

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
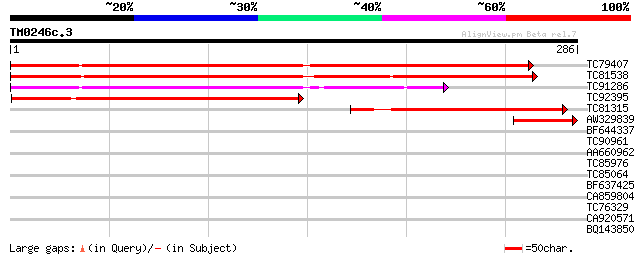
Query= TM0246c.3
(286 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79407 similar to GP|20197174|gb|AAM14957.1 hypothetical protei... 284 2e-77
TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associat... 247 4e-66
TC91286 weakly similar to PIR|E84578|E84578 probable senescence-... 179 1e-45
TC92395 similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_11... 149 2e-36
TC81315 weakly similar to GP|19699320|gb|AAL91270.1 AT3g12090/T2... 119 1e-27
AW329839 similar to PIR|T47500|T475 hypothetical protein F9K21.1... 60 1e-09
BF644337 29 2.4
TC90961 similar to GP|20067056|gb|AAM09517.1 putative glucosyltr... 29 2.4
AA660962 28 4.0
TC85976 similar to PIR|T09845|T09845 L-ascorbate peroxidase (EC ... 28 5.3
TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12 ... 28 5.3
BF637425 27 6.9
CA859804 27 9.0
TC76329 homologue to GP|2780194|emb|CAA05979.1 adenine nucleotid... 27 9.0
CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {A... 27 9.0
BQ143850 27 9.0
>TC79407 similar to GP|20197174|gb|AAM14957.1 hypothetical protein
{Arabidopsis thaliana}, partial (87%)
Length = 1053
Score = 284 bits (727), Expect = 2e-77
Identities = 134/264 (50%), Positives = 185/264 (69%)
Frame = +2
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M+ SN+LIG+LNFLT +LSIPIL GIWL +A + +C +FL+ P+II+GV +++VSL G
Sbjct: 119 MKISNNLIGILNFLTLILSIPILLTGIWLHKQATS-ECERFLEKPIIILGVFLLIVSLMG 295
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F G C R T+L+ YL MFL+I VL F IFA+VVT+KG+G + N+GY +Y L DYS
Sbjct: 296 FIGGCCRVTWLLWFYLFFMFLLIVVLFVFTIFAFVVTNKGAGESLSNKGYKEYRLGDYSN 475
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV + W +I SC++ K+C S + + AD FYL+ L ++QSGCCKP
Sbjct: 476 WLQKRVNDNGNWNRIKSCLQSGKLCIDF---HSQFLNDTADKFYLQHLNALQSGCCKPSN 646
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCG+ YQN T W + +G NPDC W+ND LC+ C SCKAG+L +LK +W+KV+V+
Sbjct: 647 DCGFTYQNPTNWTMPAGGTYTNPDCDTWTNDPKVLCFNCKSCKAGLLDNLKTNWKKVAVV 826
Query: 241 NIVVMIILVIVYIIAYAAYRNNKR 264
NI+ +I L+IVY I A+RNN+R
Sbjct: 827 NIIFLIFLIIVYSIGCCAFRNNRR 898
>TC81538 similar to GP|9758506|dbj|BAB08914.1 senescence-associated protein
5-like protein {Arabidopsis thaliana}, partial (92%)
Length = 1073
Score = 247 bits (630), Expect = 4e-66
Identities = 120/266 (45%), Positives = 177/266 (66%)
Frame = +1
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++IG +N + +LSIPI+G GIWL++ +T C+KFLQWP+II+GV ++ VSLAG
Sbjct: 40 MAMSNNIIGFINLVAVILSIPIIGAGIWLTNEPADT-CVKFLQWPVIILGVLLLFVSLAG 216
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
G+ +R + L+ YLV M ++I +L+ +IF Y+VT +G G NR Y++Y L D+SG
Sbjct: 217 LIGSFWRISCLLIFYLVAMLVLIILLVCLVIFVYMVTLRGHGMIEPNRAYLEYRLDDFSG 396
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
+L+ RV S W I SC+ + +C ++ N A F+ +LT +QSGCCKPPT
Sbjct: 397 FLKRRVRSSFKWDAIRSCLSQTNMCGEL-----NQSFRMAQDFFNARLTPMQSGCCKPPT 561
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
C Y + N T W + +A+ DC +WSNDQ LCY CDSCKAG+LA+L+K W++ +VI
Sbjct: 562 QCAYTFVNPTYW-ISPINNAADMDCLQWSNDQTTLCYNCDSCKAGLLANLRKEWKRANVI 738
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMD 266
I+ +++L++VY+I A+RN K D
Sbjct: 739 LIITVVVLIVVYLIGCFAFRNAKTED 816
>TC91286 weakly similar to PIR|E84578|E84578 probable senescence-associated
protein 5 [imported] - Arabidopsis thaliana, partial
(71%)
