

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.15
(1070 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
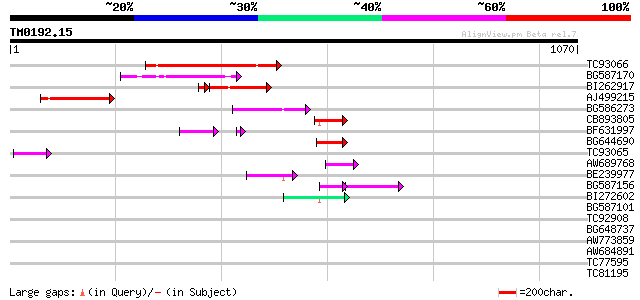
Score E
Sequences producing significant alignments: (bits) Value
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 192 6e-49
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 127 2e-29
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 107 2e-25
AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 102 8e-22
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 87 3e-17
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 63 7e-10
BF631997 weakly similar to GP|18542925|gb Putative pol polyprote... 52 1e-07
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 55 1e-07
TC93065 49 8e-06
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 49 1e-05
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 47 3e-05
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 40 4e-05
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 44 4e-04
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 42 0.001
TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol po... 42 0.001
BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberos... 41 0.002
AW773859 41 0.002
AW684891 40 0.004
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 37 0.041
TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Ar... 34 0.34
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 192 bits (488), Expect = 6e-49
Identities = 108/260 (41%), Positives = 159/260 (60%), Gaps = 4/260 (1%)
Frame = +1
Query: 257 EACERCLL-GMQRRVSFSTCKEWRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSR 315
E C+ L G +++VSFST R K +L+ IH+D+ GP + +S+ RY + DDF R
Sbjct: 16 EFCKHLLFFGNRKKVSFSTATH-RTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPR 192
Query: 316 MTWVYFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQ 375
WVYFL+ K+E F FKK++ LVE Q+GK++K L +D E+ S +F++FC + GI R
Sbjct: 193 KVWVYFLRYKNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARH 372
Query: 376 LTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHN-TFWWAEAVYTAVYILNRCPTNAVQ 434
T+ PQQNGV+ER RT++E AR ML G+ N W EA TA +++NR P +A+
Sbjct: 373 KTIPRNPQQNGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALD 552
Query: 435 NKTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVY- 493
K P + W G +LR+FG Y V D KL + IFL Y+++SKGYR++
Sbjct: 553 FKVPEDIWSGNLVDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWC 723
Query: 494 -NLQTKKLTISRDVEVDGNA 512
+ +++KL +SRDV + +A
Sbjct: 724 SDPKSQKLILSRDVTFNEDA 783
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 127 bits (319), Expect = 2e-29
Identities = 79/231 (34%), Positives = 119/231 (51%), Gaps = 3/231 (1%)
Frame = -3
Query: 209 YVANIAMKAQVDDSWLWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQR 268
Y + + ++ LWH R GH + + L L+ LP + N+ CE C+LG
Sbjct: 629 YKCSFTSSSSLNKDALWHARLGHPHGRALNLM--------LPGVVFENKNCEACILGKHC 474
Query: 269 RVSF---STCKEWRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAK 325
+ F ST E + +LI+TD+ + S DN++YF+ F D+ S+ TW+ + +K
Sbjct: 473 KNVFPRTSTVYE----NCFDLIYTDLW-TAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSK 309
Query: 326 SEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQN 385
