+ EL ++ + + H E L Q +Y EI AV ++ G FVYG+GGTGK
Sbjct: 776 NRLIVEELRYNRESNLKEKHEEWKQMLTPEQRGVYNEITEAVFNNLGGVFFVYGFGGTGK 835
Query: 1032 TFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSP 1091
TF+W TL+ +R +I+LNVASSGIASLLL GGRTAHS F IPLN DE S C I S
Sbjct: 836 TFIWKTLSATIRYRDQIVLNVASSGIASLLLEGGRTAHSRFGIPLNPDEFSVCKIKPKSD 895
Query: 1092 KAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQ 1151
A L+K +SL+IWDEAPM+SR+ FEALD++ DI++ + N FGGKVVV GGDFRQ
Sbjct: 896 LANLVKKASLVIWDEAPMMSRFCFEALDKSFSDIIKNTD----NTVFGGKVVVFGGDFRQ 951
Query: 1152 ILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNS-ETSDVEKIKVFAEWVLD 1210
+ PVI RAEIVM+++N+S LW CKVL LT+N RL +N+ ++ ++I+ F++W+L
Sbjct: 952 VFPVINGAGRAEIVMSSLNASYLWDNCKVLKLTKNTRLLANNLSETEAKEIQEFSDWLLA 1011
Query: 1211 IGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY--PEIMENVGCAKYYEDKAIL 1268
+GDG + + NDG A I +P+D L+ + P+ I IY P+I+ + K+++ +AIL
Sbjct: 1012 VGDGRINESNDGVAIIDIPEDLLITNADKPIESITNEIYGDPKILHEITDPKFFQGRAIL 1071
Query: 1269 APTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEFLNDIRCSGIPD 1328
A + V+ +N+Y+L ER YLS+ S+ T+ + IT +FLN I+ G+P+
Sbjct: 1072 ASKNEDVNTINEYLLDQLHAEERIYLSADSI-DPTDSDSLSNPVITPDFLNSIKLPGLPN 1130
Query: 1329 HKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDL 1388
H L LK GAPV+L+RNLD GLCNGTR+ +T L +V VI+G +G + I ++L
Sbjct: 1131 HSLRLKVGAPVLLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIGHIILIPTVNL 1190
Query: 1389 MPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKT 1448
PT+ +P K +RRQFPL ++F MTINKS+G++L HVGLYLP PVFSHGQLYVA+SRV +
Sbjct: 1191 TPTNTKLPFKMRRRQFPLSVAFVMTINKSEGQSLEHVGLYLPKPVFSHGQLYVALSRVTS 1250
Query: 1449 RAGLKILICNED 1460
+ GLKILI ++D
Sbjct: 1251 KKGLKILILDKD 1262
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 855 bits (2210), Expect = 0.0
Identities = 493/1260 (39%), Positives = 704/1260 (55%), Gaps = 134/1260 (10%)
Query: 181 LLPAEGDNPKFAQLNIYDTRNEFENRLNHLSDSTGKCSLNSDLVVELMAMV--DEFNVLA 238
+LP ++P++ + + +N L +SD T C + LV + + D ++
Sbjct: 102 VLPMLKESPEYMWWLLTSDHPDAKN-LEQISDHTTCCFHSHLLVARWIDQLKRDVVHLCL 160
Query: 239 KSFRRVRDHVQEANSNQVALRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTSDCGRDIIL 298
+ R +EAN + +R+ R D + YNLP+V EVAALI DFD + +DI+L
Sbjct: 161 IAHDRFNIEKEEANFH---MRIISKRETDGRVYNLPSVAEVAALIPGDFDDNLDKKDIVL 217
Query: 299 RTSSGNLQRIYDTHSSFLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYKRTTISLREWV 358
+ SG L+RI++ H YPL+FP GE+G+ +G +S K+ S+R+W
Sbjct: 218 QMKSGKLRRIHECH-------YPLLFPKGEDGY--RLGIKKTPTKTSKGKK---SMRQWF 265
Query: 359 AFRLQERQFECKRITLSRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIH 418
+RLQER+ E + S+RLLQQ + S N Q +++A
Sbjct: 266 DYRLQERKDEKHILLRSKRLLQQFMTKLRST----------NKQ---------AVQDASD 306
Query: 419 RGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIER 478
GD DLS G ++P SFTGG YM N DA+G C+ +G+PDLF+T TCNPKWPEI R
Sbjct: 307 AGDNDLSNKGKSYIIPPSFTGGPAYMQQNYLDAIGTCKHFGFPDLFITFTCNPKWPEITR 366
Query: 479 HVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHIL 538
V R L+A DRPD+ CR+++MKLD LM +L K F MYT+EFQKRGLPHAHI+
Sbjct: 367 FVKERKLNAEDRPDIICRIYKMKLDNLMDDLTKNHIFA-----MYTVEFQKRGLPHAHII 421
Query: 539 LWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRC 598
+W+ + K H+ + +D +I AE+PD + +P LYQ VS M+HGPC + NSPCM+NG+C
Sbjct: 422 VWMDPRYKFHTADHVDKIIFAEIPDKEKHPELYQAVSECMIHGPCRLVNPNSPCMENGKC 481
Query: 599 SKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINI 658
SK++PKN VE TS D GYP+YRRR++G + DN +VVPYN LL KY+AHIN+
Sbjct: 482 SKYYPKNHVENTSLDYKGYPIYRRRDSGRFIEKNKYQCDNWYVVPYNDVLLRKYRAHINV 541
Query: 659 EYCNKSNCIKYLFKYINKGVDRVTMSMSVTQSSNEETTVVDEIKQYHDCRYLSPCEAVWR 718
E+CN+S IKYLFKY+NKG DRVT + N + ++++ Y DCR
Sbjct: 542 EWCNQSVSIKYLFKYVNKGPDRVTQNN--VGEINNDPQERNQVQDYFDCR---------- 589
Query: 719 TFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPL 778
+ IH + V K ++H KQ + E + VL R ++ F+AW E N + P
Sbjct: 590 --GYPIHYRQTSVTKLTFHEKGKQSVYVKEGETAESVLYRVNNDETQFIAWFELNKRDPE 647
Query: 779 GRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSF 838
L Y + P+ + +N VPP + Y+LR+L+N R F
Sbjct: 648 AAKLLYEQIPNFYT-------------------INHVPPKIDDAYHLRILINNIRAPKGF 688
Query: 839 EDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMA 898
+D+++V VH T+R AC AL LL DD+EYI GI + + S Y+R+ FV L+ +++
Sbjct: 689 DDIKTVEGVVHKTYRDACYALGLLDDDKEYIHGIEEANFWCSPKYVRKSFVIMLISESLS 748
Query: 899 DPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFD 958
P VW TW +L + +R ++E+ + + L +
Sbjct: 749 SPVVVWEHTWKILFEDFQRKVRD--------------------KLERPDLWRYKMLLEPG 788
Query: 959 NMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIY-EEIISAVNSDG 1017
+ P N L+ +E N++ + K H L L K+Y +EI+ V +D
Sbjct: 789 DEPAF---------NPLIIDERNYNRESLKKKHDNWLKTLTPEHKKVYHDEIMDDVLNDK 839
Query: 1018 GEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLN 1077
G F+Y +GGTGKTFLW L+ +R + LNVASS IASLLL GGRTAHS F IPL
Sbjct: 840 GGVFFLYAFGGTGKTFLWKVLSAAIRCKGDTCLNVASSSIASLLLEGGRTAHSRFGIPLT 899
Query: 1078 ADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKP 1137
E S C + +GS AEL+ + LIIWDE + P
Sbjct: 900 PHETSTCNMERGSDLAELVTAAKLIIWDE----------------------------DMP 931
Query: 1138 FGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSD 1197
FG KV++ GGDFRQIL VIP R IV +++NSS LW+ CKVL LT+NMRL + + ++
