

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0223.9
(324 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
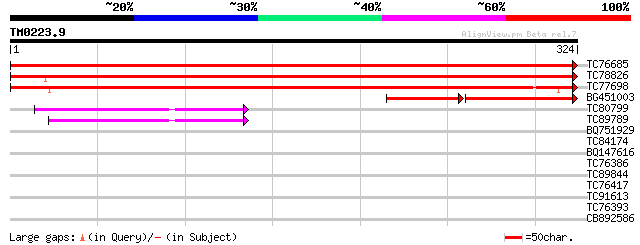
Score E
Sequences producing significant alignments: (bits) Value
TC76685 similar to PIR|A84912|A84912 probable galactinol synthas... 543 e-155
TC78826 similar to PIR|G96607|G96607 probable galactinol synthas... 514 e-146
TC77698 homologue to GP|5541885|emb|CAB51130.1 putative galactin... 490 e-139
BG451003 similar to GP|2462751|gb|A nearly identical to rice wat... 94 1e-38
TC80799 similar to GP|15810137|gb|AAL07212.1 unknown protein {Ar... 74 8e-14
TC89789 similar to GP|17065238|gb|AAL32773.1 Unknown protein {Ar... 72 2e-13
BQ751929 39 0.003
TC84174 similar to GP|18087513|gb|AAL58891.1 AT5g18480/F20L16_20... 39 0.003
BQ147616 similar to GP|18087513|gb AT5g18480/F20L16_200 {Arabido... 38 0.005
TC76386 similar to GP|12049598|emb|CAC19855. mitochondrial succi... 30 1.6
TC89844 similar to GP|3258570|gb|AAC24380.1| Unknown protein {Ar... 29 2.2
TC76417 homologue to GP|12049598|emb|CAC19855. mitochondrial suc... 29 2.2
TC91613 similar to GP|19347756|gb|AAL86302.1 unknown protein {Ar... 29 2.2
TC76393 similar to GP|12049598|emb|CAC19855. mitochondrial succi... 29 2.8
CB892586 homologue to GP|21698930|dbj casein kinase 2 catalytic ... 28 4.8
>TC76685 similar to PIR|A84912|A84912 probable galactinol synthase
[imported] - Arabidopsis thaliana, partial (79%)
Length = 1356
Score = 543 bits (1398), Expect = e-155
Identities = 250/325 (76%), Positives = 282/325 (85%), Gaps = 1/325 (0%)
Frame = +1
Query: 1 MAPDLTTAATPITTTEQPLATRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILP 60
M+PD+ TAAT IT T+ RAFVTFLAGNGDYVKGVVGLAKGLRKV+++YPLVVA+LP
Sbjct: 88 MSPDIITAATNITNTQSKATRRAFVTFLAGNGDYVKGVVGLAKGLRKVKTMYPLVVAVLP 267
Query: 61 DVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDG 120
DVP+EHR IL SQGCIVREI PVYPP+NQTQFA AYYVINYSKLRIW F EY KMIYLDG
Sbjct: 268 DVPQEHRNILTSQGCIVREIEPVYPPENQTQFAMAYYVINYSKLRIWAFEEYDKMIYLDG 447
Query: 121 DIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPP 180
DIQVFENIDHLFD+P+N FYAV DCFCE SWRHTKQY I YCQQCPDKV WP+NFGP+PP
Sbjct: 448 DIQVFENIDHLFDLPNNYFYAVMDCFCEASWRHTKQYEIGYCQQCPDKVQWPANFGPKPP 627
Query: 181 LYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVLAML 240
LYFNAG FVYEPN+ TYHDLL+ + T PTSFAEQDFLN+YF+DKYKPIPNVYNLVLAML
Sbjct: 628 LYFNAGMFVYEPNMATYHDLLQKLQVTKPTSFAEQDFLNIYFKDKYKPIPNVYNLVLAML 807
Query: 241 WRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDWNY- 299
WRHPENV+L+KVKVVHYCA GSKPWR+TG EENM R+DIKMLVKKWWD+YEDE+LD+
