

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0400a.3
(419 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
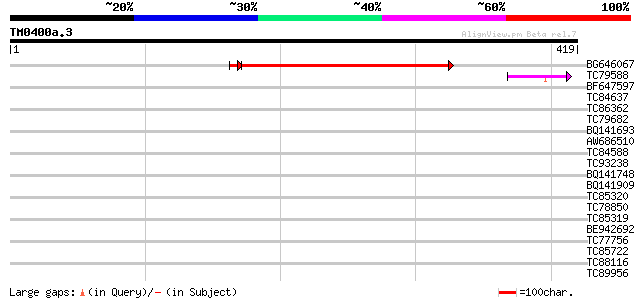
BG646067 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsi... 283 4e-77
TC79588 weakly similar to GP|11994114|dbj|BAB01117. gene_id:MYF2... 53 2e-07
BF647597 weakly similar to GP|19071648|gb| putative protein tyro... 31 1.0
TC84637 weakly similar to SP|O49858|C823_SOYBN Cytochrome P450 8... 30 1.3
TC86362 similar to GP|10177062|dbj|BAB10504. gene_id:MKP11.2~unk... 30 1.7
TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 30 2.3
BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Sch... 29 3.9
AW686510 similar to PIR|A86323|A86 protein F14D16.3 [imported] -... 28 5.0
TC84588 28 6.6
TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage l... 28 6.6
BQ141748 homologue to SP|P21997|SSGP_ Sulfated surface glycoprot... 28 6.6
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 28 6.6
TC85320 similar to GP|1532175|gb|AAB07885.1| similar to protein ... 28 6.6
TC78850 homologue to GP|10177413|dbj|BAB10544. fibrillarin-like ... 28 6.6
TC85319 similar to GP|8778824|gb|AAF79823.1| T6D22.5 {Arabidopsi... 28 6.6
BE942692 similar to GP|8778824|gb| T6D22.5 {Arabidopsis thaliana... 28 6.6
TC77756 similar to PIR|T06796|T06796 glycine-rich RNA-binding pr... 28 6.6
TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 28 8.6
TC88116 weakly similar to PIR|E96803|E96803 probable lipase 416... 28 8.6
TC89956 similar to GP|22136076|gb|AAM91116.1 unknown protein {Ar... 28 8.6
>BG646067 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsis thaliana},
partial (32%)
Length = 516
Score = 283 bits (725), Expect(2) = 4e-77
Identities = 138/157 (87%), Positives = 146/157 (92%)
Frame = +2
Query: 172 KMGRLTAGVQYEPQHGNAKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVKSSQFVASF 231
++GR+TAGVQYEP H NAKLSNLMNWSCAMAYGVGS SPLSPSFNFSLELVKSSQFVASF
Sbjct: 29 RIGRVTAGVQYEPHHENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASF 208
Query: 232 YQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNISDSTFRIGASWQANK 291
YQHMVVQRRVKNPLEEN V+GITNYIDFGFELQTSVDDAIA NNISDSTF+I ASWQANK
Sbjct: 209 YQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAIAANNISDSTFQIAASWQANK 388
Query: 292 NFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDR 328
NFL+KAK GP+ S+MALAFKSWWKPSFTFSIS T R
Sbjct: 389 NFLVKAKAGPKSSTMALAFKSWWKPSFTFSISGTSSR 499
Score = 23.1 bits (48), Expect(2) = 4e-77
Identities = 9/10 (90%), Positives = 9/10 (90%)
Frame = +1
Query: 163 LPKSAWLVSK 172
LPKS WLVSK
Sbjct: 1 LPKSVWLVSK 30
>TC79588 weakly similar to GP|11994114|dbj|BAB01117.
