

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.4
(606 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
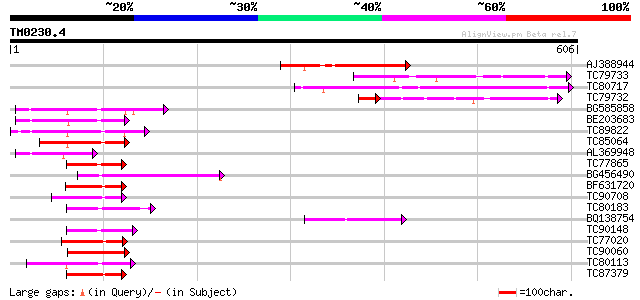
Sequences producing significant alignments: (bits) Value
AJ388944 weakly similar to PIR|H84769|H847 probable DnaJ protein... 130 1e-30
TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM02... 108 8e-24
TC80717 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical... 106 3e-23
TC79732 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical... 86 2e-21
BG585858 similar to GP|15983799|gb At2g05250/F5G3.15 {Arabidopsi... 96 4e-20
BE203683 weakly similar to GP|14587221|d contains ESTs AU108104(... 90 3e-18
TC89822 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12 ... 78 9e-15
TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12 ... 70 2e-12
AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321... 64 1e-10
TC77865 similar to GP|17473565|gb|AAL38258.1 dnaJ-like protein {... 64 2e-10
BG456490 weakly similar to GP|22296361|dbj AHM1(AT hook-containi... 64 2e-10
BF631720 weakly similar to GP|10798648|em putative DNAJ protein ... 61 1e-09
TC90708 weakly similar to PIR|S26703|S26703 dnaJ protein homolog... 57 2e-08
TC80183 similar to GP|4508077|gb|AAD21421.1| 62114 {Arabidopsis ... 57 3e-08
BQ138754 weakly similar to GP|14587221|db contains ESTs AU108104... 57 3e-08
TC90148 similar to GP|6403503|gb|AAF07843.1| putative DnaJ prote... 55 6e-08
TC77020 similar to PIR|D96795|D96795 probable DnaJ protein 1979... 54 2e-07
TC90060 similar to GP|23428796|gb|AAM23263.1 DnaJ-like protein {... 53 3e-07
TC80113 weakly similar to GP|21554000|gb|AAM63081.1 unknown {Ara... 53 3e-07
TC87379 similar to PIR|T04618|T04618 heat shock protein homolog ... 52 9e-07
>AJ388944 weakly similar to PIR|H84769|H847 probable DnaJ protein [imported]
- Arabidopsis thaliana, partial (9%)
Length = 448
Score = 130 bits (327), Expect = 1e-30
Identities = 76/144 (52%), Positives = 90/144 (61%), Gaps = 5/144 (3%)
Frame = +2
Query: 290 RSSRKRVENKRGLEIGEVRTLKLP----IKENAVKSKRGSEVGEERSLKKNVKLAIKEKP 345
R R+E R GE+ + + +K ++S+R + EE +A+ +
Sbjct: 26 RRKASRLERHRDTSGGELEAMVVADLDSVKRKGLRSERHIDTSEEEL----EVMAVADLD 193
Query: 346 EASGKRKRLELEECGDANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYD-DDDGM 404
S KRK L E+ D +G ELEVMAV DSDFYDFDKDRVERSFKKGQVWA YD DDDGM
Sbjct: 194 --SVKRKGLRSEKHMDTSGEELEVMAVADSDFYDFDKDRVERSFKKGQVWAVYDGDDDGM 367
Query: 405 PRHYALIDETVSANPFEVRISWLD 428
PR Y LIDETVSANPF ISWLD
Sbjct: 368 PRQYVLIDETVSANPFNXMISWLD 439
>TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM021B04.9 -
Arabidopsis thaliana, partial (2%)
Length = 1253
Score = 108 bits (269), Expect = 8e-24
Identities = 79/247 (31%), Positives = 116/247 (45%), Gaps = 14/247 (5%)
Frame = +2