Sbjct: 932 FGRKVILFGGDFRQILHVIPAAGRELIVKSSLNSSYLWQHCKVLKLTKNMRLLQDIDINE 991
Query: 1198 VEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIYPEIMENVG 1257
+I+ F +W+L +G+G L + +DG I++P D L+ + NP+ I++A+Y
Sbjct: 992 AREIEDFLKWILTVGEGKLNEPSDGVTHIQIPDDILIPEGDNPIESIIKAVYGTTFAQKR 1051
Query: 1258 CAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEF 1317
K+++ KAIL PT D V+ +N ++LS G ER Y SS S+ ++ + T +F
Sbjct: 1052 DPKFFQHKAILCPTNDDVNSINDHMLSKLTGEERIYRSSNSI-DPSDTRADKNPIYTPDF 1110
Query: 1318 LNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHV 1377
LN I+ SG+ +H L LK G PVML+RN GL NGTR+ + L +V G +++GT V
Sbjct: 1111 LNKIKISGLANHLLRLKVGCPVMLLRNFYPHGGLMNGTRLQIVRLGDKLVQGRILTGTRV 1170
Query: 1378 GRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHG 1437
G+ V I RM L P+D +P K +RR FPL ++FAMTINKSQG++L +VG+YLP VFSHG
Sbjct: 1171 GKLVIIPRMSLTPSDRRLPFKMKRRHFPLSVAFAMTINKSQGQSLGNVGMYLPKAVFSHG 1230
Score = 47.8 bits (112), Expect = 5e-05
Identities = 21/50 (42%), Positives = 33/50 (66%)
Query: 81 AILWYHERAQKAKNAISPDFSICCMKGKISLPYLQDAPTLLSNLLTNIDP 130
AI WY ER + K + +P ++ CCM+G+I LP L+++P + LLT+ P
Sbjct: 73 AIFWYGERLNRRKKSANPVYTGCCMQGQIVLPMLKESPEYMWWLLTSDHP 122
>At1g52960 hypothetical protein
Length = 924
Score = 843 bits (2178), Expect = 0.0
Identities = 436/927 (47%), Positives = 596/927 (64%), Gaps = 29/927 (3%)
Query: 546 KLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKN 605
KL + E D +I AE+PD K P LY V + M+HGPCGV NSPCM+NG+C K+FPK+
Sbjct: 6 KLSTAEDTDKIITAEIPDKKKKPGLYAVVKDCMIHGPCGVGHPNSPCMENGKCKKYFPKS 65
Query: 606 FVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSN 665
+ + T D+DG+PVYRRR+TGI + G DN +V+PYN K+ ++YQAHIN+E CN+S
Sbjct: 66 YSDTTKVDNDGFPVYRRRDTGIYVEKNGFQCDNRYVIPYNEKVSLRYQAHINVELCNQSG 125
Query: 666 CIKYLFKYINKGVDRVTMSMSVTQSSNEETTV---VDEIKQYHDCRYLSPCEAVWRTFSF 722
IKYLFKY++KG DRVT VT N++ T DE+K Y DCRY+S CEA+WR F F
Sbjct: 126 SIKYLFKYVHKGHDRVT----VTVEPNDQDTAKKEKDEVKDYFDCRYVSACEAMWRIFKF 181
Query: 723 YIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPL---- 778
IH + PV+K +H KQ + + + V++R + + FLAW + N K P
Sbjct: 182 PIHYRTTPVVKLFFHEEGKQPVYYKPGETTESVMDRLSSEATQFLAWFQLNKKPPSRTIR 241
Query: 779 ------------GRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLR 826
L + E P+ F ++ K+K++ R++G IGR+NFVP + YYLR
Sbjct: 242 ANAKKLPKAAPDPTKLLFEEIPNHFTWNSKEKKFMIRERGFAIGRINFVPRTIEDAYYLR 301
Query: 827 MLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRE 886
+LLN++RG TS++DL++V VH +FR A AL LL DD+EYI+GI D + S Y+R
Sbjct: 302 ILLNIKRGVTSYKDLKTVKGVVHKSFRDAVFALGLLDDDKEYINGIKDAKFWCSAKYVRR 361
Query: 887 LFVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKL 946
LFV LL ++ P VW ETW +L++ I R+ +L + +E +Q CL EI +L
Sbjct: 362 LFVIMLLSESLTKPEMVWDETWRILSEDIERRKRKEWKRPDLQLSDEERQQYCLQEIARL 421
Query: 947 LMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIY 1006
L NG SL + MP + S+ ++ N + +E +D + + H E L + Q KIY
Sbjct: 422 LTKNGVSLSKWKQMPQI-SDEHVEKCNHFILDERKYDRAYLIEKHEEWLTMVTSEQKKIY 480
Query: 1007 EEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGR 1066
+EI+ AV D G FVYG+GGTGKTFLW L+ +RS+ I LNVASSGIA+LLL GGR
Sbjct: 481 DEIMDAVLHDRGGVFFVYGFGGTGKTFLWKLLSAAIRSKGDISLNVASSGIAALLLDGGR 540
Query: 1067 TAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIM 1126
T HS F IP+N +E S C I +GS EL+K ++LIIWDE PM+S++ FE+LDRTLRDIM
Sbjct: 541 TTHSRFGIPINPNESSTCNISRGSDLGELVKEANLIIWDETPMMSKHCFESLDRTLRDIM 600
Query: 1127 RICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTEN 1186
NP +KPFGGK +V GGDFRQ+LPVI R EIV A +NSS +W CKVL LT+N
Sbjct: 601 N--NPG--DKPFGGKGIVFGGDFRQVLPVINGAGREEIVFAALNSSYIWEHCKVLELTKN 656
Query: 1187 MRLFSNSETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVR 1246
MRL +N + I+ F++W+LD+GDG + NDG A I +P++ L+ ++PV I+
Sbjct: 657 MRLLANISEHEKRDIEYFSKWILDVGDGKISQPNDGIALIDIPEEFLINGDNDPVESIIE 716
Query: 1247 AIYPEIMENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDV 1306
A+Y K+++ +AIL PT + V+ +N++++S+ G ER YLSS S+ +
Sbjct: 717 AVYGNTFMEEKDPKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSSDSI-DPADTS 775
Query: 1307 GIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNV 1366
D + +FLN +R SG+P+H L LK G PVML+RN+D + GLCNGTR+ VT + V
Sbjct: 776 SANNDAYSADFLNSVRVSGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMADTV 835
Query: 1367 VGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVG 1426
+ I+G VG+ V I RM + P+D +P K +RRQFPL ++FAMTINKSQG+TL VG
Sbjct: 836 IQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRRQFPLSVAFAMTINKSQGQTLESVG 895
Query: 1427 LYLPNPVFSHGQLYVAISRVKTRAGLK 1453
LYLP PVFSHGQLYVAISRV ++ G K
Sbjct: 896 LYLPRPVFSHGQLYVAISRVTSKTGTK 922
>At2g07620 putative helicase
Length = 1241
Score = 820 bits (2119), Expect = 0.0
Identities = 465/1123 (41%), Positives = 644/1123 (56%), Gaps = 108/1123 (9%)
Query: 84 WYHERAQKAKNAISPDFSICCMKGKISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSYN 143
WY ER K + + P FS+CC++G + LP+L ++P L+ LL+ D HF +NIR+YN
Sbjct: 3 WYDERVNKRRKSKKPKFSLCCLQGSVKLPFLTESPELIRELLSCDDALRRHFRENIRAYN 62
Query: 144 SMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEF 203
+F+ TS+GGKVD S G+GP QF + G NYH IGS+LP EGD KF+QL I DT NE
Sbjct: 63 MLFSMTSLGGKVDRSNPQGKGPNQFQLHGANYHLIGSMLPGEGDYAKFSQLYIVDTGNEV 122
Query: 204 ENRLNHLSDSTGKCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSNQVALRLFRH 263