gene_id:MYF24.23~unknown protein {Arabidopsis thaliana},
partial (40%)
Length = 1742
Score = 52.8 bits (125), Expect = 2e-07
Identities = 34/84 (40%), Positives = 37/84 (43%), Gaps = 37/84 (44%)
Frame = +3
Query: 369 VVVSYNILGVDSASKHQDLYSNIRHS---------------------------------- 394
VVVSYNILGV++AS H DLYSNI
Sbjct: 255 VVVSYNILGVENASNHLDLYSNIPRRFLDWGRRKRLILEEIDSYNASILCFQEVDHFDDL 434
Query: 395 ---FQNNGFKGVGKDYTGDARDGC 415
FQNNGFK V K TG+A DGC
Sbjct: 435 DDLFQNNGFKSVYKGRTGEANDGC 506
>BF647597 weakly similar to GP|19071648|gb| putative protein
tyrosine-serine-threonine kinase {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 520
Score = 30.8 bits (68), Expect = 1.0
Identities = 25/101 (24%), Positives = 41/101 (39%), Gaps = 10/101 (9%)
Frame = +3
Query: 223 KSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQT----------SVDDAIA 272
KSS F+ FY H + RVK ++ V+G + G + + D+ +
Sbjct: 15 KSSLFLNYFYVHGLACFRVKVETSKHNVVGREEEVSSGLSYKAASSFVPLTSRTEDEVLR 194
Query: 273 TNNISDSTFRIGASWQANKNFLLKAKVGPRISSMALAFKSW 313
+NN+ + F I A +NF + +G FK W
Sbjct: 195 SNNL--TCFTINEVRAATRNFHPDSMIGK--GGFGCVFKGW 305
>TC84637 weakly similar to SP|O49858|C823_SOYBN Cytochrome P450 82A3 (EC
1.14.-.-) (P450 CP6). [Soybean] {Glycine max}, partial
(43%)
Length = 757
Score = 30.4 bits (67), Expect = 1.3
Identities = 15/51 (29%), Positives = 26/51 (50%)
Frame = +2
Query: 206 GSGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNY 256
GSG + P ++ L++V S +ASF + + P++ GITN+
Sbjct: 479 GSGRRICPGISYGLQMVHLS--LASFLHSFEISKASAEPIDMTEAFGITNH 625
>TC86362 similar to GP|10177062|dbj|BAB10504. gene_id:MKP11.2~unknown
protein {Arabidopsis thaliana}, partial (70%)
Length = 1113
Score = 30.0 bits (66), Expect = 1.7
Identities = 19/72 (26%), Positives = 35/72 (48%)
Frame = -2
Query: 8 PPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFTTRIMFK 67
PPP V+ PP F +P + R L +S +VF A+ + + F + + ++
Sbjct: 491 PPPSTAVITTPPPFFWPKV--RPSGLSTSASIVFEAPAVLAVISSLFLAMEE--EPLLLS 324
Query: 68 PIDEPHVDLIAT 79
P ++P L+A+
Sbjct: 323 PRNDPETPLLAS 288
>TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (17%)
Length = 1031
Score = 29.6 bits (65), Expect = 2.3
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +1
Query: 8 PPPPPLVVLVPPLFDFPPL 26
PPPPP + PL FPPL
Sbjct: 163 PPPPPKSIFPAPLLPFPPL 219
>BQ141693 weakly similar to GP|160968|gb|AA eggshell protein {Schistosoma
japonicum}, partial (12%)
Length = 1226
Score = 28.9 bits (63), Expect = 3.9
Identities = 12/19 (63%), Positives = 12/19 (63%)
Frame = +1
Query: 8 PPPPPLVVLVPPLFDFPPL 26
PPPPPL PP F F PL
Sbjct: 1 PPPPPLSPFFPPFFLFFPL 57
>AW686510 similar to PIR|A86323|A86 protein F14D16.3 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 535
Score = 28.5 bits (62), Expect = 5.0
Identities = 19/63 (30%), Positives = 33/63 (52%)
Frame = -3
Query: 266 SVDDAIATNNISDSTFRIGASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISAT 325
S DDA++ N S S+ A W+ K + +K+G R ++ +S+ SF+ S S+
Sbjct: 281 SKDDAVSANGRS*SSSERRAWWKQTKRSIGASKIGKR---ERMSTESFTMTSFSSSSSSI 111
Query: 326 RDR 328
D+
Sbjct: 110 SDK 102
>TC84588
Length = 492
Score = 28.1 bits (61), Expect = 6.6
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +1
Query: 8 PPPPPLVVLVPPL 20
PPPPP +L+PPL
Sbjct: 214 PPPPPSTILIPPL 252
>TC93238 SP|P03756|VE22_LAMBD EA22 gene protein. {Bacteriophage lambda},
partial (93%)
Length = 887
Score = 28.1 bits (61), Expect = 6.6
Identities = 16/50 (32%), Positives = 24/50 (48%)
Frame = -1
Query: 209 SPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYID 258
S S + +F+ + S AS HM + + NPL N VI + N +D
Sbjct: 557 SAFSQALSFTKPISLSFDANASVRHHMQILTCILNPLTSNPVIAMRNDVD 408
>BQ141748 homologue to SP|P21997|SSGP_ Sulfated surface glycoprotein 185
(SSG 185). {Volvox carteri}, partial (3%)
Length = 1293
Score = 28.1 bits (61), Expect = 6.6
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = +3
Query: 8 PPPPPLVVLVPPLF 21
PPPPPL +L PP F
Sbjct: 21 PPPPPLPLLFPPFF 62