Sbjct: 808 WRHPENVELEKVKVVHYCAAGSKPWRYTGVEENMQREDIKMLVKKWWDVYEDESLDYKQP 987
Query: 300 INPSYTEKVLIDGGGVKNVSAPSAA 324
+N ++ +++ +K V APSAA
Sbjct: 988 VNANHLASAILEASDLKVVPAPSAA 1062
>TC78826 similar to PIR|G96607|G96607 probable galactinol synthase F25P12.95
[imported] - Arabidopsis thaliana, partial (83%)
Length = 1238
Score = 514 bits (1324), Expect = e-146
Identities = 242/328 (73%), Positives = 278/328 (83%), Gaps = 4/328 (1%)
Frame = +3
Query: 1 MAPDLTTAATPITTTEQPL---ATRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVA 57
MAP +TT A T + L RAFVTFLAGN DY+KGVVGLAKGLRK +++YPLVVA
Sbjct: 15 MAPAITTTAVNATVEKPKLDGGKGRAFVTFLAGNADYIKGVVGLAKGLRKTKTMYPLVVA 194
Query: 58 ILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIY 117
ILPDVPEEHRKIL+SQGCIV+EI PVYPP NQT+FA AYYVINYSKLRI+EF EYSKMIY
Sbjct: 195 ILPDVPEEHRKILVSQGCIVKEIAPVYPPANQTEFAMAYYVINYSKLRIFEFEEYSKMIY 374
Query: 118 LDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGP 177
LDGDIQVFENIDHLFD+PD+ FYAV DCFCE +W HT QY I YCQQCPDKV WPSNFGP
Sbjct: 375 LDGDIQVFENIDHLFDLPDDYFYAVMDCFCEKTWSHTPQYKIGYCQQCPDKVQWPSNFGP 554
Query: 178 RPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVL 237
+PPLYFNAG FV++PN+ TYHDLL+ + T PT FAEQDFLNMYF+DKYKPIPNVYNLVL
Sbjct: 555 KPPLYFNAGMFVFQPNVATYHDLLEKVKITKPTPFAEQDFLNMYFKDKYKPIPNVYNLVL 734
Query: 238 AMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDW 297
AM+WRHPENV+L+KV+VVHYCA GSKPWR+TGEE+NMDR+DIKMLVKKW +IYEDETLD+
Sbjct: 735 AMMWRHPENVELEKVQVVHYCAAGSKPWRYTGEEQNMDREDIKMLVKKWKEIYEDETLDY 914
Query: 298 -NYINPSYTEKVLIDGGGVKNVSAPSAA 324
N + L++ GG+K++ A +AA
Sbjct: 915 NNNVRVERFTAALLEAGGLKSMPASNAA 998
>TC77698 homologue to GP|5541885|emb|CAB51130.1 putative galactinol synthase
{Pisum sativum}, partial (93%)
Length = 1563
Score = 490 bits (1262), Expect = e-139
Identities = 239/340 (70%), Positives = 274/340 (80%), Gaps = 16/340 (4%)
Frame = +3
Query: 1 MAPDLT-TAATPITTTEQPLAT--RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVA 57
MAP+L TAA +T +P+ RA+VTFLAGNGDYVKGV+GLAKGLRKV + YPLVVA
Sbjct: 153 MAPELVPTAAKSVTGFTKPVTIPKRAYVTFLAGNGDYVKGVIGLAKGLRKVMTAYPLVVA 332
Query: 58 ILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIY 117
+LPDVPEEHR++L +QGCIVREI PV PP+NQTQFA AYYVINYSKLRIWEFVEYSKMIY
Sbjct: 333 VLPDVPEEHREMLEAQGCIVREIEPVNPPENQTQFAMAYYVINYSKLRIWEFVEYSKMIY 512
Query: 118 LDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGP 177
LDGDIQV+ENIDHLFD+PD FYAV DCFCE +W HT QY I YCQQCP+KV WP G
Sbjct: 513 LDGDIQVYENIDHLFDLPDGHFYAVMDCFCERTWSHTPQYKIGYCQQCPEKVHWPKEMGQ 692
Query: 178 RPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVL 237
P LYFNAG F++EP+++TYHDLLKT + T PT FAEQDFLNMYF+D YKPIP VYNLVL