V FK F+A V IK+LRSD G EYTS F + GI Q + Y PQQN
Sbjct: 308 DRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQN 129
Query: 386 GVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNK 436
GV++RK + +ME+ARS++ + V TA Y++N PT ++ K
Sbjct: 128 GVAKRKNKHLMEVARSLMFQAN----------VSTACYLINWIPTKVLRIK 6
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 107 bits (268), Expect(2) = 2e-25
Identities = 55/117 (47%), Positives = 74/117 (63%)
Frame = +1
Query: 378 VAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKT 437
VA+ PQQNGV+ER RT++E R+MLK GM +FW AEAV TA Y++NR P+ + KT
Sbjct: 76 VAHTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFW-AEAVKTACYVINRSPSTVIDLKT 252
Query: 438 PIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYN 494
P+E W GK L VFG Y +R KL+ K+ + IFLGY+ KGY +++
Sbjct: 253 PMEMWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
Score = 27.7 bits (60), Expect(2) = 2e-25
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = +3
Query: 357 EYTSREFDKFCEDEGIERQLT 377
EY EF FC+ EGI RQ T
Sbjct: 12 EYVDGEFLAFCKQEGIVRQFT 74
>AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (18%)
Length = 567
Score = 102 bits (254), Expect = 8e-22
Identities = 54/140 (38%), Positives = 89/140 (63%)
Frame = +3
Query: 59 KNRHQANLAEEQEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVK 118
KN+H+ + +EQ LL + + ++ W +DSGC++HM D +FK ++ S K
Sbjct: 21 KNKHKPHQ*GSKEQ---LLAMSTFATKQPSKYWLIDSGCTHHMTHDRDLFKELNKSTISK 191
Query: 119 VRLGNGAVVESKGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDV 178
VR+ NGA +E +G GTV+V++ G + IS+VL P L ++LLS+ Q++ +GY + FE +
Sbjct: 192 VRMLNGAHIEVEGIGTVLVKSHSGYKQISNVLYAPKLNQSLLSVPQLLTKGYKVLFEHEK 371
Query: 179 CRIYDKHDKRVEIAQVKMQK 198
C I D+++K V AQ+K +K
Sbjct: 372 CVIKDQNNKEVLRAQMKARK 431
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 87.4 bits (215), Expect = 3e-17
Identities = 47/147 (31%), Positives = 80/147 (53%)
Frame = -2
Query: 421 AVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIF 480
A Y++NR PT ++++ P E +KPS ++RVFG +CY VP R+KLE ++ + +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 481 LGYSTKSKGYRVYNLQTKKLTISRDVEVDGNASWNWDEEKVEKNILIPTQRPQEEVEEEA 540
+GYST KGY+ Y+ + +++ +SRDV+ + EEK ++++ T +
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYY--EEKNQEDLRDLTSDKAGVLRVIL 351
Query: 541 ENPGEPTSPPQQQEQQQDLSPESTPRR 567
E G + Q +Q + PRR
Sbjct: 350 EGLGIKMNQDQSTRSRQPEESSNEPRR 270
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 62.8 bits (151), Expect = 7e-10
Identities = 30/65 (46%), Positives = 41/65 (62%), Gaps = 4/65 (6%)
Frame = +3
Query: 576 ETCNLAIL----EPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWV 631
+T NL +L +P +FE A K E W +M E++ E+NNTWEL D G IG+KW+
Sbjct: 3 KTQNLGMLTMTSDPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWI 182
Query: 632 YKTKL 636
+KTKL
Sbjct: 183 FKTKL 197
>BF631997 weakly similar to GP|18542925|gb Putative pol polyprotein {Oryza
sativa}, partial (6%)
Length = 650
Score = 51.6 bits (122), Expect(2) = 1e-07
Identities = 27/75 (36%), Positives = 41/75 (54%)
Frame = -2
Query: 320 YFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVA 379
+ +K KSE F + + + G+ V + + G E S EF++ C++E I RQ TV
Sbjct: 643 FMMKHKSEAFYFSRNGRFWSRVKQGRR*NVFKLNNGLEICSAEFNELCKEEHITRQYTVR 464