ENR + +T K + DL+ +L+ M+D N ++FR + + N +++
Sbjct: 123 ENRSTNPKKATFKNQVRKDLIEKLIRMLDACNPYIENFRLAKYKLDSNNGEPFYMQIVSD 182
Query: 264 RVN-DPKTYNLPTVDEVAALIVEDFDTSDCGRDIILRTS-SGNLQRIYDTHSSFLPLQYP 321
RV D +TY P EVAALI DF RDII++ +G L RI H S++P+QYP
Sbjct: 183 RVGKDGRTYCNPRTSEVAALIPGDFRPKMHTRDIIVQDKKTGQLSRISKVHPSYVPMQYP 242
Query: 322 LIFPYGEEGFSDEI--GFDGINYDSSIYKRTTISLREWVAFRLQERQFECKRITLSRRLL 379
LIF YGE+ F I G+ G R ++ +C
Sbjct: 243 LIFNYGEDDFRPGIQKGYTG---------------------RTGKQANKC---------- 271
Query: 380 QQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTG 439
++ Y+ IES RL Y++ NQ +R ++ A G+T++S G I +P SFTG
Sbjct: 272 ----INGYTTIESNRLAYIKFNQSNLRCKNYDTVKAAREAGNTNMSEQGKSIRIPQSFTG 327
Query: 440 GRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLACRVFR 499
G R+M N+ DAM IC+ YG+P+LF+T TCNPK PEI + R L+A DRPD+ C +F+
Sbjct: 328 GPRHMLNSYYDAMAICKMYGFPNLFITFTCNPKLPEITSYCKERKLNAEDRPDVVCWIFK 387
Query: 500 MKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICA 559
MKLD LMK+L + G+ T+ FQKRGLPHAHILL++ K + + ID+++ A
Sbjct: 388 MKLDSLMKDLTEDHLLGK------TVMFQKRGLPHAHILLFMDKSCKFPTSDDIDNILSA 441
Query: 560 ELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPV 619
E+PD P LY+ V++ M+HGPCG SPC+ +G+CSKFFPK VE+T+ DGYP+
Sbjct: 442 EIPDKAKDPKLYEVVNDCMIHGPCGAANKESPCIVDGKCSKFFPKKLVEQTTVGKDGYPI 501
Query: 620 YRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVD 679
YRRR + + G+ DN +VVPYN L ++Y+AHIN+E+C +S IKYLFKYINKG D
Sbjct: 502 YRRRESEHFVEKGGIKCDNTYVVPYNRMLSLRYRAHINVEWCKQSGSIKYLFKYINKGQD 561
Query: 680 RVTM--------SMSVTQSSNEE--TTVVD---EIKQYHDCRYLSPCEAVWRTFSFYIHD 726
RV + S V S +++ V+D EIK Y DCRY+S EAVWR F F I
Sbjct: 562 RVAIVVEPKDKTSNMVLFSGSQKLLVAVIDDDKEIKDYFDCRYVSASEAVWRIFKFPIQY 621
Query: 727 KWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLGRGLTYAE 786
+ PV+K SYHLP KQ + F + ID++ E+ + MF+ +++ N + R Y E
Sbjct: 622 RTTPVMKLSYHLPGKQPLCFEDTQNIDELSEKKANEDFMFIGFLKLNQECEFARQFIYTE 681
Query: 787 FPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNN 846
P F +D + K+W R++G IGRMN+ YY+++LL + G S ED+R+ +
Sbjct: 682 IPPYFTWDGQNKQWKLRERGFYIGRMNYASIKMEPEYYMKILLGIVCGPKSDEDIRTYKD 741
Query: 847 HVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRE 906
V F F G R F+ N + D S
Sbjct: 742 VVRRKF-------------------------FFLGIIFRIYFLCCFWINVLLDQSMC--- 773
Query: 907 TWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSN 966
+ LV+ + L+EIE +L+ NG +L+DF +MP +
Sbjct: 774 -----------------GKMLLVLSDAEKINYALLEIEDMLLCNGSTLEDFKHMP-KPTK 815
Query: 967 ILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGY 1026
+ N + E N++ ++ + H + NK+ Q +IY+EI+ AV + G FVYG+
Sbjct: 816 EGTDHSNRFITEEKNYNVEKLKEDHDDWFNKMTSEQKEIYDEIMKAVLENSGGIFFVYGF 875
Query: 1027 GGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGI 1086
GGTGKTF+W TL+ +R + I +NVASSGIA LLL GGRTAHS F IP+N D+ + C I
Sbjct: 876 GGTGKTFMWKTLSAAVRMKGLISVNVASSGIAFLLLQGGRTAHSRFGIPINPDDFTTCHI 935
Query: 1087 VQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLG 1146
V S A +LK +SLIIWDEAPM+SRY FE+LDR+L D++ + KPFGGKVVV G
Sbjct: 936 VPNSDLANMLKEASLIIWDEAPMMSRYCFESLDRSLNDVIGNVD----GKPFGGKVVVFG 991
Query: 1147 GDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRL 1189
GDFRQ+L VI RAEIV+A +NSS LW C VLTLT+NM L
Sbjct: 992 GDFRQVLHVIHGAGRAEIVLAALNSSYLWEHCNVLTLTKNMSL 1034
Score = 173 bits (438), Expect = 8e-43
Identities = 86/188 (45%), Positives = 127/188 (66%), Gaps = 1/188 (0%)
Query: 1292 TYLSSYSVWSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGL 1351
TYLS+ S+ + + T FLN I+ SG+ +H L LK G PVML++N+D GL
Sbjct: 1053 TYLSADSI-DPQDPASLNNPVFTPYFLNSIKLSGLSNHNLTLKIGTPVMLLKNIDPKGGL 1111
Query: 1352 CNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFA 1411
CNGTR+ VT + +++ VI+G V +V I + + P+D +P + +RRQFP+ ++FA
Sbjct: 1112 CNGTRLQVTQMGNHILEARVITGDRVRDKVIIIKAQISPSDTKLPFRMRRRQFPIAVAFA 1171
Query: 1412 MTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIV 1471
M I KSQG++L V +YLP PVFSHGQLYVA+SRV ++ GLK+LI +++ + + T N+V
Sbjct: 1172 MRIKKSQGQSLKEVEIYLPRPVFSHGQLYVALSRVTSKKGLKVLIVDKEGNTQSQTMNVV 1231
Query: 1472 FKEVFQRI 1479
FKE+FQ +
Sbjct: 1232 FKEIFQNL 1239
>At3g31980 hypothetical protein
Length = 1099
Score = 619 bits (1596), Expect = e-177
Identities = 320/725 (44%), Positives = 473/725 (65%), Gaps = 9/725 (1%)
Query: 758 RNKVKKSMFLAWMEANCKYPLGRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPP 817
R + SMF+A+++ N + R TY E P F +D + K+W R++G CIGRMN+
Sbjct: 379 RKANEDSMFMAFLKLNQECEFARQFTYTEIPQYFTWDGQNKQWKLRERGFCIGRMNYASI 438
Query: 818 GSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVAT 877
YY+R+LL + G TS ED+R+ + V++T+++ACLA +L DD+ YID I++ +
Sbjct: 439 KMDPEYYMRILLGIVCGPTSDEDIRTYKDVVYETYKEACLARGILTDDQAYIDTIVEGSL 498
Query: 878 FASGSYIRELFVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQ 937
+ G ++R LF LL +A P +VW + +L + I R+ ++N +LV+ + +
Sbjct: 499 YFFGDHLRNLFSMMLLDKCLARPEYVWEKCSRILIEDIETKKRKQYDNPDLVLTDAERRN 558
Query: 938 LCLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNK 997
L+EIE +L+ NG +L+DF +MP + + N + E N++ ++ + H + NK
Sbjct: 559 YALLEIEDMLLCNGSTLQDFKDMP-KPTKEGTDHSNRFITEEKNYNIEKLREDHDDWFNK 617
Query: 998 LNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGI 1057
+ Q IY+EII AV + G FVYG+GGT KTF+W TL+ +R I +NVASSGI
Sbjct: 618 MTSEQKGIYDEIIKAVLENSGGIFFVYGFGGTSKTFMWKTLSAAVRMRGLISVNVASSGI 677
