

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.5
(471 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
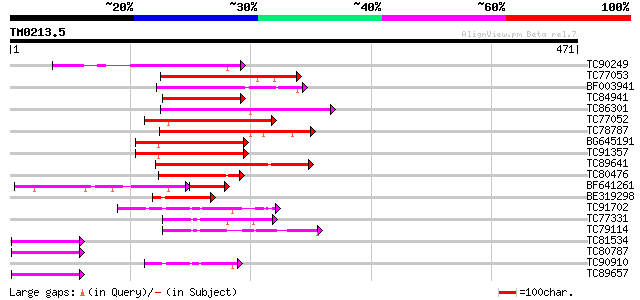
Score E
Sequences producing significant alignments: (bits) Value
TC90249 similar to PIR|T01241|T01241 probable MYB family transcr... 100 1e-21
TC77053 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 M... 97 1e-20
BF003941 homologue to GP|9759223|dbj| contains similarity to Myb... 96 2e-20
TC84941 homologue to GP|18874263|gb|AAL78741.1 MYB-like transcri... 96 3e-20
TC86301 similar to GP|8809622|dbj|BAA97173.1 Myb-related transcr... 96 4e-20
TC77052 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 M... 95 5e-20
TC78787 similar to GP|13569996|gb|AAK31280.1 putative Myb-relate... 91 1e-18
BG645191 similar to GP|2194137|gb| ESTs gb|R29947 gb|H76702 come... 91 1e-18
TC91357 similar to GP|2194137|gb|AAB61112.1| ESTs gb|R29947 gb|H... 90 2e-18
TC89641 similar to GP|9988432|dbj|BAB12698.1 hypothetical protei... 90 2e-18
TC80476 similar to GP|9988432|dbj|BAB12698.1 hypothetical protei... 88 6e-18
BF641261 similar to PIR|F96527|F96 protein F27J15.20 [imported] ... 54 1e-17
BE319298 similar to GP|17385689|dbj putative Myb-related transcr... 61 1e-09
TC91702 homologue to GP|21593278|gb|AAM65227.1 contains similari... 57 1e-08
TC77331 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~un... 57 1e-08
TC79114 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~un... 56 3e-08
TC81534 similar to PIR|T01241|T01241 probable MYB family transcr... 54 1e-07
TC80787 similar to GP|18175700|gb|AAL59913.1 unknown protein {Ar... 54 1e-07
TC90910 homologue to GP|21689883|gb|AAM67502.1 unknown protein {... 53 3e-07
TC89657 weakly similar to PIR|T08573|T08573 hypothetical protein... 53 3e-07
>TC90249 similar to PIR|T01241|T01241 probable MYB family transcription
factor [imported] - Arabidopsis thaliana, partial (44%)
Length = 1012
Score = 100 bits (249), Expect = 1e-21
Identities = 61/163 (37%), Positives = 88/163 (53%), Gaps = 2/163 (1%)
Frame = +3
Query: 36 GRLPAEIMERYRLLVLHVDNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGP 95
G+ +++++YR LV V IE G P +P +SS
Sbjct: 582 GKTVFDVIKQYRELVEDVSEIEAGNVP----------IPGYLASS--------------- 686
Query: 96 APDFIVDRVPEPVPNDPSSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQT 155
V E D + + +T + S ++ +K V WT++EHR FL GL KYGKG W+
Sbjct: 687 ----FTFEVVEKQNYDGNRRRHVTVRGSDHERKKGVPWTEEEHRRFLMGLLKYGKGDWRN 854
Query: 156 ISREFLPSKTPTQIASHAQKYYLR--LNSTPKRRKRASIHDLT 196
ISR F+ +KTPTQ+ASHAQKYY+R ++S K ++R SIHD+T
Sbjct: 855 ISRNFVVTKTPTQVASHAQKYYIRQKVSSGGKDKRRPSIHDIT 983
>TC77053 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 Myb-like
DNA-binding domain. EST gb|Z18152 comes from this
gene., partial (43%)
Length = 794
Score = 97.1 bits (240), Expect = 1e-20
Identities = 49/132 (37%), Positives = 80/132 (60%), Gaps = 15/132 (11%)
Frame = +1
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ ++R+ WT++EH+ FL GL+K GKG W+ ISR ++ ++TPTQ+ASHAQKY+LR ++ +
Sbjct: 316 ERKRRIPWTEEEHKLFLVGLQKVGKGDWRGISRNYVKTRTPTQVASHAQKYFLRRSNLNR 495
Query: 186 RRKRASIHDLTIDDTDLVP-------QQNQVPQLNAPMQE--------LSPQQNGVQLDA 230