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 28.1 bits (61), Expect = 6.6
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = +3
Query: 7 KPPPPPLVVLVPPLFDFPPLAARNRMLESSYDV 39
K PPP V L+PP+F F P N ++++ ++
Sbjct: 21 KSPPPHPVFLLPPIFFFLPSLFLNGKIKTTEEI 119
>TC85320 similar to GP|1532175|gb|AAB07885.1| similar to protein disulfide
isomerase {Arabidopsis thaliana}, partial (78%)
Length = 746
Score = 28.1 bits (61), Expect = 6.6
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Frame = -1
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNY-IDFGFELQT 265
S SP+S SF+FS+ L SS F Q++ +G +TN+ + F F L
Sbjct: 320 STSPISISFSFSIALPTSSHNEPKFLQYL-----------HHGTQNLTNHAVSFSFILSE 174
Query: 266 SVDD 269
V +
Sbjct: 173 KVSE 162
>TC78850 homologue to GP|10177413|dbj|BAB10544. fibrillarin-like
{Arabidopsis thaliana}, partial (87%)
Length = 1177
Score = 28.1 bits (61), Expect = 6.6
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = -1
Query: 8 PPPPPLVVLVPPLFDFPP 25
PPPPPL L PPL PP
Sbjct: 121 PPPPPLPPLSPPLPPNPP 68
>TC85319 similar to GP|8778824|gb|AAF79823.1| T6D22.5 {Arabidopsis
thaliana}, partial (62%)
Length = 881
Score = 28.1 bits (61), Expect = 6.6
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Frame = -1
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNY-IDFGFELQT 265
S SP+S SF+FS+ L SS F Q++ +G +TN+ + F F L
Sbjct: 371 STSPISISFSFSIALPTSSHNEPKFLQYL-----------HHGTQNLTNHAVSFSFILSE 225
Query: 266 SVDD 269
V +
Sbjct: 224 KVSE 213
>BE942692 similar to GP|8778824|gb| T6D22.5 {Arabidopsis thaliana}, partial
(62%)
Length = 588
Score = 28.1 bits (61), Expect = 6.6
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Frame = -2
Query: 207 SGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNY-IDFGFELQT 265
S SP+S SF+FS+ L SS F Q++ +G +TN+ + F F L
Sbjct: 281 STSPISISFSFSIALPTSSHNEPKFLQYL-----------HHGTQNLTNHAVSFSFILSE 135
Query: 266 SVDD 269
V +
Sbjct: 134 KVSE 123
>TC77756 similar to PIR|T06796|T06796 glycine-rich RNA-binding protein -
garden pea, partial (89%)
Length = 813
Score = 28.1 bits (61), Expect = 6.6
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = -3
Query: 8 PPPPPLVVLVPPLFDFPP 25
PPPPPL PPL+ PP
Sbjct: 511 PPPPPLYPPPPPLYPPPP 458
>TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (61%)
Length = 897
Score = 27.7 bits (60), Expect = 8.6
Identities = 13/42 (30%), Positives = 22/42 (51%)
Frame = -2
Query: 133 SEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKSAWLVSKMG 174
+ +GL G R+G G+ S G++ F + + S WL +G
Sbjct: 608 NSSFGLCGSRFGLGSSSLGLSSYRFGPRLDNSFSLWLSGWLG 483
>TC88116 weakly similar to PIR|E96803|E96803 probable lipase 4162-5963
[imported] - Arabidopsis thaliana, partial (45%)
Length = 917
Score = 27.7 bits (60), Expect = 8.6
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = +1
Query: 394 SFQNNGFKGVGKDYTGDARDGCERQ 418
S Q NG KG+ D+ AR GC+ Q
Sbjct: 529 SEQKNGMKGIRGDWKFSARAGCQSQ 603
>TC89956 similar to GP|22136076|gb|AAM91116.1 unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 742
Score = 27.7 bits (60), Expect = 8.6
Identities = 24/67 (35%), Positives = 27/67 (39%), Gaps = 7/67 (10%)
Frame = +3
Query: 261 FELQTSVDDAIATNNISDSTFRIGASWQANKNF-------LLKAKVGPRISSMALAFKSW 313
FE T V IAT N++ TF GAS KN LL K S +LA W
Sbjct: 180 FERNTIVGGRIATVNVAGETFEAGASILHPKNLHAVNYTKLLNLKAREPSSGFSLAI--W 353
Query: 314 WKPSFTF 320
F F
Sbjct: 354 DGNKFVF 374
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,107,248
Number of Sequences: 36976
Number of extensions: 191468
Number of successful extensions: 1702
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1429
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1663
length of query: 419
length of database: 9,014,727
effective HSP length: 99
effective length of query: 320
effective length of database: 5,354,103
effective search space: 1713312960
effective search space used: 1713312960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0400a.3