Query: 368 EVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYA------LIDETVSANPFE 421
E V D F FD +R F+ GQ+WA Y D+D +P++Y ID +
Sbjct: 347 ESFEVPDPSFNQFDAERSHEKFEAGQIWAFYGDEDELPKYYGQIKCVRRIDSKIELQVIY 526
Query: 422 VRISWLDLQSNADGKIVSREKMGFHIPCGRFKV---ARKDSINSVNIFSHVVDCDRVA-R 477
+ W+ K++ E I CGRFK+ + + N+ N SH V V
Sbjct: 527 LTDCWV------PKKVIRWEDKDMIISCGRFKINPSGKLCTYNNTNSVSHQVHASAVRNN 688
Query: 478 EVYKIYPRKGSVWALY--GEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEK 535
+ Y+IYPRKG +WALY TL + K C +V +T +M + +LEK
Sbjct: 689 KEYEIYPRKGEIWALYRGWRTTLK------RSDLKNCEYDIVEVTEDADM-WTDVLFLEK 847
Query: 536 VDGYKTVFKRQ--DTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPAS 593
V GY +VFK + + GS + + + SH+IPA K +E L WELDPA+
Sbjct: 848 VSGYSSVFKGKLSNGGSKMTMTIDRTELLRFSHKIPAFK--LTEEHGSNLRGFWELDPAA 1021
Query: 594 LPSYLLT 600
+P + L+
Sbjct: 1022VPHHYLS 1042
>TC80717 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical protein
{Arabidopsis thaliana}, partial (28%)
Length = 1198
Score = 106 bits (264), Expect = 3e-23
Identities = 79/309 (25%), Positives = 148/309 (47%), Gaps = 11/309 (3%)
Frame = +1
Query: 305 GEVRTLKLPIKENAVKSKRGSEVGEERSLK------KNVKLAIKE--KPEASGKRKRLEL 356
G++R LK IKE ++ S + + + + +N ++ KE +PE + R +++
Sbjct: 79 GKIRPLKW-IKEKCLQMVGCSIITQVQLVSLADVAAQNGEMRNKENAQPEKTVSRNKMKT 255
Query: 357 EECGDANG--GELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDET 414
E+ +++ D +F DF+K R + F GQ WA YD+ D MPR YA I +
Sbjct: 256 EQLNPQRKETSNPDIICCPDPEFSDFEKVRKKDCFAVGQYWAVYDNTDCMPRFYARIKKV 435
Query: 415 VSANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDR 474
S PF + +WL+ +I G + CG++++ + +FSH V C +
Sbjct: 436 HS--PFGLEYTWLEPNPVRKDEI-DWHDAGLPVACGKYRLGHSQISRDIVMFSHEVHCIK 606
Query: 475 -VAREVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYL 533
R Y +YP KG WA++ + + + + V L++++E +G+ ++YL
Sbjct: 607 GSGRGSYLVYPMKGETWAIFRHWDIGWSSEPEKNSEYQF-EFVEVLSDFDESDGVKVSYL 783
Query: 534 EKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPAS 593
KV G+ ++F++ ++ + ++ SH++P+ E + +ELDPA
Sbjct: 784 SKVKGFVSLFQQTVQNGISLCCIPPTELYRFSHRVPSFVMTG-KEREGVPSGSYELDPAG 960
Query: 594 LPSYLLTIG 602
LP + +G
Sbjct: 961 LPMSVFQVG 987
>TC79732 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical protein
{Arabidopsis thaliana}, partial (19%)
Length = 654
Score = 85.9 bits (211), Expect(2) = 2e-21
Identities = 60/205 (29%), Positives = 108/205 (52%), Gaps = 5/205 (2%)
Frame = +1
Query: 392 GQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQ-SNADGKIVSREKMGFHIPCG 450
G+ YDD DGMPR YALI + S F+++I+WL+ + + + +EK+ CG
Sbjct: 70 GRYGPVYDDIDGMPRFYALIKKVFSTG-FKLQITWLEPDPDDEEERRWVKEKL--PSACG 240
Query: 451 RFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWALYG----EATLDADGRHFA 506
++++ + + +FSH++ ++V R +K+YPRKG WAL+ + +DA+
