

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.8
(334 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
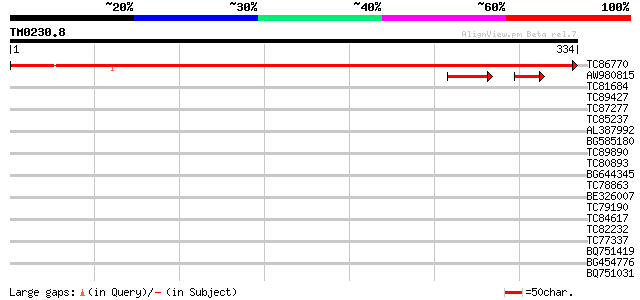
Score E
Sequences producing significant alignments: (bits) Value
TC86770 similar to GP|13872875|dbj|BAB43982. contains EST AU0965... 582 e-167
AW980815 homologue to GP|13872875|dbj contains EST AU096506(S140... 54 2e-11
TC81684 weakly similar to GP|21593080|gb|AAM65029.1 unknown {Ara... 31 0.76
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 30 1.00
TC87277 similar to PIR|T45588|T45588 arm repeat containing prote... 30 1.7
TC85237 similar to GP|4200165|emb|CAA76145.1 neutral invertase {... 29 2.2
AL387992 weakly similar to GP|507345|gb|AA TonB {Haemophilus inf... 29 2.2
BG585180 29 2.9
TC89890 weakly similar to GP|17065226|gb|AAL32767.1 Unknown prot... 28 3.8
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 28 4.9
BG644345 weakly similar to GP|19697333|gb putative protein poten... 28 4.9
TC78863 weakly similar to GP|18481710|gb|AAL73532.1 hypothetical... 28 4.9
BE326007 similar to GP|13676415|d hypothetical protein {Glycine ... 28 6.5
TC79190 similar to PIR|E96542|E96542 scarecrow-like protein [imp... 28 6.5
TC84617 similar to GP|1151236|gb|AAB68209.1| Lpg18p {Saccharomyc... 28 6.5
TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protei... 27 8.4
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 27 8.4
BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania maj... 27 8.4
BG454776 weakly similar to PIR|T49896|T4 glycine/proline-rich pr... 27 8.4
BQ751031 27 8.4
>TC86770 similar to GP|13872875|dbj|BAB43982. contains EST
AU096506(S14064)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (73%)
Length = 2061
Score = 582 bits (1499), Expect = e-167
Identities = 293/336 (87%), Positives = 311/336 (92%), Gaps = 2/336 (0%)
Frame = +2
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK- 59
DD EG RN+DDDNFIDDTGVEP YG Y+EP SPG+APQAEEGEEDDEI DLFKMGKKK
Sbjct: 590 DDNEGARNMDDDNFIDDTGVEPALYG-YDEPRSPGDAPQAEEGEEDDEIKDLFKMGKKKK 766
Query: 60 -NERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEF 118
NERSPAEIALLVENV+AELEVTAEEDAELNRQ KPA+NKLKKLPLL EVLSKKQLQLEF
Sbjct: 767 KNERSPAEIALLVENVMAELEVTAEEDAELNRQHKPAVNKLKKLPLLIEVLSKKQLQLEF 946
Query: 119 LDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIM 178
LDHGVL LLK+WLEPLPDGSLPNINIRTAILKILND PIDLE DRREQLKRSGLGKVIM
Sbjct: 947 LDHGVLNLLKSWLEPLPDGSLPNINIRTAILKILNDLPIDLEHYDRREQLKRSGLGKVIM 1126
Query: 179 FLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAP 238
FLS+SDEEINVNR+L K+LVDKWSRPIFNKSTRFEDMRN ED+R P+RRPSVKKPA KA
Sbjct: 1127FLSRSDEEINVNRRLAKDLVDKWSRPIFNKSTRFEDMRNTEDDRVPYRRPSVKKPAAKAA 1306
Query: 239 GMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHD 298
GMQSRD DLDLDL QPRSG+SSSRQHASRPEATP+DFVIRPQSK+DP+E+RARAKQA+ D
Sbjct: 1307GMQSRDGDLDLDLSQPRSGESSSRQHASRPEATPLDFVIRPQSKIDPDEIRARAKQATQD 1486
Query: 299 QQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
Q RMKMNKKLQQLRAPKK+QLQATKLSVEGRGM KY
Sbjct: 1487QHRMKMNKKLQQLRAPKKKQLQATKLSVEGRGMAKY 1594