Query: 1058 ASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEA 1117
ASLLL GGRTAHS F IP+N D+ + C IV S A +LK +SLIIWDEAPM+SRY FE+
Sbjct: 678 ASLLLQGGRTAHSRFGIPINPDDFTTCHIVPNSDLANMLKEASLIIWDEAPMMSRYCFES 737
Query: 1118 LDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRF 1177
LDR+L D++ + KPFGGKVVV GGDFRQ+LPVI RAEIV+A +NSS LW
Sbjct: 738 LDRSLNDVIGNID----GKPFGGKVVVFGGDFRQVLPVIHGAGRAEIVLAALNSSYLWEH 793
Query: 1178 CKVLTLTENMRLFSNS-ETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQ 1236
CKVLTLT+NMRL SN + + E+IK F+ W+L +GDG + + NDG+ I +P++ L++
Sbjct: 794 CKVLTLTKNMRLMSNDLDKDEAEEIKEFSNWLLAVGDGRVSEPNDGEVLIDIPEELLIKD 853
Query: 1237 SSNPVADIVRAIYPEI--MENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYL 1294
+++P+ I +A+Y ++ ++ K+++ +AIL P V+ +N +L G TYL
Sbjct: 854 ANDPIEAITKAVYGDLDLLQPNNDPKFFQQRAILCPRNTDVNTINDIMLDKLNGELVTYL 913
Query: 1295 SSYSVWSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNG 1354
S+ S+ + + +T +FLN I+ SG+P+H L LK G PVML+RN+D GLCNG
Sbjct: 914 SADSI-DPQDAASLNNPVLTPDFLNSIKLSGLPNHNLTLKIGTPVMLLRNIDPKGGLCNG 972
Query: 1355 TRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTI 1414
TR+ VT + +++ VI+G VG +V I + + P+D +P + +RRQFP+ ++FAMTI
Sbjct: 973 TRLQVTQMGNHILEARVITGDRVGDKVIIIKSQISPSDTKLPFRMRRRQFPIAVAFAMTI 1032
Query: 1415 NKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKE 1474
NKSQG++L VG+YLP PVFSHGQLYVA+SRV ++ GLK+LI +++ + + T N+VFKE
Sbjct: 1033 NKSQGQSLKEVGIYLPKPVFSHGQLYVALSRVTSKKGLKVLIVDKEGNTQSQTMNVVFKE 1092
Query: 1475 VFQRI 1479
+FQ I
Sbjct: 1093 IFQNI 1097
Score = 286 bits (731), Expect = 8e-77
Identities = 157/370 (42%), Positives = 213/370 (57%), Gaps = 32/370 (8%)
Query: 230 MVDEFNVLAKSFRRVRDHVQEANSNQVALRLFRHRVN-DPKTYNLPTVDEVAALIVEDFD 288
M+D N + FR +D + N +++ RV D KTY PT EV ALI DF
Sbjct: 1 MLDASNPYVEKFRLAKDKLDSNNGEPFYMQIVSDRVGKDGKTYCNPTTSEVTALIPGDFR 60
Query: 289 TSDCGRDIILRTS-SGNLQRIYDTHSSFLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIY 347
RDII++ +G+L RI + H S++P+QYPLIF YGE+GF I G +
Sbjct: 61 PEMPTRDIIVQDKKTGHLSRISEVHPSYVPMQYPLIFNYGEDGFRPGIQ-KGYTGRTGKQ 119
Query: 348 KRTTISLREWVAFRLQERQFECKRITLSRRLLQQLVVDCYSMIESQRLYYLRNNQETIRR 407
IS+R+W AFR+QER E + + S+RL QQ +VD Y+
Sbjct: 120 ANKCISMRQWYAFRIQERSDEAQTLLRSKRLFQQFLVDGYTT------------------ 161
Query: 408 DFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTM 467
G+T++S G I +P SFTGG RYM N+ D M I + YG+P+LF+T
Sbjct: 162 -----------GGNTNMSEQGKSIRIPQSFTGGPRYMLNSYYDVMAITKMYGFPNLFITF 210
Query: 468 TCNPKWPEIERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEF 527
TCNPKWPEI R+ R L+A DRPD C +F+MKLD LMK+L + G+ +S MYT+EF
Sbjct: 211 TCNPKWPEITRYCKERKLNAEDRPDGVCWIFKMKLDSLMKDLTEEHLLGKTVSAMYTVEF 270
Query: 528 QKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNR 587
QKRGLPHAHILL++ K + + ID++I AE+PD P LY+ V + M+HGPCG
Sbjct: 271 QKRGLPHAHILLFMDKSCKFPTSDDIDNIISAEIPDKSKDPKLYEVVKDCMIHGPCGAAN 330
Query: 588 TNSPCMKNGR 597
SPC+ +G+
Sbjct: 331 KESPCIVDGQ 340
>At3g30420 hypothetical protein
Length = 837
Score = 471 bits (1211), Expect = e-132
Identities = 268/639 (41%), Positives = 388/639 (59%), Gaps = 21/639 (3%)
Query: 848 VHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRET 907
VH++F+ AC A LL D+E+ D + + A +++ +R LFV L+ +++P +W
Sbjct: 199 VHESFKAACHARGLLDGDKEWHDAMDEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHC 258
Query: 908 WSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSNI 967
W +AD ++ +R N +L + + L++ L+EIE LL + +SL D+ MP + +I
Sbjct: 259 WESMADDVLRKQQRVLNFPQLELKAKELEKYTLIEIETLLRQHEKSLSDYPEMPQPEKSI 318
Query: 968 LIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYG 1027
L + N LL E + + + H +KLN Q IY++++ +V + G+ F+YG G
Sbjct: 319 LEEVNNSLLRQEFQINIDKERETHANLFSKLNEQQRIIYDDVLKSVTNKEGKLFFLYGDG 378
Query: 1028 GTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIV 1087
GTGKTFL+ T+ LRS K ++ VASS IA+LLL GGRTAHS F IP+N ED C I
Sbjct: 379 GTGKTFLYKTIISALRSNGKNVMPVASSAIAALLLPGGRTAHSWFKIPINVHEDFICDIK 438
Query: 1088 QGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGG 1147
GS A +L LIIWDEAPM R+ FEA+DRTLRDI+ + + + K GGK V+LGG
Sbjct: 439 IGSMLANVLSKVDLIIWDEAPMAHRHTFEAVDRTLRDILSVGDEKALTKTLGGKTVLLGG 498
Query: 1148 DFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSDVEKIKVFAEW 1207
DFRQILPVIP+ +R E V A IN S LW C L++NMR+ E+IK FAEW
Sbjct: 499 DFRQILPVIPQRTRQETVSAAINRSYLWESCHKYLLSQNMRV-------QPEEIK-FAEW 550
Query: 1208 VLDIGDGVLG------DYNDGDADIKVPKDTLVQQSSNPVADIVRAIYPEIMENVGCAKY 1261
+L IGDG D + + +I + K+ L+ ++ NP+ + +++ P+ +
Sbjct: 551 ILQIGDGEAPRKTHGIDDDQEEDNIIIDKNLLLPETENPLEVLCQSVSPDFTNTFQDLEN 610
Query: 1262 YEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSV---WSVTEDVGIEADWITTEFL 1318
+ A+L P + VD +N Y+LS PG + Y S+ S+ ++TE+ G E + E+L
Sbjct: 611 LKGTAVLTPRNETVDEINDYLLSKVPGLAKEYFSADSIDQDEALTEE-GFEMSY-PMEYL 668
Query: 1319 NDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVG 1378
N + G+P H+L LK G P+ML+RNL+ GLCNGTR+ VTHL V+ ++S T
Sbjct: 669 NSLEFPGLPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLTVTHLGDKVLKAEILSDTTKK 728
Query: 1379 R-QVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHG 1437
R +V I R+ L P D P +RRQFP+ + +AMT+NKSQG+TL+ V LYLP PVFSHG
Sbjct: 729 RKKVLIPRIILSPQDSKHPFTLRRRQFPVRMCYAMTVNKSQGQTLNRVALYLPKPVFSHG 788