RR+R+S+ D+T D +P Q+ V Q A + +S + GV
Sbjct: 496 RRRRSSLFDITTDTVSAIPMEEEQVKNQDNVSQCPAAPEARKINGFPFMSVYELGVNEST 675
Query: 231 PMQQVSSQQNQV 242
PM++++ Q V
Sbjct: 676 PMEELTLGQGNV 711
>BF003941 homologue to GP|9759223|dbj| contains similarity to Myb-related
transcription factor~gene_id:K19M22.10 {Arabidopsis
thaliana}, partial (30%)
Length = 545
Score = 96.3 bits (238), Expect = 2e-20
Identities = 56/134 (41%), Positives = 80/134 (58%), Gaps = 9/134 (6%)
Frame = +3
Query: 123 SPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
+P Q K+ V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R
Sbjct: 72 APEQERKKGVPWTEEEHKLFLLGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQL 251
Query: 182 STPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSS---- 237
S K ++RASIHD+T +L + + + SP QN + L QQ +
Sbjct: 252 SGGKDKRRASIHDIT--TVNLSEKIGTCSSEDTSNRSTSP-QNSILLSHQQQQQQTSTAT 422
Query: 238 ----QQNQVDALDF 247
+ NQ +A+ F
Sbjct: 423 NFRWRNNQQNAMAF 464
>TC84941 homologue to GP|18874263|gb|AAL78741.1 MYB-like transcription
factor DIVARICATA {Antirrhinum majus}, partial (28%)
Length = 590
Score = 95.9 bits (237), Expect = 3e-20
Identities = 43/69 (62%), Positives = 56/69 (80%)
Frame = +3
Query: 128 EKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRR 187
EK V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R S K +
Sbjct: 3 EKGVPWTEEEHKLFLLGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQLSGGKDK 182
Query: 188 KRASIHDLT 196
+RASIHD+T
Sbjct: 183 RRASIHDIT 209
>TC86301 similar to GP|8809622|dbj|BAA97173.1 Myb-related transcription
activator-like {Arabidopsis thaliana}, partial (56%)
Length = 1396
Score = 95.5 bits (236), Expect = 4e-20
Identities = 53/150 (35%), Positives = 85/150 (56%), Gaps = 5/150 (3%)
Frame = +2
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ +K WT++EHR FL GL K GKG W+ I+R ++ S+TPTQ+ASHAQKY++R ++ +
Sbjct: 452 ERKKGTPWTEEEHRMFLLGLNKLGKGDWRGIARNYVISRTPTQVASHAQKYFIRQSNVSR 631
Query: 186 RRKRASIHDLTID---DTDLVPQQ-NQVPQLNAPMQELSPQQNGVQLDAPMQQV-SSQQN 240
R++R+S+ D+ D DT +VPQ QL + +P LD + + S+ N
Sbjct: 632 RKRRSSLFDIVADDAPDTSMVPQDFLSANQLQTETEGNNPLPAPPPLDEECESMDSTNSN 811
Query: 241 QVDALDFPMQQLSFQQNGDPPLNFPHQVFP 270
++ P++ S Q P+ +P P
Sbjct: 812 DGESASAPLKPDSNAQASAYPVVYPAYYSP 901
>TC77052 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 Myb-like
DNA-binding domain. EST gb|Z18152 comes from this
gene., partial (43%)
Length = 1670
Score = 95.1 bits (235), Expect = 5e-20
Identities = 42/112 (37%), Positives = 74/112 (65%), Gaps = 3/112 (2%)
Frame = +3
Query: 113 SSSQKITKKISPNQNEKR---VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQI 169
S+ + + + N++ +R + WT++EH+ FL GL+K GKG W+ ISR ++ ++TPTQ+
Sbjct: 450 SADDAVPQNSARNRDRERKRGIPWTEEEHKLFLVGLQKVGKGDWRGISRNYVKTRTPTQV 629
Query: 170 ASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSP 221
ASHAQKY+LR ++ +RR+R+S+ D+T D +P + + + + +L P
Sbjct: 630 ASHAQKYFLRRSNLNRRRRRSSLFDITTDTVSAIPMEEEQVKNQDSVSQLQP 785
>TC78787 similar to GP|13569996|gb|AAK31280.1 putative Myb-related protein
{Oryza sativa}, partial (47%)
Length = 1438
Score = 90.9 bits (224), Expect = 1e-18