>AW980815 homologue to GP|13872875|dbj contains EST AU096506(S14064)~unknown
protein {Oryza sativa (japonica cultivar-group)},
partial (9%)
Length = 175
Score = 53.9 bits (128), Expect(2) = 2e-11
Identities = 24/26 (92%), Positives = 26/26 (99%)
Frame = +2
Query: 259 SSSRQHASRPEATPMDFVIRPQSKVD 284
SSSRQHASRPEATP+DFVIRPQSK+D
Sbjct: 2 SSSRQHASRPEATPLDFVIRPQSKID 79
Score = 32.3 bits (72), Expect(2) = 2e-11
Identities = 14/18 (77%), Positives = 16/18 (88%)
Frame = +3
Query: 298 DQQRMKMNKKLQQLRAPK 315
DQ +KMNKKLQQLRAP+
Sbjct: 120 DQSCLKMNKKLQQLRAPR 173
>TC81684 weakly similar to GP|21593080|gb|AAM65029.1 unknown {Arabidopsis
thaliana}, partial (29%)
Length = 732
Score = 30.8 bits (68), Expect = 0.76
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 7/59 (11%)
Frame = +3
Query: 3 QEGVRNLDDDNFIDDTGV------EPGF-YGNYNEPSSPGEAPQAEEGEEDDEINDLFK 54
+E + LD+D +DD G+ +PG + + +EP P + P+ GE+ DE + F+
Sbjct: 273 KEPIPVLDEDPNVDDCGIRLFRHSKPGIVFDHADEPQPPMKRPKLVPGEDIDEKSKKFR 449
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 30.4 bits (67), Expect = 1.00
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +1
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEIND 51
DD + V + DDDN D +G E + + P A +++ +EDD+ +D
Sbjct: 328 DDDDDVNDEDDDNDEDFSGDEDDEDADPEDDPVPNGAGGSDDDDEDDDDDD 480
>TC87277 similar to PIR|T45588|T45588 arm repeat containing protein homolog
- Arabidopsis thaliana, partial (48%)
Length = 1393
Score = 29.6 bits (65), Expect = 1.7
Identities = 33/142 (23%), Positives = 58/142 (40%), Gaps = 1/142 (0%)
Frame = +1
Query: 12 DNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPA-EIALL 70
+ + + G+EP + +EPS A E + + + G +++RS A EI LL
Sbjct: 49 EQWCEANGIEPPKRPSTSEPSKSASACTPAERSKIESLIQKLTSGGPEDQRSAAGEIRLL 228
Query: 71 VENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNW 130
A+ N + AI + +PLL +LS + + +H V LL
Sbjct: 229 ---------------AKRNADNRVAIAEAGAIPLLVGLLSVPDSRTQ--EHAVTALLNLS 357
Query: 131 LEPLPDGSLPNINIRTAILKIL 152
+ GS+ + I+ +L
Sbjct: 358 IYESNKGSIVSSGAVPGIVHVL 423
>TC85237 similar to GP|4200165|emb|CAA76145.1 neutral invertase {Daucus
carota}, partial (68%)
Length = 2255
Score = 29.3 bits (64), Expect = 2.2
Identities = 14/37 (37%), Positives = 22/37 (58%), Gaps = 5/37 (13%)
Frame = -3
Query: 130 WLEPLP-----DGSLPNINIRTAILKILNDFPIDLEQ 161
W +PLP +G LP + ++ KIL++FP +EQ
Sbjct: 687 WHQPLPWAVAINGLLPTLQLQRVK*KILDNFPFTIEQ 577
>AL387992 weakly similar to GP|507345|gb|AA TonB {Haemophilus influenzae},
partial (8%)
Length = 446
Score = 29.3 bits (64), Expect = 2.2
Identities = 23/58 (39%), Positives = 32/58 (54%), Gaps = 4/58 (6%)
Frame = +1
Query: 81 TAEEDAELNRQGKPAINKLKKLPLLT--EVL--SKKQLQLEFLDHGVLTLLKNWLEPL 134
T E +L + K +NK + PLL E+L KKQ++ E L+ G L LLK + PL
Sbjct: 151 TNPESQQLPQLKKHLLNKRRLQPLLRRPELLLHPKKQVKREQLEEGRLPLLKFIINPL 324
>BG585180
Length = 589
Score = 28.9 bits (63), Expect = 2.9
Identities = 17/57 (29%), Positives = 26/57 (44%)
Frame = +1
Query: 15 IDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIALLV 71
I DT +P + YN EEGEE+ + +GK+ SP +IA ++