Query: 1438 QLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVF 1476
QLYVA+SRV + GL +L ++ + VT NIV++EVF
Sbjct: 789 QLYVALSRVTSPKGLTVLDTSKKKEGKYVT-NIVYREVF 826
>At3g43350 putative protein
Length = 830
Score = 460 bits (1184), Expect = e-129
Identities = 265/616 (43%), Positives = 357/616 (57%), Gaps = 81/616 (13%)
Query: 839 EDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMA 898
+DL++V VH +FR A AL LL DD+EYI+ I D + S Y+R LFV LL ++
Sbjct: 53 KDLKTVKGVVHKSFRDAVFALGLLDDDKEYINAIKDANFWCSAKYVRRLFVIMLLSESLT 112
Query: 899 DPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFD 958
P VW ETW +L+ I H L + +E +Q CL EI +LL NG SL ++
Sbjct: 113 KPEMVWDETWRILSKDIEH----------LQLSDEERQQYCLQEIARLLTKNGVSLSKWN 162
Query: 959 NMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGG 1018
N ++ YG+ KIY+EI+ V D G
Sbjct: 163 RCHKFQMNTMVDYGDF--------------------------RAKKIYDEIMDVVLHDRG 196
Query: 1019 EFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNA 1078
FVYG+GGTGKTFLW L+ +RS+ I LNVASSGIA+L L GGRTAHS F IP+N
Sbjct: 197 GVFFVYGFGGTGKTFLWKLLSAAVRSKGDISLNVASSGIAALRLDGGRTAHSRFDIPINP 256
Query: 1079 DEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPF 1138
+E S C I +GS EL+K + LIIWDEAPM+S++ FE+LDRTL+DI+ NP +KP
Sbjct: 257 NESSTCNISRGSDLGELVKEAKLIIWDEAPMMSKHCFESLDRTLKDIVN--NPG--DKPL 312
Query: 1139 GGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSDV 1198
GGKV+V GGDFRQ+LPVI R EIV A +NSS +W KVL LT+NMRL ++ +
Sbjct: 313 GGKVIVFGGDFRQVLPVINGAGREEIVFAALNSSYIWEHSKVLELTKNMRLLADISEHEK 372
Query: 1199 EKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIYPE-IMENVG 1257
I+ F++W+LD+GDG + NDG A I +P++ L+ ++PV I+ A+Y ME
Sbjct: 373 RDIEDFSKWILDVGDGKISQPNDGIALIDIPEEFLINGDNDPVESIIEAVYGNTFMEEKD 432
Query: 1258 CAK----YYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWI 1313
K Y+ +AIL PT + V+ +N++++ + G ER YLSS S+ A ++
Sbjct: 433 PKKTDYPQYQGRAILCPTNEDVNSINEHMMRMLDGEERIYLSSDSIDPADISSANNAAYL 492
Query: 1314 TTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVIS 1373
+FLN++R G+P+H L LK G PVML+RN+D + GLCNGTR+ VT + ++
Sbjct: 493 -ADFLNNVRVYGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMTDTII------ 545
Query: 1374 GTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPV 1433
Q + +FAMTINKSQG+TL VGLYLP PV
Sbjct: 546 -----------------------------QARFITAFAMTINKSQGQTLESVGLYLPRPV 576
Query: 1434 FSHGQLYVAISRVKTR 1449
FSHGQLYVAISRV ++
Sbjct: 577 FSHGQLYVAISRVTSK 592
Score = 140 bits (353), Expect = 5e-33
Identities = 69/118 (58%), Positives = 86/118 (72%)
Query: 1336 GAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSM 1395
G PVML+RN+D + GLCNGTR+ VT + V+ I+G VG+ V I RM + P D +
Sbjct: 711 GCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTRL 770
Query: 1396 PIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLK 1453
P K +R+QF L ++FAMTINKSQG+TL VGLYLP PVFSHGQLYVAISRV ++ G K
Sbjct: 771 PFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSKTGTK 828
Score = 137 bits (345), Expect = 5e-32
Identities = 67/114 (58%), Positives = 85/114 (73%)
Query: 1336 GAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSM 1395
G PVML+RN+D + GLCNGTR+ VT + V+ I+G VG+ V I RM + P+D +
Sbjct: 595 GCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRL 654
Query: 1396 PIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTR 1449
P K +R+QF L ++FAMTINKSQG+TL VGLYLP PVFSHGQLYVAISRV ++
Sbjct: 655 PFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSK 708
>At3g31440 hypothetical protein
Length = 536
Score = 391 bits (1005), Expect = e-108
Identities = 217/455 (47%), Positives = 293/455 (63%), Gaps = 32/455 (7%)
Query: 1028 GTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIV 1087
GT KT +W TL +R +I+LN+ASSGIASLLL GGRTAHS F I LN DE S C I
Sbjct: 110 GTWKTIIWKTLFAAIRRRDQIVLNMASSGIASLLLEGGRTAHSRFGIRLNPDEFSVCKIK 169
Query: 1088 QGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGG 1147
S A L+K +SL+I D+APM+SR+ FEALD++ DI++ +NK FGGKVVV G
Sbjct: 170 PKSDLANLVKEASLVICDKAPMMSRFCFEALDKSFSDIIK----NTYNKVFGGKVVVFSG 225
Query: 1148 DFRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNS-ETSDVEKIKVFAE 1206
DFRQ+LPVI RAEIVM+++N+S LW CKVL LT+NMRL +N+ ++ ++I F++
Sbjct: 226 DFRQVLPVINGAGRAEIVMSSLNASYLWDHCKVLKLTKNMRLLANNLSETEAKEIHEFSD 285
Query: 1207 WVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY--PEIMENVGCAKYYED 1264
W+L +GDG + + ND A I +PKD L+ + P+ I IY P+I+ + K+++
Sbjct: 286 WLLAVGDGRINEPNDDVAIIDIPKDLLITNADKPIEWITNEIYGDPKILHEITDPKFFQG 345
Query: 1265 KAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEFLNDIRCS 1324
+AILAP + V+ +N+Y+L ER YLS+ S+ T+ + IT +FLN I+
Sbjct: 346 RAILAPKNEDVNTINEYLLEQLHAEERIYLSADSI-DPTDSDSLNNPVITPDFLNSIKLP 404
Query: 1325 GIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIG 1384
G GLCNG R+ +T L +V VI+G +G V I
Sbjct: 405 G------------------------GLCNGARLQITQLFTEIVEAKVITGDRIGHIVLIP 440
Query: 1385 RMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAIS 1444
++L PTD +P K +RRQFPL ++FAMTINKSQG++L HVGLYLP PVFSHGQLYVA+S
Sbjct: 441 TVNLTPTDTKLPFKMRRRQFPLSVAFAMTINKSQGQSLEHVGLYLPKPVFSHGQLYVALS 500
Query: 1445 RVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
RV ++ GLKILI +++ + T NIVFKEVFQ I
Sbjct: 501 RVTSKKGLKILILDKNGKLQKQTTNIVFKEVFQNI 535
Score = 54.7 bits (130), Expect = 4e-07
Identities = 22/55 (40%), Positives = 40/55 (72%)