Query: 380 YIPQQNGVSERKKRT 394
PQ+NGV+ER T
Sbjct: 463 NTPQKNGVAERMNIT 419
Score = 23.5 bits (49), Expect(2) = 1e-07
Identities = 8/17 (47%), Positives = 10/17 (58%)
Frame = -3
Query: 429 PTNAVQNKTPIEAWCGK 445
P+ A+ KTP E W K
Sbjct: 321 PSTAIDCKTPFEVWSNK 271
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 55.5 bits (132), Expect = 1e-07
Identities = 20/58 (34%), Positives = 38/58 (65%)
Frame = -3
Query: 580 LAILEPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYKTKLK 637
++ +EP++ + A + W+ +M+EE+ E++ W LV P GK +IG +WV++ KL+
Sbjct: 600 ISSIEPKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNKLE 427
Score = 29.6 bits (65), Expect = 6.5
Identities = 24/86 (27%), Positives = 37/86 (42%)
Frame = -1
Query: 679 NSSCSTKRLEHPSTRCQISFPQRCTRRRDLCGATTRFRVQRQ*KQSAKTKKSSIWLEASS 738
NS C ++ C+ ++R +C AT+ R K + + +IW EASS
Sbjct: 314 NSFCCIHGVQAVPNGCEECIY*WRSQRGGVCQATSWI*RCRGTKSCVQIE*DTIWSEASS 135
Query: 739 SSMVQSD*SVLHRSRFQEEQE*TYTI 764
SMV V FQ+ Q+ Y +
Sbjct: 134 KSMV*KAVKVSAEEWFQKRQDRQYPV 57
>TC93065
Length = 783
Score = 49.3 bits (116), Expect = 8e-06
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 3/74 (4%)
Frame = +2
Query: 8 SRNFSKNSQDKNPPCSIC*RLGHAEADCRYRDKPQCNYCKKFGHMEKYCYSKNRHQ---A 64
+++F + K PPC C + H + C YR +C C + GH+EK C +K Q A
Sbjct: 530 NQSFKSFGKKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQEA 709
Query: 65 NLAEEQEQDQYLLY 78
+ E ++D+ L+
Sbjct: 710 RVVEHHQEDEEPLF 751
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 48.5 bits (114), Expect = 1e-05
Identities = 23/61 (37%), Positives = 32/61 (51%)
Frame = +1
Query: 597 WVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYKTKLKP*WHYTKAQGEASSEGLLT 656
W++AM+ E K + N TW+LV P K IG KWVY+ K P K + ++G
Sbjct: 97 WLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQ 276
Query: 657 T 657
T
Sbjct: 277 T 279
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 47.4 bits (111), Expect = 3e-05
Identities = 30/107 (28%), Positives = 57/107 (53%), Gaps = 11/107 (10%)
Frame = -2
Query: 447 PSTKHL--RVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQTKKLTISR 504
P++ +L R+FG + H+ R K + + ++ +F+ YS+ KGYR Y+ ++K +SR
Sbjct: 480 PTSNNLTPRIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCYHPPSRKYFVSR 301
Query: 505 DVEVDGNASW-------NWDEEKVEKNILI--PTQRPQEEVEEEAEN 542
DV S+ + + K ++++L+ T P+ EVE +N
Sbjct: 300 DVTFHEQESYFGQN*SSSGGKYKEDESLLLLDLTFGPEIEVETGGDN 160
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 40.4 bits (93), Expect(2) = 4e-05
Identities = 16/53 (30%), Positives = 29/53 (54%)
Frame = -1
Query: 585 PESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYKTKLK 637
P S+E A + + W +++ E + KN+TW + P GK + +W++ K K
Sbjct: 558 PRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYK 400
Score = 25.8 bits (55), Expect(2) = 4e-05
Identities = 25/109 (22%), Positives = 48/109 (43%)
Frame = -3
Query: 635 KLKP*WHYTKAQGEASSEGLLTTT*HRLQ*DISTSGSS*YNKSSNSSCSTKRLEHPSTRC 694
+++ *W + + SS+ + + L *DI TS + +N + + + C
Sbjct: 409 QVQG*WVD*EEEN*TSSKRVYSDIWRGLH*DICTSSQATHN*NCFKLGCEPWMGIVANGC 230
Query: 695 QISFPQRCTRRRDLCGATTRFRVQRQ*KQSAKTKKSSIWLEASSSSMVQ 743
+ R T L ++TR + ++ + K+ +W EA + SMVQ
Sbjct: 229 EECISTRRT*G*SLYVSSTRS*TSSEERECTEVKEGYLWAEAITKSMVQ 83
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 43.5 bits (101), Expect = 4e-04