Sbjct: 693 PPSLYFNAGMFLFEPSIDTYHDLLKTLKVTPPTPFAEQDFLNMYFKDIYKPIPFVYNLVL 872
Query: 238 AMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDW 297
AMLWRHPENV+L KVKVVHYCA GSKPWR+TG+EENM R+DI+MLVKKWWDIY D +LD+
Sbjct: 873 AMLWRHPENVELHKVKVVHYCAAGSKPWRYTGKEENMQREDIRMLVKKWWDIYNDSSLDY 1052
Query: 298 NYINPSYTEKVLIDG-------------GGVKNVSAPSAA 324
N N S + +V +G G V+ V+APSAA
Sbjct: 1053NK-NLSGSGEVQTNGFEIEPFVQALSEVGRVQYVTAPSAA 1169
>BG451003 similar to GP|2462751|gb|A nearly identical to rice water stress
induced protein gp|D26537|537404 {Arabidopsis
thaliana}, partial (26%)
Length = 575
Score = 93.6 bits (231), Expect(2) = 1e-38
Identities = 40/44 (90%), Positives = 44/44 (99%)
Frame = +2
Query: 216 DFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKVVHYCA 259
DFLN+YF+DKYKPIPNVYNLVLAMLWRHPENV+L+KVKVVHYCA
Sbjct: 5 DFLNIYFKDKYKPIPNVYNLVLAMLWRHPENVELEKVKVVHYCA 136
Score = 84.0 bits (206), Expect(2) = 1e-38
Identities = 37/65 (56%), Positives = 49/65 (74%), Gaps = 1/65 (1%)
Frame = +1
Query: 261 GSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDWNY-INPSYTEKVLIDGGGVKNVS 319
GSKPWR+TG EENM R+DIKMLV KWWD+YEDE+LD+ +N ++ +++ +K V
Sbjct: 229 GSKPWRYTGVEENMQREDIKMLVXKWWDVYEDESLDYKQPVNANHLASAILEASDLKVVP 408
Query: 320 APSAA 324
APSAA
Sbjct: 409 APSAA 423
>TC80799 similar to GP|15810137|gb|AAL07212.1 unknown protein {Arabidopsis
thaliana}, partial (41%)
Length = 1723
Score = 73.9 bits (180), Expect = 8e-14
Identities = 42/122 (34%), Positives = 63/122 (51%)
Frame = +3
Query: 15 TEQPLATRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKILLSQG 74
T + L +A+ T L YV G + A+ +R S LV+ + + HR L + G
Sbjct: 1092 TMEMLQEKAYATILHSAHVYVCGAIAAAQSIRMSGSTRDLVILVDETISGYHRSGLEAAG 1271
Query: 75 CIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDM 134
VR I + PK + AY NYSK R+W+ +Y K+I++D D+ + NID LF M
Sbjct: 1272 WKVRTIKRIRNPKAEKD---AYNEWNYSKFRLWQLTDYDKIIFIDADLLILRNIDFLFGM 1442
Query: 135 PD 136
P+
Sbjct: 1443 PE 1448
>TC89789 similar to GP|17065238|gb|AAL32773.1 Unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 848
Score = 72.4 bits (176), Expect = 2e-13
Identities = 38/114 (33%), Positives = 61/114 (53%)
Frame = +1
Query: 23 AFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKILLSQGCIVREIVP 82
A+ T L YV G + A+ +R S LV+ + + + HR L + G + I
Sbjct: 193 AYATILHSAHIYVCGAITAAQSIRMSGSTRDLVILVDETISDYHRSGLAAAGWKIHTIQR 372
Query: 83 VYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPD 136
+ PK + + AY NYSK R+W+ +Y K+I++D D+ + NID LF+MP+
Sbjct: 373 IRNPKAEPE---AYNEWNYSKFRLWQLTDYDKIIFIDADLLILRNIDFLFEMPE 525
>BQ751929
Length = 771