Query: 823 YYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVAT 877
Y+LR+LLN+ R TS+ ++++ + V+ TF++AC A +L DD+ +IDG+++ T
Sbjct: 5 YFLRVLLNIVRRPTSYAEIKTYDGVVYKTFKEACFARGILDDDQVFIDGLVEATT 59
>At1g64410 unknown protein
Length = 753
Score = 387 bits (994), Expect = e-107
Identities = 207/429 (48%), Positives = 280/429 (65%), Gaps = 5/429 (1%)
Query: 796 KKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQA 855
K+K++ R++G IGR+NFVP + YYLR+LLN++RG TS +DL++V V+ +FR A
Sbjct: 321 KEKKFMIRERGFAIGRINFVPRTIEDAYYLRILLNIKRGVTSSKDLKTVKAVVYKSFRDA 380
Query: 856 CLALSLLKDDREYIDGILDVATFASGSYIRELFVSFLLGNAMADPSHVWRETWSVLADGI 915
AL LL DD+EYI+ I D + S Y+R LFV LL ++ P VW ETW +L++ I
Sbjct: 381 VFALGLLDDDKEYINEIKDANFWCSAKYVRRLFVIMLLSESLTKPEMVWDETWKILSEDI 440
Query: 916 VHALRRTHNNLELVIDNEHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNIL 975
R+ +L + +E +Q CL EI +LL NG SL + MP + S+ ++ N
Sbjct: 441 ERKKRKEWKRPDLQLSDEERQQYCLQEIARLLTKNGVSLSKWKQMPQI-SDEHVEKCNHF 499
Query: 976 LFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLW 1035
+ +E +D +++ H E L + Q KIY+EI+ V D G FVYG+GGTGKTFLW
Sbjct: 500 ILDERKYDRAYLTEKHEEWLTMVTLEQKKIYDEIMDVVLHDRGGVFFVYGFGGTGKTFLW 559
Query: 1036 TTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAEL 1095
L+ +RS+ I LNVASSGIA+LLL GGRTAHS F IP+N +E S C I +G EL
Sbjct: 560 KLLSAAIRSKGDISLNVASSGIAALLLDGGRTAHSRFGIPINPNESSTCNISRGLDFGEL 619
Query: 1096 LKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPV 1155
+K ++LIIWDEA M+S++ FE+LDRTLRDIM NP +KPFGGKV+V GGDFRQ+L V
Sbjct: 620 VKEANLIIWDEAHMMSKHCFESLDRTLRDIMN--NPG--DKPFGGKVIVFGGDFRQVLSV 675
Query: 1156 IPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSDVEKIKVFAEWVLDIGDGV 1215
I R EIV A +NSS +W CKVL LT+NMRL +N + I+ F++W+LD+GDG
Sbjct: 676 INGAGREEIVFAALNSSYIWEHCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGK 735
Query: 1216 LGDYNDGDA 1224
+ NDG A
Sbjct: 736 ISQPNDGIA 744
Score = 148 bits (374), Expect = 2e-35
Identities = 74/160 (46%), Positives = 100/160 (62%), Gaps = 1/160 (0%)
Query: 522 MYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHG 581
MYTIEFQKRGLPHAHILL++ KL + E D VI AE+PD K P LY V + M+HG
Sbjct: 179 MYTIEFQKRGLPHAHILLFMHPTSKLSTAEDTDKVITAEIPDKKKKPELYAVVKDCMIHG 238
Query: 582 PCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFV 641
PCGV NSPCM+NG+C K+FPK++ + T D+DG+PVYRRR+TGI + G D V
Sbjct: 239 PCGVGHPNSPCMENGKCKKYFPKSYSDTTKVDNDGFPVYRRRDTGIYVEKNGHDRVTVTV 298
Query: 642 VPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVDRV 681
P + K + + +Y + S K++ + + R+
Sbjct: 299 EPNDQDTAKKEKDEVK-DYFDCSKEKKFMIRERGFAIGRI 337
Score = 70.5 bits (171), Expect = 7e-12
Identities = 39/84 (46%), Positives = 48/84 (56%), Gaps = 7/84 (8%)
Query: 125 LTNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPA 184
L + D + HF +NIR+YN +F+FTSIGGKVD + G+GP F I G+L P
Sbjct: 65 LQSDDELAKHFRENIRAYNMLFSFTSIGGKVDHCLPKGRGPNMFAIQ-------GALKPK 117
Query: 185 EGDNPKFAQLNIYDTRNEFENRLN 208
KF QL I DT NE NR N
Sbjct: 118 SVAKAKFQQLYIVDTENEVNNRYN 141
>At5g34960 putative protein
Length = 1033
Score = 334 bits (856), Expect = 3e-91
Identities = 213/542 (39%), Positives = 286/542 (52%), Gaps = 104/542 (19%)
Query: 941 MEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFDTVE-MSKLHVECLNKLN 999
+ EK+++ NG +L+D D P + I N L+ EL ++ + + H E L
Sbjct: 592 VSFEKIMLSNGATLEDIDEFP-KPTRDGIDNSNRLIVEELRYNRESNLKEKHEEWKQMLT 650
Query: 1000 GGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIAS 1059
Q +Y EI AV + +I+LNVASSGIAS
Sbjct: 651 PEQRGVYNEITEAV----------------------------FNNLDQIVLNVASSGIAS 682
Query: 1060 LLLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALD 1119
LLL GGRTAHS F I LN DE S C I S A L+K +SL+IWDEAPM+SR
Sbjct: 683 LLLEGGRTAHSRFGISLNPDEFSVCKIKPKSDLANLVKEASLVIWDEAPMMSR------- 735
Query: 1120 RTLRDIMRICNPECFNKPFGGKVVVLGGDFRQILPVIPKGSRAEIVMATINSSRLWRFCK 1179
Q+L VI RAEIVM+++N+S LW CK
Sbjct: 736 -------------------------------QVLLVINGAGRAEIVMSSLNASYLWDHCK 764
Query: 1180 VLTLTENMRLFSNSETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSN 1239
VL + E NDG A I +P+D L+ +
Sbjct: 765 VLKINEP---------------------------------NDGVAIIDIPEDLLITNTDK 791
Query: 1240 PVADIVRAIY--PEIMENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSY 1297
P+ I IY P+I+ + K+++ +AILAPT + V+ +N+Y+L ER YLS+
Sbjct: 792 PIESITNEIYGDPKILHEITDPKFFQGRAILAPTNEDVNTINEYLLEQLHAEERIYLSAD 851
Query: 1298 SVWSVTEDVGIEADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRI 1357
S+ T+ + IT +FLN I+ +G+P+H L LK APVML+RNLD GLCNGTR+
Sbjct: 852 SI-DPTDSNSLNNPVITPDFLNSIKLAGLPNHSLRLKVSAPVMLLRNLDPKGGLCNGTRL 910
Query: 1358 VVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKS 1417
+T L +V VI+G +G V I ++L PTD +P K +RRQFPL ++FAMTIN S
Sbjct: 911 QITQLCTQIVEAKVITGDIIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINTS 970
Query: 1418 QGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQ 1477
QG++L HVGLYLP VFSHGQLYVA+SRV ++ GLK LI ++D + T N+VFKEVFQ
Sbjct: 971 QGQSLEHVGLYLPKAVFSHGQLYVALSRVTSKKGLKFLILDKDGKLQKQTTNVVFKEVFQ 1030
Query: 1478 RI 1479
I
Sbjct: 1031 NI 1032
>At5g32630 putative protein
Length = 856
Score = 321 bits (822), Expect = 2e-87
Identities = 216/660 (32%), Positives = 316/660 (47%), Gaps = 142/660 (21%)
Query: 400 NNQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYG 459