Identities = 50/138 (36%), Positives = 86/138 (62%), Gaps = 8/138 (5%)
Frame = +2
Query: 125 NQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTP 184
++ +K V WT++EHR FL GL+K GKG W+ I+R F+ S+TPTQ+ASHAQKY++R ++
Sbjct: 422 SERKKGVPWTEEEHRLFLVGLQKLGKGDWRGIARNFVVSRTPTQVASHAQKYFIRQSNAT 601
Query: 185 KRRKRASIHDLTID---DTDLVPQQNQV--PQLNA-PMQELSPQQNGVQLDAPMQ--QVS 236
+R++R+S+ D+ D D+ +P++ + P N+ P E S + L + + + +
Sbjct: 602 RRKRRSSLFDMAPDVCPDSTSMPEEQVLLPPSENSQPCNEKSQPSLNLSLKSEYEPMETT 781
Query: 237 SQQNQVDALDFPMQQLSF 254
S++N +A + M F
Sbjct: 782 SEENIEEANETTMGSNGF 835
>BG645191 similar to GP|2194137|gb| ESTs gb|R29947 gb|H76702 come from this
gene. {Arabidopsis thaliana}, partial (40%)
Length = 649
Score = 90.5 bits (223), Expect = 1e-18
Identities = 41/99 (41%), Positives = 68/99 (68%), Gaps = 5/99 (5%)
Frame = +1
Query: 105 PEPVPNDPSSSQKITKKI-----SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISRE 159
P P ++P+ + ++ I + ++ V WT++EH+ FL GL++ GKG W+ ISR
Sbjct: 313 PPPQDSNPADAGYVSDDIVHASGRSRERKRGVPWTEEEHKLFLLGLQQVGKGDWRGISRN 492
Query: 160 FLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTID 198
F+ ++TPTQ+ASHAQKY+LR ++ +RR+R+S+ D+T D
Sbjct: 493 FVKTRTPTQVASHAQKYFLRRHNQNRRRRRSSLFDITTD 609
>TC91357 similar to GP|2194137|gb|AAB61112.1| ESTs gb|R29947 gb|H76702 come
from this gene. {Arabidopsis thaliana}, partial (40%)
Length = 766
Score = 90.1 bits (222), Expect = 2e-18
Identities = 41/99 (41%), Positives = 68/99 (68%), Gaps = 5/99 (5%)
Frame = +3
Query: 105 PEPVPNDPSSSQKITKKI-----SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISRE 159
P P ++P+ + ++ I + ++ V WT++EH+ FL GL++ GKG W+ ISR
Sbjct: 420 PPPQDSNPAHAGYVSHDIVHASGRSRERKRGVPWTEEEHKLFLLGLQQVGKGDWRGISRN 599
Query: 160 FLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTID 198
F+ ++TPTQ+ASHAQKY+LR ++ +RR+R+S+ D+T D
Sbjct: 600 FVKTRTPTQVASHAQKYFLRRHNQNRRRRRSSLFDITTD 716
>TC89641 similar to GP|9988432|dbj|BAB12698.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (44%)
Length = 1080
Score = 89.7 bits (221), Expect = 2e-18
Identities = 47/133 (35%), Positives = 81/133 (60%), Gaps = 2/133 (1%)
Frame = +3
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
+ + +K V WT++EHR FL GL+K GKG W+ ISR ++ ++TPTQ+ASHAQKY++RL
Sbjct: 345 VRTQERKKGVPWTEEEHRKFLVGLEKLGKGDWRGISRNYVTTRTPTQVASHAQKYFIRLA 524
Query: 182 STPKRRKRASIHDLT-IDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQ-QVSSQQ 239
+ K+++R+S+ D+ T+ N + + + + V+ DA + ++S Q
Sbjct: 525 TLNKKKRRSSLFDMVGSGKTNKTVDPNNSSKSKSG-DSVCRHDHEVEKDATLSLLINSLQ 701
Query: 240 NQVDALDFPMQQL 252
Q + D+ MQ++
Sbjct: 702 QQTKSDDYDMQKI 740
>TC80476 similar to GP|9988432|dbj|BAB12698.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (34%)
Length = 778
Score = 88.2 bits (217), Expect = 6e-18
Identities = 40/72 (55%), Positives = 57/72 (78%)
Frame = +1
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
P + +K V WT++EHR FL GL+K GKG W+ IS+ F+ S+TPTQ+ASHAQKY+LRL +T
Sbjct: 325 PQERKKGVPWTEEEHRMFLVGLEKLGKGDWRGISKNFVTSRTPTQVASHAQKYFLRL-AT 501
Query: 184 PKRRKRASIHDL 195
+++R+S+ DL
Sbjct: 502 INKKRRSSLFDL 537
>BF641261 similar to PIR|F96527|F96 protein F27J15.20 [imported] -