Sbjct: 241 KYQLGKTVTTKDQPMFSHLILYEKV-RSTFKVYPRKGETWALFKNWDIKWYMDAESHQKY 417
Query: 507 DEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISH 566
D + V L++Y E G+ ++YL K+ G+ ++F R G + ++ SH
Sbjct: 418 D-----LEFVEILSDYVEGAGVFVSYLAKLKGFMSLFSRITKGGGCSFQIPPAELFRFSH 582
Query: 567 QIPARKSPCIDETPELLEDCWELDP 591
++P+ K + E + +ELDP
Sbjct: 583 RVPSFKMTGL-ERAGVPVGAFELDP 654
Score = 34.7 bits (78), Expect(2) = 2e-21
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +3
Query: 374 DSDFYDFDKDRVERSFKKGQVWA 396
D +F DFDKD+ E F GQ+WA
Sbjct: 15 DPEFSDFDKDKKEECFASGQIWA 83
>BG585858 similar to GP|15983799|gb At2g05250/F5G3.15 {Arabidopsis thaliana},
partial (19%)
Length = 851
Score = 95.9 bits (237), Expect = 4e-20
Identities = 65/186 (34%), Positives = 96/186 (50%), Gaps = 23/186 (12%)
Frame = +3
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------ 60
+EEAL+ AE + + + A YA +A+ L P LEGIS++VT+ + A+
Sbjct: 300 KEEALKAIENAEKRF-SHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVTCNG 476
Query: 61 --DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
DWY+++G+ P N AV+KQYKK++ LLHPD N ++ AF LV EA+ LS S
Sbjct: 477 ELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNK---CVGADGAFHLVSEAWARLSGS 647
Query: 119 ALKK-----GYDAELRKK----------EAPTFWTACSACRLLHQFERRYVGHSLVCPNC 163
K G + K TFWT C++ ++ +++ R+YV L C NC
Sbjct: 648 YDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSWQVQYEYLRKYVNKKLSCKNC 827
Query: 164 NKSFEA 169
F A
Sbjct: 828 RGIFIA 845
>BE203683 weakly similar to GP|14587221|d contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (12%)
Length = 565
Score = 89.7 bits (221), Expect = 3e-18
Identities = 53/131 (40%), Positives = 79/131 (59%), Gaps = 9/131 (6%)
Frame = +3
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATD----- 61
+EEALR K +AE K++ + + A K+A +AQRL+P LE I++M+ + + +
Sbjct: 171 KEEALRAKEIAEKKME-NRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQKVFG 347
Query: 62 ----WYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSD 117
WY +L +E A ++KQ++K +L LHPDKN A +E AFKL+GEA RVLSD
Sbjct: 348 DEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNK---FAGAEAAFKLIGEAQRVLSD 518
Query: 118 SALKKGYDAEL 128
+ YD +L
Sbjct: 519 REKRTRYDMKL 551
>TC89822 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12
{Arabidopsis thaliana}, partial (35%)
Length = 539
Score = 78.2 bits (191), Expect = 9e-15
Identities = 61/167 (36%), Positives = 85/167 (50%), Gaps = 18/167 (10%)
Frame = +2
Query: 1 MAPSKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT 60
M SK E E RL + E LQ + K + + A Q P LEG +++ + +L A
Sbjct: 53 MNTSKAEAE--RLLEIGEELLQ-KRDLKGSREIANLVQETEPLLEGSDQILAIVDVLEAA 223
Query: 61 -----------DWYTVLGVEPFANS-NAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLV 108
DWY VL ++ + N ++KQY+ L+LLLHPDKN A E AFKLV
