

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0249.1
(68 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
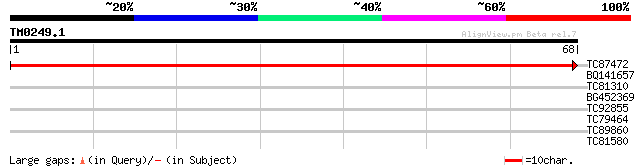
Score E
Sequences producing significant alignments: (bits) Value
TC87472 similar to GP|10177334|dbj|BAB10683. lysosomal Pro-X car... 80 2e-16
BQ141657 25 4.4
TC81310 weakly similar to GP|22136934|gb|AAM91811.1 unknown prot... 25 5.8
BG452369 PIR|T04588|T04 hypothetical protein F23E13.80 - Arabido... 25 7.5
TC92855 25 7.5
TC79464 similar to GP|10177175|dbj|BAB10444. contains similarity... 25 7.5
TC89860 similar to PIR|T05209|T05209 hypothetical protein F24J7.... 25 7.5
TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 24 9.9
>TC87472 similar to GP|10177334|dbj|BAB10683. lysosomal Pro-X carboxypeptidase
{Arabidopsis thaliana}, partial (75%)
Length = 1802
Score = 80.1 bits (196), Expect = 2e-16
Identities = 38/68 (55%), Positives = 50/68 (72%)
Frame = +3
Query: 1 DIEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKW 60
+I L++ SNIIF NGL DPWSGG VL+NIS+++V++V +EGA H DLR ST+ DP W
Sbjct: 1359 NIHLALRKFGSNIIFSNGLLDPWSGGSVLQNISESIVSLVTEEGAPHIDLRASTENDPDW 1538
Query: 61 LKDVRIKE 68
L + R E
Sbjct: 1539 LVEQRATE 1562
>BQ141657
Length = 970
Score = 25.4 bits (54), Expect = 4.4
Identities = 17/67 (25%), Positives = 28/67 (41%), Gaps = 5/67 (7%)
Frame = +3
Query: 6 LKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHR-----DLRYSTKEDPKW 60
+ RS N +Y + W G +KN +T + GA H + RY+ W
Sbjct: 582 ISRSPRNARYYKSNTEAWGGAKTIKNERQT------RSGARHELHLLSNYRYNIH---VW 734
Query: 61 LKDVRIK 67
L +R++
Sbjct: 735 LCTIRVR 755
>TC81310 weakly similar to GP|22136934|gb|AAM91811.1 unknown protein
{Arabidopsis thaliana}, partial (45%)
Length = 984
Score = 25.0 bits (53), Expect = 5.8
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +1
Query: 7 KRSASNIIFYNGLRDPW 23
K + S I+F NG +DPW
Sbjct: 451 KIAGSKIVFTNGSQDPW 501
>BG452369 PIR|T04588|T04 hypothetical protein F23E13.80 - Arabidopsis
thaliana, partial (2%)
Length = 585
Score = 24.6 bits (52), Expect = 7.5
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 9 SASNIIFYNGLRDPW 23
+ S IIF NG +DPW
Sbjct: 263 TGSKIIFTNGSQDPW 307
>TC92855
Length = 625
Score = 24.6 bits (52), Expect = 7.5
Identities = 8/20 (40%), Positives = 14/20 (70%)
Frame = -2
Query: 13 IIFYNGLRDPWSGGGVLKNI 32
I+ + GLRD W GG+++ +
Sbjct: 357 ILIHQGLRDGWCCGGLMRKM 298
>TC79464 similar to GP|10177175|dbj|BAB10444. contains similarity to ATP/GTP
nucleotide-binding protein~gene_id:MCI2.1 {Arabidopsis
thaliana}, partial (69%)
Length = 1739
Score = 24.6 bits (52), Expect = 7.5
Identities = 12/35 (34%), Positives = 21/35 (59%)
Frame = -1
Query: 20 RDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYST 54
+DP+S GVL N+S+ + A + H ++YS+
Sbjct: 1301 QDPYSDLGVLGNMSRVEILFCASQ---HTKMKYSS 1206
>TC89860 similar to PIR|T05209|T05209 hypothetical protein F24J7.50 -
Arabidopsis thaliana, partial (16%)
Length = 1139
Score = 24.6 bits (52), Expect = 7.5
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +2
Query: 3 EKDLKRSASNIIFYNGL 19
EKD K+SAS +FY G+
Sbjct: 476 EKDQKKSASQPLFYKGV 526
>TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (84%)
Length = 827
Score = 24.3 bits (51), Expect = 9.9
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -2
Query: 27 GVLKNISKTLVAIVAKEGAHH 47
G+ K ISK + ++ EGA H
Sbjct: 181 GIKKKISKVVASVFTSEGAKH 119
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,082,069
Number of Sequences: 36976
Number of extensions: 18168
Number of successful extensions: 72
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 72
length of query: 68
length of database: 9,014,727
effective HSP length: 44
effective length of query: 24
effective length of database: 7,387,783
effective search space: 177306792
effective search space used: 177306792
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0249.1