Length = 759
Score = 179 bits (453), Expect = 1e-45
Identities = 92/221 (41%), Positives = 132/221 (59%)
Frame = +2
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++ VLN + L SIPI+ GIWL+S+ +N +C+ +WP++IIG+ + +V+L G
Sbjct: 98 MGVSNNITAVLNIIAILASIPIIASGIWLASKPDN-ECIANFRWPIVIIGILVFLVALTG 274
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F GA Y L+ LYL M L+IA+L+ ++FA+VVT V +RGY ++ L +S
Sbjct: 275 FIGAYYNKEGLLALYLFAMALLIALLLIILVFAFVVTRPDGSYVVPDRGYKEFRLDGFSS 454
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL RV W KI C+ S VC K+ N I AD F+ ++ +QSGCCKPPT
Sbjct: 455 WLRHRVTGSGSWRKIMPCLAASDVCIKL---TQNYI--TADQFFNSHISPLQSGCCKPPT 619
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDS 221
CGY Y + +W M A+ C+ W+NDQ LCY C++
Sbjct: 620 VCGYSYVSPIMWTNPVNPM-ADSRCNLWNNDQKQLCYNCNA 739
>TC92395 similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_110
{Arabidopsis thaliana}, partial (54%)
Length = 575
Score = 149 bits (375), Expect = 2e-36
Identities = 70/147 (47%), Positives = 97/147 (65%)
Frame = +1
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IG+LN LT L SIPI+G G+W++ ++T C FLQ PL++IG ++V+SLAGF
Sbjct: 118 RFSNTVIGLLNLLTLLASIPIIGAGLWMAR--SSTTCANFLQTPLLVIGFIVLVISLAGF 291
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC+ + LYLV+M L+I L+G IF + VT KG G V R Y +Y+L DYS W
Sbjct: 292 IGACFHVACALWLYLVIMLLLIVALLGLTIFGFGVTSKGGGVEVPGRSYSEYHLTDYSPW 471
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKM 148
L++R+ YW I +C+ SK C K+
Sbjct: 472 LKKRIQDPRYWNTIKNCILGSKTCDKL 552
>TC81315 weakly similar to GP|19699320|gb|AAL91270.1 AT3g12090/T21B14_110
{Arabidopsis thaliana}, partial (25%)
Length = 629
Score = 119 bits (298), Expect = 1e-27
Identities = 51/109 (46%), Positives = 73/109 (66%)
Frame = +3
Query: 173 SGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKK 232
+GCCKPPT C Y N+ + +M+ + DC KWSN+ LCY CDSCKAGVL +++
Sbjct: 30 TGCCKPPTACNY--------NMEAVMMTQDSDCYKWSNEPTLLCYECDSCKAGVLEDIRR 185
Query: 233 SWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHP 281
+W K+SV+ + ++I+L+ +Y I A+RN +R + D PYGE RMTK P
Sbjct: 186 NWHKLSVLTVTMLILLIGIYSIGCCAFRNARRAETDYPYGENRMTKVRP 332
>AW329839 similar to PIR|T47500|T475 hypothetical protein F9K21.180 -
Arabidopsis thaliana, partial (11%)
Length = 229
Score = 60.1 bits (144), Expect = 1e-09
Identities = 27/32 (84%), Positives = 28/32 (87%)
Frame = +2
Query: 255 AYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 286
AY AYRNNK+MDNDEPYGEARMTKS PS F L
Sbjct: 2 AYYAYRNNKKMDNDEPYGEARMTKSQPSAFHL 97
>BF644337
Length = 567
Score = 28.9 bits (63), Expect = 2.4
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 1/82 (1%)
Frame = -3
Query: 53 IMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGY-M 111
+ +VS GF G C R +F+ + V+ GF+ +V+ S RV + + +
Sbjct: 544 VKIVSFCGF*GLCLR-----------VFVGVKVV*GFL---WVLRLFESSERVKRKSFCL 407
Query: 112 DYYLQDYSGWLEERVASHSYWG 133
LQ+ GW E V WG
Sbjct: 406 HEGLQEEQGWFFEGVYREMCWG 341
>TC90961 similar to GP|20067056|gb|AAM09517.1 putative glucosyltransferase
{Phaseolus lunatus}, partial (9%)
Length = 1130
Score = 28.9 bits (63), Expect = 2.4
Identities = 22/78 (28%), Positives = 39/78 (49%), Gaps = 2/78 (2%)
Frame = +1
Query: 123 EERVASHSYWGKIASCVRDSKVCAKMGRV-DSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
E R AS + + + SC+ +K+ + D ++V+Y+R ++ + CCK +
Sbjct: 256 EYRKASSN*YNQTLSCIN*P*SSSKLFNIKDHTTRGSTSNVYYIRCVSWLGYQCCKESRN 435