Sbjct: 82 ISDTLPKPDLFYKYNNNIIINNGLMLEEGEEEMSRDQSEALGKQLCHASPNDIAKMI 252
>TC89890 weakly similar to GP|17065226|gb|AAL32767.1 Unknown protein
{Arabidopsis thaliana}, partial (76%)
Length = 1013
Score = 28.5 bits (62), Expect = 3.8
Identities = 22/58 (37%), Positives = 27/58 (45%)
Frame = -1
Query: 96 INKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILN 153
I K+ KLPLL L+ L ++ NW PLPD I I+ AILK N
Sbjct: 767 IFKINKLPLLP---------LDILKSSSK*VISNWPRPLPDDF---IGIKLAILKKSN 630
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 28.1 bits (61), Expect = 4.9
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 6/57 (10%)
Frame = +3
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQA------EEGEEDDEIND 51
DD + V++ DDD +D + G E P + P+A ++GE+DD+ D
Sbjct: 342 DDDDDVQDEDDDGEEEDYSGDEG-----EEEGDPEDDPEANGAGGSDDGEDDDDDGD 497
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 28.1 bits (61), Expect = 4.9
Identities = 25/97 (25%), Positives = 44/97 (44%), Gaps = 4/97 (4%)
Frame = +1
Query: 228 PSVKKPAN-KAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDP- 285
P ++P+ + PG Q Q Q+ +Q AS P TP P S++ P
Sbjct: 136 PQAQQPSLFQTPGQQQASPFQTPGQQQSSPFQTPGQQQAS-PFQTPGQQQPSPFSQITPF 312
Query: 286 --EEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQ 320
++ + + +Q QQ+ ++ QQL+ ++ QLQ
Sbjct: 313 SQQQQQLQFQQQQQQQQQQLQQQQQQQLQQQQQLQLQ 423
>TC78863 weakly similar to GP|18481710|gb|AAL73532.1 hypothetical protein
{Sorghum bicolor}, partial (13%)
Length = 1830
Score = 28.1 bits (61), Expect = 4.9
Identities = 13/45 (28%), Positives = 24/45 (52%)
Frame = +1
Query: 6 VRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEIN 50
+ +DD +D GV +Y ++ +SP P++E E+D + N
Sbjct: 1006 ITGMDDPRSMDKVGVAQQWYDELDDLASPKADPESEVIEDDLDQN 1140
>BE326007 similar to GP|13676415|d hypothetical protein {Glycine max},
partial (8%)
Length = 544
Score = 27.7 bits (60), Expect = 6.5
Identities = 23/96 (23%), Positives = 43/96 (43%), Gaps = 7/96 (7%)
Frame = +1
Query: 10 DDDNF-IDDTGVEPGFYGNYNE------PSSPGEAPQAEEGEEDDEINDLFKMGKKKNER 62
DDD F I++ G + ++ E PSS + E+D E + + +
Sbjct: 58 DDDIFEIENGKARGGGFDSFKEGDADCSPSSSWKRVDNNTDEDDSEDSGSDRAESSSPDA 237
Query: 63 SPAEIALLVENVVAELEVTAEEDAELNRQGKPAINK 98
S A+I +++ + L+V A + A L+ G A ++
Sbjct: 238 SMADIMPMLDELHPLLDVDAPQPAHLSHDGSDAASE 345
>TC79190 similar to PIR|E96542|E96542 scarecrow-like protein [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1669
Score = 27.7 bits (60), Expect = 6.5
Identities = 20/75 (26%), Positives = 32/75 (42%), Gaps = 5/75 (6%)
Frame = +2
Query: 258 QSSSRQHASRPEATPMDFVIRPQSKVD-----PEEVRARAKQASHDQQRMKMNKKLQQLR 312
Q+S H+ P +P +RPQ ++ PEE + HD R KM++ +R
Sbjct: 122 QNSPSTHSFSPNNSPGS-TLRPQHSLEFVNGSPEEEDSYLIYHDHDDLRHKMSELESVMR 298
Query: 313 APKKRQLQATKLSVE 327
P L+ V+
Sbjct: 299 GPNVEMLEMYDTKVQ 343
>TC84617 similar to GP|1151236|gb|AAB68209.1| Lpg18p {Saccharomyces
cerevisiae}, partial (86%)
Length = 769
Score = 27.7 bits (60), Expect = 6.5
Identities = 25/98 (25%), Positives = 39/98 (39%)
Frame = +1