N +T+ R S +++A G T ++ G+++++P+SFTGG RYM + DAM IC YG
Sbjct: 157 NECKTLTRS-KSSLKKASEAGTTSINEEGNKVLIPTSFTGGPRYMVQSYYDAMAICEHYG 215
Query: 460 YPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAI 519
+PDLF+T TCNPKWP I R+ A LS DRPD+ R+F++KLD LMK+L G +
Sbjct: 216 FPDLFITFTCNPKWPGINRYCQAIGLSVDDRPDIVARIFKIKLDSLMKDLTDGNAGKDS- 274
Query: 520 SWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMV 579
M+T+EFQKRGL HA LL++ K KL + + ID +I AE+PD P Y+ + N M+
Sbjct: 275 --MHTVEFQKRGLLHAPTLLFMDAKSKLPTTDDIDKLISAEIPDKDKEPEFYEVIKNSMI 332
Query: 580 HGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNG 639
HGPCG NSPCM +++EK F DN
Sbjct: 333 HGPCGAANMNSPCM--------VEDDYIEKGGF----------------------KCDNS 362
Query: 640 FVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVDRVTMSMSVTQSSNEETTVVD 699
+VVPYN KL ++YQAHIN+E +SN IK + ++ + N D
Sbjct: 363 YVVPYNQKLSLRYQAHINVEC--QSNRIK-------------QKAATLGEPPNSTEKKKD 407
Query: 700 EIKQYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERN 759
EIK + DC L + + + S Y + S+H KQ F+ A I+DVLER
Sbjct: 408 EIKDWFDCSCLEDLQ-ISTSASIYSSS------RLSFHCEWKQPAYFDPNAIIEDVLERI 460
Query: 760 KVKKSMFLAWMEANCKYPLGRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGS 819
+ SMF+AW+ N +G K G +GR+N+
Sbjct: 461 SNEDSMFMAWLTLNRNNDVG------------------------KNGFSLGRINYGARKM 496
Query: 820 GELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFA 879
+ YYL++LLN+ RG S ED+++ N ++ +F++AC A +L DD+ IDG+L+
Sbjct: 497 EDEYYLQVLLNIVRGPMSCEDIKTFNGVLYPSFKEACFARGILDDDQVNIDGLLEA---- 552
Query: 880 SGSYIRELFVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHNNLELVIDNEHLKQLC 939
+L + ++
Sbjct: 553 -----------------------------------------------KLKLTVAEIRNYT 565
Query: 940 LMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFNELNFD-TVEMSKLHVECLNKL 998
L EIEK+++ NG +LKD + S I N L+ +EL ++ + + H E L
Sbjct: 566 LQEIEKIMLFNGATLKDIQDF-MQPSRESIDNSNRLVVDELRYNIDSNLKEKHDEWFQML 624
Query: 999 NGGQAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIA 1058
N IY+EI A+ +D GGT KT +W TL R +I+LNVASSG++
Sbjct: 625 NTEHRGIYDEITGAIFND---------LGGTEKTSMWKTLAAAFRCRGQIVLNVASSGLS 675
Score = 144 bits (364), Expect = 3e-34
Identities = 76/154 (49%), Positives = 104/154 (67%), Gaps = 7/154 (4%)
Query: 1326 IPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGR 1385
IP+ L+ + G P+ +T LCNGTR+ +T + VV VI+G +G + I
Sbjct: 709 IPEELLITEAGNPIE-------ATSLCNGTRLHITQIAKQVVQAKVITGDIIGDIILIPL 761
Query: 1386 MDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISR 1445
++L P+D +P K +RRQFPL +FAMTINKSQG++L GLYLP VFSHGQLYVA+SR
Sbjct: 762 INLTPSDTKLPFKMRRRQFPLSDAFAMTINKSQGQSLEQAGLYLPKLVFSHGQLYVALSR 821
Query: 1446 VKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
V +++GLKILI ++D + T N+VFKE+FQ I
Sbjct: 822 VTSKSGLKILILDKDGDIQKQTTNVVFKELFQNI 855
Score = 98.6 bits (244), Expect = 2e-20
Identities = 55/146 (37%), Positives = 82/146 (55%), Gaps = 7/146 (4%)
Query: 150 SIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEFENRLNH 209
S+GG+VD S+ G+GP F + NYH IGSL P GD K++QL I DT N+ ENR
Sbjct: 3 SLGGRVDNSMPKGKGPNMFRLQEGNYHLIGSLKPKPGDYAKYSQLYIVDTENKVENRATV 62
Query: 210 LSDSTG------KCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSNQVALRLFRH 263
+ G K L +++ L+ M+++ N + F++ R+ +Q N +R+ +
Sbjct: 63 TNKGKGGQNTVAKQKLKKEVIEALIEMLNKVNPYVEKFKQSRERIQADNDELFHMRIVAY 122
Query: 264 RVN-DPKTYNLPTVDEVAALIVEDFD 288
R D +TYN+PT +VAALI FD
Sbjct: 123 RKGVDRRTYNMPTSSKVAALIPGGFD 148
Score = 46.2 bits (108), Expect = 1e-04
Identities = 17/51 (33%), Positives = 34/51 (66%)
Query: 1191 SNSETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPV 1241
S + + I++F +W+L +GDG + + NDG+A I +P++ L+ ++ NP+
Sbjct: 672 SGLSVEEAKDIQLFYDWLLVVGDGRINEPNDGEALIDIPEELLITEAGNPI 722
>At3g42320 putative protein
Length = 541
Score = 301 bits (772), Expect = 1e-81
Identities = 174/438 (39%), Positives = 231/438 (52%), Gaps = 94/438 (21%)
Query: 440 GRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARCLSAYDRPDLACRVFR 499
G RYM NN DAM IC+ +G P +F+T TCNPKW EI R AR L + DRPD+ C++F+
Sbjct: 138 GPRYMANNYLDAMAICQHFGLPSVFITFTCNPKWHEIVRFYKARNLKSEDRPDIICQIFK 197
Query: 500 MKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICA 559
MKLD LM +L G+ +S MYTIEFQKRGL HAHILL+L +KL++ E D VI A
Sbjct: 198 MKLDNLMYDLTTKYLLGKTVSCMYTIEFQKRGLLHAHILLFLHATNKLYTAEDTDKVITA 257
Query: 560 ELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPV 619
E+PD K LY V + M+HGPCGV NSPCM NG+C K+F K++ + T D+DG+PV
Sbjct: 258 EIPDKKKKLELYVLVKDCMIHGPCGVGHPNSPCMVNGQCKKYFSKSYYDTTKVDNDGFPV 317
Query: 620 YRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNCIKYLFKYINKGVD 679
YRR NTGI Y K+ IKYLFKY++K D
Sbjct: 318 YRRHNTGI--------------------------------YAEKNGLIKYLFKYVHKEHD 345
Query: 680 RVTMSMSVTQSSNEETTVVDEIKQYHDCRYLSPCEAVWRTFSFYIHDKWPP---VLKQSY 736
RVT VT N + T E + + K PP V +
Sbjct: 346 RVT----VTIEPNNQDTTKKEKDELN---------------------KKPPSRRVHASAK 380
Query: 737 HLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANCKYPLGRGLTYAEFPSSFVYDKK 796
+LPK P L E P+ F ++ K
Sbjct: 381 NLPKAA----------------------------------PFPTKLLVEEIPNHFTWNSK 406
Query: 797 KKEWHPRKKGICIGRMNFVPPGSGELYYLRMLLNVQRGCTSFEDLRSVNNHVHDTFRQAC 856
+K++ +++G IGR NFVP + YYLR+LLN++RG TS++DL++V +H++FR A
Sbjct: 407 EKKFMIKERGFAIGRTNFVPHTIEDAYYLRILLNIKRGVTSYKDLKTVKGVIHESFRDAV 466