Arabidopsis thaliana, partial (46%)
Length = 679
Score = 54.3 bits (129), Expect(2) = 1e-17
Identities = 45/158 (28%), Positives = 79/158 (49%), Gaps = 12/158 (7%)
Frame = +3
Query: 5 WTPAQDRALELALAT---VPDDAPDRWEQIAAAVGRLPAE-IMERYRLLVLHVDNIEEGP 60
W+ +++A E A+A +D+ ++WE+IA++V E + + Y++LV V IEEG
Sbjct: 96 WSYEEEKAFENAIAMHWIEKEDSKEQWEKIASSVPNKSMEQVKQHYQVLVDDVSAIEEGH 275
Query: 61 A--PDFIVDRVPEPVPNDPSSSQKIT----KKISPNIEEGPAPDFIVDRVPEPVPNDPSS 114
P++ + E + + S K T K+ S N G + + S
Sbjct: 276 VSLPNYANEL--ETINSSNKDSSKATTSSDKRSSCNFGSG------FSGLGHDSTHHSSG 431
Query: 115 SQKITKKISPNQNEKR--VLWTDDEHRNFLRGLKKYGK 150
+++ S ++ E+R + WT++EHR FL GL K+GK
Sbjct: 432 KGGLSRSSSSSEQERRKGIPWTEEEHRLFLLGLDKFGK 545
Score = 53.1 bits (126), Expect(2) = 1e-17
Identities = 21/33 (63%), Positives = 29/33 (87%)
Frame = +2
Query: 150 KGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
KG W++ISR F+ S+TPTQ+A HAQKY++R+NS
Sbjct: 542 KGDWRSISRNFVISRTPTQVAXHAQKYFIRVNS 640
>BE319298 similar to GP|17385689|dbj putative Myb-related transcription
factor {Oryza sativa (japonica cultivar-group)}, partial
(20%)
Length = 425
Score = 60.8 bits (146), Expect = 1e-09
Identities = 29/53 (54%), Positives = 39/53 (72%)
Frame = +2
Query: 119 TKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIAS 171
T I P++ K + WT++EH FLRGL+K GKG W+ ISR+F +KTPTQ+AS
Sbjct: 263 TSTIRPSK--KGMPWTEEEHMIFLRGLEKLGKGNWRGISRDFETTKTPTQVAS 415
>TC91702 homologue to GP|21593278|gb|AAM65227.1 contains similarity to
MYB-related DNA-binding protein {Arabidopsis thaliana},
partial (28%)
Length = 676
Score = 57.4 bits (137), Expect = 1e-08
Identities = 48/143 (33%), Positives = 66/143 (45%), Gaps = 7/143 (4%)
Frame = +1
Query: 90 NIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYG 149
N+ P P VP DP+ +KI K + ++ R WTD EH FL L+ +
Sbjct: 196 NLPPPPPPPSTTAAAASTVPEDPN--KKIRKPYTITKS--RESWTDQEHDKFLEALQLFD 363
Query: 150 KGQWQTISREFLPSKTPTQIASHAQKYYLRLNST-------PKRRKRASIHDLTIDDTDL 202
+ W+ I F+ SKT QI SHAQKY+L++ + P R KR + H
Sbjct: 364 R-DWKKIEA-FVGSKTVIQIRSHAQKYFLKVQKSGTSEHVPPPRPKRKAAH--------- 510
Query: 203 VPQQNQVPQLNAPMQELSPQQNG 225
P + P+ NAP SPQ G
Sbjct: 511 -PYPQKAPK-NAP--TASPQVMG 567
>TC77331 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~unknown
protein {Arabidopsis thaliana}, partial (25%)
Length = 1835
Score = 57.4 bits (137), Expect = 1e-08
Identities = 39/116 (33%), Positives = 57/116 (48%), Gaps = 21/116 (18%)
Frame = +1
Query: 128 EKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLR-------- 179
++R WTD+EH+ FL LK YG+G W+ I E + SKT QI SHAQK++ +
Sbjct: 355 KQREKWTDEEHQKFLEALKLYGRG-WRQI-EEHIGSKTAIQIRSHAQKFFSKVVREPDGS 528
Query: 180 -------LNSTPKRRKRASIHDLTIDDTD-----LVPQQNQV-PQLNAPMQELSPQ 222
++ P R KR +H D LVP +++ P +N + E Q
Sbjct: 529 AESPIQPIDIPPPRPKRKPLHPYPRKSVDSFKGQLVPNESETSPSINLSVAENDTQ 696
>TC79114 similar to GP|10177075|dbj|BAB10517. gene_id:MKP11.15~unknown
protein {Arabidopsis thaliana}, partial (32%)
Length = 759
Score = 55.8 bits (133), Expect = 3e-08
Identities = 40/139 (28%), Positives = 67/139 (47%), Gaps = 6/139 (4%)
Frame = +1
Query: 128 EKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRR 187