Score = 38.9 bits (89), Expect = 0.003
Identities = 34/139 (24%), Positives = 55/139 (39%)
Frame = +3
Query: 93 AHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWR 152
A A + + +K +E +Y +++ D D QV N+DHLF P AV + W
Sbjct: 396 ASATWADSLNKFHAFELTDYKRVLIFDADTQVLNNMDHLFQQP-KAAVAVPRAY----WL 560
Query: 153 HTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSF 212
+ K P ++ + EPN T HD+L + + T
Sbjct: 561 NEKD-------------------TPPAEQVLSSHVMLLEPNATT-HDVL--VKESLRTGK 674
Query: 213 AEQDFLNMYFRDKYKPIPN 231
+ D +N FRD +P+
Sbjct: 675 LDMDVVNTMFRDTAMILPH 731
>TC84174 similar to GP|18087513|gb|AAL58891.1 AT5g18480/F20L16_200
{Arabidopsis thaliana}, partial (38%)
Length = 673
Score = 38.9 bits (89), Expect = 0.003
Identities = 30/131 (22%), Positives = 52/131 (38%)
Frame = +2
Query: 103 KLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYC 162
+L+I+ Y+K++YLD D V NI+ LF FC + +H+++
Sbjct: 5 QLKIFNMTNYNKVVYLDADTIVVRNIEELFKCGK---------FC-ANLKHSER------ 136
Query: 163 QQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYF 222
N+G V EP+ ++D++ + + +Q FLN Y
Sbjct: 137 --------------------LNSGVMVVEPSTTLFNDMMSKVKTLPSYTGGDQGFLNSY- 253
Query: 223 RDKYKPIPNVY 233
Y PN +
Sbjct: 254 ---YSGFPNAH 277
>BQ147616 similar to GP|18087513|gb AT5g18480/F20L16_200 {Arabidopsis
thaliana}, partial (39%)
Length = 712
Score = 38.1 bits (87), Expect = 0.005
Identities = 27/109 (24%), Positives = 45/109 (40%), Gaps = 27/109 (24%)
Frame = +1
Query: 184 NAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRD----------------KYK 227
N+G V EP+ ++D++ + T + +Q FLN Y+ D K +
Sbjct: 172 NSGVMVVEPSETLFNDMVNKIKTTKSYTGGDQGFLNSYYSDFPNARVFDPNLSPEELKSR 351
Query: 228 PIPNVYNL-----------VLAMLWRHPENVQLDKVKVVHYCAFGSKPW 265
P+P + L +LA W E +++V+HY KPW
Sbjct: 352 PVPEMERLSTLYNADVGLYMLANKWMVDEK----ELRVIHYTLGPLKPW 486
>TC76386 similar to GP|12049598|emb|CAC19855. mitochondrial succinate
dehydrogenase iron-sulphur subunit {Arabidopsis
thaliana}, partial (60%)
Length = 707
Score = 29.6 bits (65), Expect = 1.6
Identities = 22/85 (25%), Positives = 41/85 (47%), Gaps = 3/85 (3%)
Frame = -2
Query: 21 TRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYP-LVVAILPDVP--EEHRKILLSQGCIV 77
TR+F+T + GNG V+ V +++ + P + +A P +P +E RK+ L
Sbjct: 496 TRSFITNICGNGVIVESVPASGIFVKQASPLQPSMFIAHEPQIPSRQERRKVKLGSMSFF 317
Query: 78 REIVPVYPPKNQTQFAHAYYVINYS 102
+ ++T H+ +I+YS
Sbjct: 316 -----ILIRASRTMGPHSLRLISYS 257
>TC89844 similar to GP|3258570|gb|AAC24380.1| Unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 680
Score = 29.3 bits (64), Expect = 2.2
Identities = 15/42 (35%), Positives = 21/42 (49%)
Frame = +1