Query: 182 CGY-IYQNETVWNLGSGL 198
Y I+ +WNLGS L
Sbjct: 436 *EYIIHYLWCLWNLGSYL 489
>AA660962
Length = 685
Score = 28.1 bits (61), Expect = 4.0
Identities = 18/47 (38%), Positives = 24/47 (50%)
Frame = -3
Query: 25 GGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGACYRNTFL 71
GG+ S + +NTD F Q PL I VS +S A +G Y F+
Sbjct: 389 GGLKQSLQCHNTDLDSFHQQPLSTIQVSSPRLSPASHSGRPYHFVFV 249
>TC85976 similar to PIR|T09845|T09845 L-ascorbate peroxidase (EC 1.11.1.11)
glyoxysomal - upland cotton, partial (95%)
Length = 1296
Score = 27.7 bits (60), Expect = 5.3
Identities = 14/32 (43%), Positives = 18/32 (55%), Gaps = 2/32 (6%)
Frame = +2
Query: 140 RDSKVCAKMGRVD--SNGIPEPADVFYLRKLT 169
RDSKVC + GR+ G+ D+FY LT
Sbjct: 524 RDSKVCTRDGRLPDAKQGVSHLRDIFYRMGLT 619
>TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12
{Arabidopsis thaliana}, partial (29%)
Length = 645
Score = 27.7 bits (60), Expect = 5.3
Identities = 15/56 (26%), Positives = 31/56 (54%)
Frame = +2
Query: 47 IIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSG 102
I++ S++++ L F+ RN ++++YL + L+ L+GF +Y + G G
Sbjct: 407 IMLSSSLLMLGL--FSPIR*RNLIMIKIYLFSLGLIYRFLVGFSKISYRFVEPGLG 568
>BF637425
Length = 321
Score = 27.3 bits (59), Expect = 6.9
Identities = 16/60 (26%), Positives = 32/60 (52%)
Frame = +2
Query: 69 TFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGWLEERVAS 128
T+L L++VV +++ V+ F+++ + S R+ + Y+ YY + WL E + S
Sbjct: 11 TWLANLFMVVTPMLVVVV--FLLYLVLFLPSSSTRKSMR--YISYYPFLFLAWLTE*IGS 178
>CA859804
Length = 525
Score = 26.9 bits (58), Expect = 9.0
Identities = 12/47 (25%), Positives = 26/47 (54%)
Frame = -1
Query: 71 LMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQD 117
++ LY +++ L+I V F ++ + SGR + R + +Y+L +
Sbjct: 504 IILLYYLIIILLIMVWEIFAFLLLIIIEMCSGRTLRTR*FTNYFLME 364
>TC76329 homologue to GP|2780194|emb|CAA05979.1 adenine nucleotide
translocator {Lupinus albus}, partial (81%)
Length = 1497
Score = 26.9 bits (58), Expect = 9.0
Identities = 15/51 (29%), Positives = 21/51 (40%)
Frame = +3
Query: 172 QSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSC 222
+S CC P D + E +W L +S + D S + F C SC
Sbjct: 123 RSECCTP--DRSFYVPKEVIWELLQPCISVSNDASM*CCNNSFPYLCCSSC 269
>CA920571 similar to GP|12322018|gb unknown protein; 3519-5443 {Arabidopsis
thaliana}, partial (48%)
Length = 827
Score = 26.9 bits (58), Expect = 9.0
Identities = 12/45 (26%), Positives = 25/45 (54%), Gaps = 1/45 (2%)
Frame = +1
Query: 239 VINIVVMIILVIVYIIAYAAYRNNKRMDNDEP-YGEARMTKSHPS 282
++ +I L+ + I++ Y+ +++ + P YGE R +HPS
Sbjct: 208 ILQASAIITLITMKILSMVIYQGHQQWPFETPRYGEGRTRHAHPS 342
>BQ143850
Length = 848
Score = 26.9 bits (58), Expect = 9.0
Identities = 13/37 (35%), Positives = 23/37 (62%), Gaps = 1/37 (2%)
Frame = -2
Query: 126 VASHSYWGKIA-SCVRDSKVCAKMGRVDSNGIPEPAD 161
VASH +A SC++ + C+ +G+ S+G+P +D
Sbjct: 244 VASH*SLSPLA*SCIKGEESCSPLGKEQSHGMPCVSD 134
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,281,483
Number of Sequences: 36976
Number of extensions: 187418
Number of successful extensions: 1105
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1088
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1094
length of query: 286
length of database: 9,014,727
effective HSP length: 95
effective length of query: 191
effective length of database: 5,502,007
effective search space: 1050883337
effective search space used: 1050883337
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0246c.3