Query: 857 LALSLLKDDREYIDGILD 874
AL LL DD+EY++G D
Sbjct: 467 FALGLLHDDKEYMNGFKD 484
Score = 105 bits (261), Expect = 3e-22
Identities = 54/105 (51%), Positives = 68/105 (64%)
Query: 108 KISLPYLQDAPTLLSNLLTNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQ 167
+I LP L+D+P L +LLT+ D S HF +NIR+YN++F+FTSIG KVD + G P
Sbjct: 6 QIVLPSLKDSPEFLWHLLTSDDELSKHFRENIRAYNTLFSFTSIGSKVDHFLPKGPQPNM 65
Query: 168 FVISGQNYHRIGSLLPAEGDNPKFAQLNIYDTRNEFENRLNHLSD 212
F I G+NYH IG+L P KF QL I DT NE NR N +SD
Sbjct: 66 FAIQGENYHLIGALKPKSSAKAKFQQLYIADTENEVNNRYNIMSD 110
>At4g07800 hypothetical protein
Length = 448
Score = 297 bits (760), Expect = 3e-80
Identities = 193/548 (35%), Positives = 272/548 (49%), Gaps = 121/548 (22%)
Query: 305 LQRIYDTHSSFLPLQYPLIFPYGEEGFSDEIGFDGINYDSSIYKRTTISLREWVAFRLQE 364
++RI H S+L LQYPL+F YGE+G++ I Y+S K+ ++S
Sbjct: 3 VKRISQIHISYLTLQYPLMFCYGEDGYTPAIEKC---YNSGSTKKKSVS----------- 48
Query: 365 RQFECKRITLSRRLLQQLVVDCYSMIESQRLYYLRNNQETIRRDFLSGIEEAIHRGDTDL 424
RL QQ ++D Y++IES RL Y++ NQ +R
Sbjct: 49 ------------RLFQQFLIDAYTIIESNRLAYIKFNQSKLR------------------ 78
Query: 425 SMVGSRIVLPSSFTGGRRYMFNNCQDAMGICREYGYPDLFLTMTCNPKWPEIERHVSARC 484
SSFT RYM DAM IC+ YG+PD FL+
Sbjct: 79 ----------SSFTSASRYMLQTYYDAMAICKHYGFPD-FLS------------------ 109
Query: 485 LSAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTK 544
R++LD LM++L + + +++ +F+KRGLP A ILL++
Sbjct: 110 --------------RLRLDSLMRDLTERNLLRKTVAF----KFKKRGLPRARILLFMEAN 151
Query: 545 DKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPK 604
KL + D + LY+ + N M+HG CG TNS CM +G+CSK +PK
Sbjct: 152 RKLPIAD-----------DIETEQELYELIKNSMIHGLCGSANTNSLCMVDGQCSKLYPK 200
Query: 605 NFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKS 664
E T +DGYPVYRRR + GV DN +VVPYN + +KYQAHIN+E+CN++
Sbjct: 201 KHQELTKVGADGYPVYRRRPIDGCIEKGGVKCDNMYVVPYN-RFSLKYQAHINVEWCNQN 259
Query: 665 NCIKYLFKYINKGVDRVTMSMS-VTQSSNEETTVV-----------DEIKQYHDCRYLSP 712
IKYLFKYINKG DRV + V + +N TT + D IK + D Y+S
Sbjct: 260 GSIKYLFKYINKGPDRVVFIVELVKEETNSNTTTLGDETVTTKKKKDGIKDWFDYIYVSA 319
Query: 713 CEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEA 772
EA+WR F F I + PV K S+H+ KQ F+ +A + DVLER + S F+ W+
Sbjct: 320 SEAIWRIFKFPIQHRSTPVQKLSFHVEGKQPAYFDAKAKMVDVLERVSNEDSQFMVWLTL 379
Query: 773 NCKYPLG------RGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLR 826
N K +G R YAE P+ F +D + K + R +G +GR+N+V + YL
Sbjct: 380 NKKNVVGKNGKRARNCLYAEIPTYFTWDGENKPFKKRTRGFFLGRINYVLRKMEDENYLI 439
Query: 827 MLLNVQRG 834
+LLN+ RG
Sbjct: 440 VLLNIVRG 447
>At3g32320 hypothetical protein
Length = 494
Score = 234 bits (596), Expect = 4e-61
Identities = 122/300 (40%), Positives = 177/300 (58%), Gaps = 22/300 (7%)
Query: 546 KLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKN 605
K + + +D +I AE+PD + P YQ VS ++HGPCG+ NSPCMKNG+CSK++PKN
Sbjct: 169 KFPTADHVDMIIFAEIPDKEKDPEQYQAVSECIIHGPCGLVNPNSPCMKNGKCSKYYPKN 228
Query: 606 FVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSN 665
VE TS D++GYP+YRRR+TG + DN +VVPYN LL KY+AHIN+E+CN+S
Sbjct: 229 HVENTSLDNEGYPIYRRRDTGRFIKKNKYQCDNWYVVPYNDVLLRKYKAHINVEWCNQSV 288
Query: 666 CIKYLFKYINKGVDRVTMSMS------VTQSSNEETTVVD-----EIKQYHDCRYLSPCE 714
+KYLFKY+NKG DRVT+S+ V++ +N T D +++ Y DCR
Sbjct: 289 SVKYLFKYVNKGPDRVTVSVEPHRKEVVSEQNNVGETNNDQQERNQVQDYFDCRIR---- 344
Query: 715 AVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMFLAWMEANC 774
+ IH + + K ++H KQ + E + VL R ++ F AW E N
Sbjct: 345 ------GYPIHYRQTSITKLTFHEKGKQPVYVKEGETAESVLYRVNNDETQFTAWFELNK 398
Query: 775 KYPLGRGLTYAEFPSSFVYDKKKKEWHPRK-KGICIGRMNFVPPGSGELYYLRMLLNVQR 833
+ P L Y + P+ + ++ K K++ RK G +GR+N VPP + Y+LR+L+N R
Sbjct: 399 RDPEAAKLLYEQIPNFYTWNGKDKDFRGRKMPGFVVGRINHVPPKIDDAYHLRILINNMR 458
Score = 67.4 bits (163), Expect = 6e-11
Identities = 43/116 (37%), Positives = 58/116 (49%), Gaps = 7/116 (6%)
Query: 198 DTRNEFENRLNHLSDST------GKCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQ-E 250
DT NE +NR+ +S GK + V L+ M+D+ N +FR RD E
Sbjct: 2 DTDNELDNRIKVMSKGNNGGMEGGKQKYKKETVSALLKMLDKINPHVANFRIARDRFNIE 61
Query: 251 ANSNQVALRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTSDCGRDIILRTSSGNLQ 306
+R+ R D + YNLP+V EVAALI DFD + RDI+L+ SG Q
Sbjct: 62 KEEENFHMRIISRRETDGRVYNLPSVAEVAALIPGDFDDNLDKRDIVLQMKSGKNQ 117
Score = 41.6 bits (96), Expect = 0.003
Identities = 20/52 (38%), Positives = 27/52 (51%)
Query: 401 NQETIRRDFLSGIEEAIHRGDTDLSMVGSRIVLPSSFTGGRRYMFNNCQDAM 452
NQ +R +++ GD DLS G ++P SFTGG YM N DA+
Sbjct: 116 NQTKLRNTNKQAVQDTSDAGDNDLSNKGKSCIIPPSFTGGPAYMQQNYLDAI 167
Score = 39.7 bits (91), Expect = 0.013
Identities = 16/40 (40%), Positives = 26/40 (65%)
Query: 1183 LTENMRLFSNSETSDVEKIKVFAEWVLDIGDGVLGDYNDG 1222
L NMRL + T++ +I+ F +W+L +G+G L + NDG
Sbjct: 453 LINNMRLLQDINTNEAREIEEFFKWILAVGEGKLNEPNDG 492
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,836,343
Number of Sequences: 26719
Number of extensions: 1586157
Number of successful extensions: 3506
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 3237
Number of HSP's gapped (non-prelim): 116
length of query: 1482
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1370
effective length of database: 8,326,068
effective search space: 11406713160
effective search space used: 11406713160
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0060.4