Sbjct: 224 EKPLNLNNHHLDWYAVLQIDRNSQDLNRIKKQYRTLALLLHPDKNPFSYA---ELAFKLV 394
Query: 109 GEAFRVLSDSALK----KGYDAELRKK--EAPTFWTACSACRLLHQF 149
+A+ VLSD K KG++ EL K FWTAC C ++++
Sbjct: 395 KDAWAVLSDPVQKAQYDKGFEFELLGKGNGNVNFWTACPYCYHMYEY 535
>TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12
{Arabidopsis thaliana}, partial (29%)
Length = 645
Score = 70.5 bits (171), Expect = 2e-12
Identities = 40/106 (37%), Positives = 65/106 (60%), Gaps = 9/106 (8%)
Frame = +1
Query: 32 KYAKRAQRLFPELEGISEMVTSLTILSAT--------DWYTVLGVEPFANS-NAVRKQYK 82
++A AQ P LEG +++ + +L A+ DWY++L ++ ++ + ++KQY+
Sbjct: 178 EFAILAQETEPLLEGSDQILAIIDVLIASEKRVNNNPDWYSILQIDRRSDDLDLIKKQYR 357
Query: 83 KLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKGYDAEL 128
+L+LLLHPDK+ H A + AFKLV +A+ VLSD K YD +L
Sbjct: 358 RLALLLHPDKSRFHFA---DHAFKLVADAWAVLSDPVKKSHYDKDL 486
>AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (5%)
Length = 510
Score = 64.3 bits (155), Expect = 1e-10
Identities = 35/98 (35%), Positives = 56/98 (56%), Gaps = 11/98 (11%)
Frame = +2
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTI---------- 56
+EEALR K +AE K++ S + A +A +AQ+L+P+LE I++M+ +
Sbjct: 197 KEEALRAKDIAEKKME-SKDFTGARTFAHKAQKLYPDLENIAQMLVVCDVHCSAEQKLLG 373
Query: 57 -LSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKN 93
+ DWY VL ++ + + KQY+K +L LH DKN
Sbjct: 374 NTNVVDWYKVLQIDRNDHDGIINKQYRKFALQLHTDKN 487
>TC77865 similar to GP|17473565|gb|AAL38258.1 dnaJ-like protein {Arabidopsis
thaliana}, partial (70%)
Length = 1393
Score = 63.9 bits (154), Expect = 2e-10
Identities = 33/65 (50%), Positives = 44/65 (66%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
++Y +LGVE + VRK Y+KLSL +HPDKN A +EEAFKLV +AF+ LS+
Sbjct: 475 NYYDILGVEKSCTVDDVRKSYRKLSLKVHPDKNK---APGAEEAFKLVSKAFQCLSNEES 645
Query: 121 KKGYD 125
K+ YD
Sbjct: 646 KRKYD 660
>BG456490 weakly similar to GP|22296361|dbj AHM1(AT hook-containing MAR
binding protein1)-like protein {Oryza sativa (japonica
cultivar-group)}, partial (6%)
Length = 531
Score = 63.5 bits (153), Expect = 2e-10
Identities = 56/171 (32%), Positives = 79/171 (45%), Gaps = 14/171 (8%)
Frame = +3
Query: 73 NSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKGYDAELRKKE 132
N + R+ Y K ++LL P + +EA V EA+ VLSD + YD
Sbjct: 12 NRDLARQHYAKFAILLDPTSSEKF--PFQDEALARVREAWHVLSDPGKRTVYDRGGGVTA 185
Query: 133 APT--FWTACSACRLLHQFERRYVGHSLVCPNCNKSFE--AVEAVLSDGSS-DEGEKVGV 187
A T FWTAC C L+Q+E++Y SL+C C K+F AV++ + G++ EGE+
Sbjct: 186 ATTAAFWTACPYCWNLYQYEKKYEDCSLMCQTCMKTFHGVAVKSPVKVGATVVEGEEKRQ 365
Query: 188 RRNEKREVVLG-VAAPNGVKGKVGGEKGLKGKVGDGDG--------VGDVG 229
K V L NG + +G + V D DG VGD G
Sbjct: 366 YYKCKARVPLKFYEVKNGDESLMGENEAEFVYVSDDDGDWEKEWGNVGDAG 518
>BF631720 weakly similar to GP|10798648|em putative DNAJ protein {Nicotiana