++R WTD+EH+ FL LK YG+ W+ I E + +KT QI SHAQK++ ++N
Sbjct: 271 KQREKWTDEEHKKFLEALKLYGRA-WRKI-EEHVGTKTAVQIRSHAQKFFSKIN------ 426
Query: 188 KRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDF 247
D +DT +V + ++P + + P P + V +N++ L+
Sbjct: 427 -----RDTDGNDTTMV-ETIEIPPPRPKRKPIHPY--------PRKLVEIPKNEISNLEQ 564
Query: 248 PMQQLSF------QQNGDP 260
P++ S Q+N P
Sbjct: 565 PLRSNSLVSLDFGQENNSP 621
>TC81534 similar to PIR|T01241|T01241 probable MYB family transcription
factor [imported] - Arabidopsis thaliana, partial (30%)
Length = 1227
Score = 53.9 bits (128), Expect = 1e-07
Identities = 26/62 (41%), Positives = 37/62 (58%), Gaps = 1/62 (1%)
Frame = +2
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S WTPA+++ E ALA D PDRW ++AA + G+ ++M +Y+ L V NIE G
Sbjct: 890 STKWTPAENKLFENALAVYDKDTPDRWHKVAAMIPGKSVVDVMNQYKELEADVCNIEAGL 1069
Query: 61 AP 62
P
Sbjct: 1070VP 1075
>TC80787 similar to GP|18175700|gb|AAL59913.1 unknown protein {Arabidopsis
thaliana}, partial (22%)
Length = 802
Score = 53.9 bits (128), Expect = 1e-07
Identities = 29/62 (46%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Frame = +3
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S W+ QD+A E AL T P+D DRWE+IA V G+ EI + Y LLV + IE G
Sbjct: 417 SSEWSWEQDKAFENALVTYPEDDSDRWEKIAVDVPGKTMEEIKQHYELLVDDIGQIEAGC 596
Query: 61 AP 62
P
Sbjct: 597 VP 602
Score = 29.3 bits (64), Expect = 3.4
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +1
Query: 123 SPNQNEKRVLWTDDEHRNFLRGL 145
S + K + WT++EHR FL GL
Sbjct: 733 SDQERRKGIAWTEEEHRLFLSGL 801
>TC90910 homologue to GP|21689883|gb|AAM67502.1 unknown protein {Arabidopsis
thaliana}, partial (32%)
Length = 683
Score = 52.8 bits (125), Expect = 3e-07
Identities = 32/88 (36%), Positives = 50/88 (56%), Gaps = 7/88 (7%)
Frame = +2
Query: 113 SSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASH 172
+S +K+ K + ++ R W+D+EH FL L+ + + W+ I +F+ SKT QI SH
Sbjct: 326 ASGKKVRKPYTITKS--RESWSDEEHDKFLEALQLFDR-DWKKIE-DFVGSKTVIQIRSH 493
Query: 173 AQKYYLRLNST-------PKRRKRASIH 193
AQKY+L++ P R KR +IH
Sbjct: 494 AQKYFLKVQKNGTLAHVPPPRPKRKAIH 577
>TC89657 weakly similar to PIR|T08573|T08573 hypothetical protein T22F8.150
- Arabidopsis thaliana, partial (60%)
Length = 909
Score = 52.8 bits (125), Expect = 3e-07
Identities = 28/62 (45%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Frame = +1
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
+ W+ QD+A E AL T P+D DRWE+IA V G+ EI + Y LLV + IE G
Sbjct: 634 TSEWSWEQDKAFENALVTYPEDDSDRWEKIAVDVPGKTMEEIKQHYELLVDDIGQIEAGC 813
Query: 61 AP 62
P
Sbjct: 814 VP 819
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,924,103
Number of Sequences: 36976
Number of extensions: 248859
Number of successful extensions: 1257
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 972
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1221
length of query: 471
length of database: 9,014,727
effective HSP length: 100
effective length of query: 371
effective length of database: 5,317,127
effective search space: 1972654117
effective search space used: 1972654117
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0213.5