Query: 230 PNVYNLVLAMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEE 271
PN +N+ +AM W + Q V HY GS+ + GEE
Sbjct: 487 PNKHNIRMAMFWL-AQGCQPGDSLVFHYSGHGSQQRNYNGEE 609
>TC76417 homologue to GP|12049598|emb|CAC19855. mitochondrial succinate
dehydrogenase iron-sulphur subunit {Arabidopsis
thaliana}, partial (81%)
Length = 1200
Score = 29.3 bits (64), Expect = 2.2
Identities = 17/54 (31%), Positives = 30/54 (55%), Gaps = 3/54 (5%)
Frame = -1
Query: 21 TRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYP-LVVAILPDVP--EEHRKILL 71
TR+F+T + GNG V+ V +++ + P + +A P +P +E RK+ L
Sbjct: 513 TRSFITNICGNGVIVESVPASGIFVKQASPLQPSMFIAHEPQIPSRQERRKVKL 352
>TC91613 similar to GP|19347756|gb|AAL86302.1 unknown protein {Arabidopsis
thaliana}, partial (57%)
Length = 657
Score = 29.3 bits (64), Expect = 2.2
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Frame = -3
Query: 246 NVQLDKVKVVHYCAFGSKPW-RFTGEEENMDRKDIKMLVKKWWDIYEDETLDWNYI 300
N + D V+ FGSKPW + + + +N+ +K IK + +W + + W Y+
Sbjct: 292 NFRFDPFLVLTLFFFGSKPWFKKSIQTQNLLKKIIKKINFFFWVYFFERRN*WGYV 125
>TC76393 similar to GP|12049598|emb|CAC19855. mitochondrial succinate
dehydrogenase iron-sulphur subunit {Arabidopsis
thaliana}, partial (37%)
Length = 637
Score = 28.9 bits (63), Expect = 2.8
Identities = 16/54 (29%), Positives = 30/54 (54%), Gaps = 3/54 (5%)
Frame = -1
Query: 21 TRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYP-LVVAILPDVP--EEHRKILL 71
TR+F+T + GNG V+ + +++ + P + +A P +P +E RK+ L
Sbjct: 622 TRSFITNICGNGVIVESIPASGIFVKQASPLQPSMFIAHEPQIPSRQERRKVKL 461
>CB892586 homologue to GP|21698930|dbj casein kinase 2 catalytic subunit
{Nicotiana tabacum}, partial (18%)
Length = 617
Score = 28.1 bits (61), Expect = 4.8
Identities = 23/68 (33%), Positives = 31/68 (44%), Gaps = 2/68 (2%)
Frame = +2
Query: 193 NLETYHDLLKTCEATTPTSFAEQDFLNMYFRDK--YKPIPNVYNLVLAMLWRHPENVQLD 250
N+ T H+L C FAE F N+YF +K K I YN + LW+ +N L
Sbjct: 341 NMHTVHNL--RCIG*RLDHFAEYQFCNIYF*NKK*LKSIVR*YNGL--FLWKGEKNFTLS 508
Query: 251 KVKVVHYC 258
V + C
Sbjct: 509 PVGIRK*C 532
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,546,995
Number of Sequences: 36976
Number of extensions: 187289
Number of successful extensions: 910
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 904
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 910
length of query: 324
length of database: 9,014,727
effective HSP length: 96
effective length of query: 228
effective length of database: 5,465,031
effective search space: 1246027068
effective search space used: 1246027068
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0223.9