tabacum}, partial (25%)
Length = 630
Score = 61.2 bits (147), Expect = 1e-09
Identities = 30/66 (45%), Positives = 45/66 (67%)
Frame = +3
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
T++YT LGV+P A+ + ++K Y+KL++ HPDKN + A EE FK + EA+ VLSD
Sbjct: 60 TEYYTRLGVQPNASQDEIKKAYRKLAVKYHPDKNPGNKEA--EEMFKKISEAYEVLSDEQ 233
Query: 120 LKKGYD 125
++ YD
Sbjct: 234 KRQMYD 251
>TC90708 weakly similar to PIR|S26703|S26703 dnaJ protein homolog YDJ1 -
yeast (Saccharomyces cerevisiae), partial (43%)
Length = 1098
Score = 57.4 bits (137), Expect = 2e-08
Identities = 32/81 (39%), Positives = 46/81 (56%)
Frame = +2
Query: 45 EGISEMVTSLTILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEA 104
+G+ + V + ++ T Y LGV+P AN +RK YK +L HPDKN + A EE
Sbjct: 224 KGL*KSVPTYKMVKETKLYDTLGVKPEANDQELRKAYKTNALKYHPDKNAHNPEA--EEK 397
Query: 105 FKLVGEAFRVLSDSALKKGYD 125
FK + A+ +LSDS + YD
Sbjct: 398 FKEISHAYEILSDSQKRAVYD 460
>TC80183 similar to GP|4508077|gb|AAD21421.1| 62114 {Arabidopsis thaliana},
partial (59%)
Length = 1063
Score = 56.6 bits (135), Expect = 3e-08
Identities = 35/96 (36%), Positives = 48/96 (49%)
Frame = +2
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D Y +LGV ANS+ ++K Y KLSL HPDKN S + F + A+ +L D A
Sbjct: 518 DCYDLLGVTQSANSSEIKKAYYKLSLKHHPDKNPD---PESRKIFVKIANAYEILKDEAT 688
Query: 121 KKGYDAELRKKEAPTFWTACSACRLLHQFERRYVGH 156
++ YD + E + TA Q+ R Y GH
Sbjct: 689 REKYDYAIAHPEEVFYNTA--------QYYRAYYGH 772
>BQ138754 weakly similar to GP|14587221|db contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (4%)
Length = 639
Score = 56.6 bits (135), Expect = 3e-08
Identities = 29/109 (26%), Positives = 54/109 (48%)
Frame = -2
Query: 316 ENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEECGDANGGELEVMAVVDS 375
+NA SKR + +K+ ++ K ++++C A+ E + D+
Sbjct: 467 KNARTSKRSKPSMSAEDIVSILKVKVETSNLTEVKDSLDDMDDC-HASASTPEAFEIPDA 291
Query: 376 DFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRI 424
F++F+ R F+ GQ+WA Y D+DGMP++Y I + V+ E+ +
Sbjct: 290 QFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTGPTIELHV 144
>TC90148 similar to GP|6403503|gb|AAF07843.1| putative DnaJ protein
{Arabidopsis thaliana}, partial (36%)
Length = 894
Score = 55.5 bits (132), Expect = 6e-08
Identities = 30/76 (39%), Positives = 42/76 (54%)
Frame = +2
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D Y VLGV+ A+ ++K + KLSL HPDKN A ++E F + A+ +LSD
Sbjct: 149 DPYKVLGVDKSASQREIQKAFHKLSLQYHPDKNK---AKGAQEKFAQINNAYEILSDEQK 319
Query: 121 KKGYDAELRKKEAPTF 136
+K YD +K P F
Sbjct: 320 RKNYDLYGDEKGNPGF 367
>TC77020 similar to PIR|D96795|D96795 probable DnaJ protein 19794-17391
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1848
Score = 53.9 bits (128), Expect = 2e-07
Identities = 27/71 (38%), Positives = 45/71 (63%)
Frame = +3