Query: 203 RPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSR 262
R FN S +D+R R + KKP KAP +Q + L L + R R
Sbjct: 361 RKFFNLSKE-DDVRKFVIRREITPKNQNKKPYTKAPKIQRLVTPLTLQRKRHRISLKRRR 537
Query: 263 QHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQ 300
A++ D ++ ++K E K+ S Q+
Sbjct: 538 AKAAKAAKEEYDVLLAKRNKEQKERKADLKKRRSAVQK 651
>TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 1181
Score = 27.3 bits (59), Expect = 8.4
Identities = 18/73 (24%), Positives = 34/73 (45%)
Frame = +3
Query: 244 DSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMK 303
DSD DL L +++S + P+D ++ +E+ + Q D++ +
Sbjct: 51 DSD-DLGLQSKTGLFGDGNEYSSDCDVKPVDITDVEPVEIHEKELEHSSAQQKLDRELKE 227
Query: 304 MNKKLQQLRAPKK 316
++KKL+Q A K
Sbjct: 228 LDKKLEQKEAEMK 266
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 27.3 bits (59), Expect = 8.4
Identities = 18/49 (36%), Positives = 25/49 (50%), Gaps = 1/49 (2%)
Frame = +3
Query: 1 DDQEGV-RNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDE 48
DD++G ++ DDD DD E G G +E E EE E++DE
Sbjct: 603 DDEDGDDQDEDDDEDEDDDDEEEG--GEEDEEEGVDEEDNEEEEEDEDE 743
>BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania major}, partial
(1%)
Length = 766
Score = 27.3 bits (59), Expect = 8.4
Identities = 28/95 (29%), Positives = 40/95 (41%), Gaps = 5/95 (5%)
Frame = +2
Query: 223 APFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHAS-RPEATPMDFVIRP-- 279
AP RR +PA + LDL L QPR R HA P T ++ R
Sbjct: 215 APRRRLPHPRPAPR----------LDLHLHQPRHRVRRQRHHAQHHPHPTVLERHRRRHP 364
Query: 280 -QSKVDPEEVRARAKQASH-DQQRMKMNKKLQQLR 312
+ + P + RAR + H + R ++ + QLR
Sbjct: 365 LRREAQPHDPRARGPEGQHPGRHRGRLRRGPVQLR 469
>BG454776 weakly similar to PIR|T49896|T4 glycine/proline-rich protein -
Arabidopsis thaliana, partial (7%)
Length = 680
Score = 27.3 bits (59), Expect = 8.4
Identities = 17/82 (20%), Positives = 29/82 (34%)
Frame = +1
Query: 191 RKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDLD 250
R +T+ RP +TR + P RP+ ++ P +R +
Sbjct: 361 RPMTRRTTRPTPRPTTRPTTRHTTRLTTKPTPRPTTRPTTRRTTRPTPRPTTRPTTRRTT 540
Query: 251 LPQPRSGQSSSRQHASRPEATP 272
P PR + + +RP P
Sbjct: 541 RPTPRPTTRPTTRLMTRPTTRP 606
>BQ751031
Length = 278
Score = 27.3 bits (59), Expect = 8.4
Identities = 18/50 (36%), Positives = 20/50 (40%), Gaps = 3/50 (6%)
Frame = +2
Query: 227 RPSVKKPANKAPGMQSRD---SDLDLDLPQPRSGQSSSRQHASRPEATPM 273
RP P P M S S D P+P SS RQ ASR P+
Sbjct: 11 RPCSAPPQRPVPSMASTTAPRSPTDALKPRPTLSPSSRRQRASRAPMAPL 160
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.133 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,568,061
Number of Sequences: 36976
Number of extensions: 89330
Number of successful extensions: 467
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 454
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 462
length of query: 334
length of database: 9,014,727
effective HSP length: 97
effective length of query: 237
effective length of database: 5,428,055
effective search space: 1286449035
effective search space used: 1286449035
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0230.8