Query: 56 ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
++ +++Y VLGV P A+ ++K Y + +HPDKN AA + F+++GEA++VL
Sbjct: 213 MVKESEYYDVLGVSPTASEAEIKKAYYIKARQVHPDKNPNDPLAA--QNFQVLGEAYQVL 386
Query: 116 SDSALKKGYDA 126
SD ++ YDA
Sbjct: 387 SDPTQRQAYDA 419
>TC90060 similar to GP|23428796|gb|AAM23263.1 DnaJ-like protein {Glycine
max}, partial (90%)
Length = 804
Score = 53.1 bits (126), Expect = 3e-07
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Frame = +1
Query: 62 WYTVLGVEPFANSNAVRKQYKKLSLLLHPDK--NNTHVAAASEEAFKLVGEAFRVLSDSA 119
+Y+VLG+ A+S+ +R Y+KL++ HPDK N A ++ F+ + EA+ VLSD +
Sbjct: 241 YYSVLGIRSDASSSDIRTAYRKLAMRWHPDKFARNPTTAGEAKRRFQQIQEAYSVLSDES 420
Query: 120 LKKGYDAEL 128
+ YDA L
Sbjct: 421 KRSMYDAGL 447
>TC80113 weakly similar to GP|21554000|gb|AAM63081.1 unknown {Arabidopsis
thaliana}, partial (17%)
Length = 1166
Score = 53.1 bits (126), Expect = 3e-07
Identities = 39/126 (30%), Positives = 64/126 (49%), Gaps = 10/126 (7%)
Frame = +2
Query: 19 SKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSA---------TDWYTVLGVE 69
+KL ++ + A +A RA+ P + ++ + L A DWY +L +
Sbjct: 197 NKLLSARDLHGARSFAIRARESDPTFDASELLLAVIDTLLAGESRINDHHRDWYGILQIL 376
Query: 70 PFA-NSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKGYDAEL 128
+ N + + QY++L+LLL P++N A S AF LV +A+ VLS+ A K YD++L
Sbjct: 377 RYTTNIDHIANQYRRLALLLDPNRNPF---AFSGHAFSLVHDAWSVLSNPAKKAMYDSDL 547
Query: 129 RKKEAP 134
R P
Sbjct: 548 RLLTTP 565
Score = 34.7 bits (78), Expect = 0.11
Identities = 12/36 (33%), Positives = 21/36 (58%)
Frame = +2
Query: 135 TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV 170
+FWT C C + +++ + Y +L C +C + F AV
Sbjct: 800 SFWTLCPYCYVHYEYPKEYEDCTLRCQSCRRGFHAV 907
>TC87379 similar to PIR|T04618|T04618 heat shock protein homolog F20O9.160 -
Arabidopsis thaliana, partial (75%)
Length = 1431
Score = 51.6 bits (122), Expect = 9e-07
Identities = 27/65 (41%), Positives = 40/65 (61%)
Frame = +2
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +L V+ A + ++K Y+KL++ HPDKN + A E FKL+ EA+ VLSD
Sbjct: 92 DYYEILEVDKNATDDELKKAYRKLAMKWHPDKNPDNKNDA-ETKFKLISEAYEVLSDPQK 268
Query: 121 KKGYD 125
+ YD
Sbjct: 269 RAIYD 283
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.134 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,436,656
Number of Sequences: 36976
Number of extensions: 211570
Number of successful extensions: 1146
Number of sequences better than 10.0: 124
Number of HSP's better than 10.0 without gapping: 1063
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1102
length of query: 606
length of database: 9,014,727
effective HSP length: 102
effective length of query: 504
effective length of database: 5,243,175
effective search space: 2642560200
effective search space used: 2642560200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0230.4