Identities = 30/134 (22%), Positives = 54/134 (39%), Gaps = 10/134 (7%)
Frame = +1
Query: 517 DEEKVEKNILI--PTQRPQEEVEEEAENPGEPTSPPQQQEQQQDLSPES---TPRRVRFL 571
+ +V +NI I P E+ +P P PP + + R +
Sbjct: 37 NSNEVAQNIQISSPASPTSNELANITTSPTPPPPPPPTARPHNNHTTNVGGIITRSQNNI 216
Query: 572 VDVYETCNLAI-----LEPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDII 626
V + NL + +EP + A + W M++E ++ NNTW+LV ++++
Sbjct: 217 VKTIKKLNLHVRPLCPIEPSNITQALRDHDWRSIMQDEFDALQNNNTWDLVSRSFAQNLV 396
Query: 627 GVKWVYKTKLKP*W 640
G K + + K W
Sbjct: 397 GCKLGFSYQTKSGW 438
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 42.4 bits (98), Expect = 0.001
Identities = 31/112 (27%), Positives = 59/112 (52%), Gaps = 1/112 (0%)
Frame = +2
Query: 282 DVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKS-EVFRVFKKFKALVE 340
+V ++ D GP +S NN+Y ++ D S+ WV + + + + V K FK+++
Sbjct: 305 EVFDVWGIDFMGPFPSSY--NNKYILVAVDYVSK--WVEAIASPTNDATVVVKMFKSVIF 472
Query: 341 KQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKK 392
+ G +V+ SD G + ++ F+K + G+ ++ AY PQ+ +ER K
Sbjct: 473 PRFGVP-RVVISDGGSHFINKVFEKLLKKNGVRHKVATAYHPQK---AERSK 616
>TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol polyprotein
{Oryza sativa}, partial (1%)
Length = 638
Score = 42.0 bits (97), Expect = 0.001
Identities = 19/51 (37%), Positives = 34/51 (66%)
Frame = +2
Query: 583 LEPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYK 633
+EP+++ A++ + W+ AME+EI+++E+NNT L GK + + VYK
Sbjct: 275 IEPKNYVQAAQ*QEWLAAMEQEIQVLEENNTSTLEPLREGKKWVDCRPVYK 427
>BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberosum}, partial
(10%)
Length = 804
Score = 41.2 bits (95), Expect = 0.002
Identities = 38/145 (26%), Positives = 66/145 (45%), Gaps = 5/145 (3%)
Frame = +3
Query: 490 YRVYNLQTKKLTISRDVEVD--GNASWNWDEEKVEKNIL---IPTQRPQEEVEEEAENPG 544
+RV+ + KK R ++ +++ N++ E + N + IPT + + ++
Sbjct: 393 FRVFTTRYKKQEGKRMQQIQDCSDSNSNFEIENTQDNEICEPIPTVEFDDRLISLRKDKR 572
Query: 545 EPTSPPQQQEQQQDLSPESTPRRVRFLVDVYETCNLAILEPESFEAASKQEVWVKAMEEE 604
TS P + D + +P + F+ ++ + I P S + W KA+ E
Sbjct: 573 SCTSHPICKFISYD---KLSPHHLAFVSNLDK-----IQIPNSVHESLTNP*WKKAILGE 728
Query: 605 IKMIEKNNTWELVDCPHGKDIIGVK 629
I +EKN TWEL D P GK +G K
Sbjct: 729 IHALEKNETWELSDLPSGKHPMGCK 803
>AW773859
Length = 538
Score = 41.2 bits (95), Expect = 0.002
Identities = 25/82 (30%), Positives = 36/82 (43%), Gaps = 1/82 (1%)
Frame = -3
Query: 224 LWHRRFGHFNTQVLKLLYQKNMMRDLPSLK-ESNEACERCLLGMQRRVSFSTCKEWRAKD 282
LWH R GH + + L L+ + P + + N C+ C +++ F RA
Sbjct: 230 LWHFRLGHLSNRKLLSLHS-----NFPFITIDQNSVCDICHYSRHKKLPFQLSTN-RASK 69
Query: 283 VLELIHTDVCGPMRTSSHDNNR 304
EL H D+ GP T S N R
Sbjct: 68 CYELFHFDIWGPFSTQSIHNQR 3
>AW684891
Length = 539
Score = 40.4 bits (93), Expect = 0.004
Identities = 18/45 (40%), Positives = 28/45 (62%)
Frame = -3
Query: 345 KHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSE 389
K I+ L S++GK Y + ++ E+ G+ ++L PQQNGVSE
Sbjct: 258 KSIERLCSNKGKVYVDQNLSQYLEENGVSQELKCVNSPQQNGVSE 124
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 37.0 bits (84), Expect = 0.041
Identities = 17/50 (34%), Positives = 29/50 (58%)
Frame = +2
Query: 352 SDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARS 401
SDRG + R + +FC G+ + L+ +Y PQ +G +ER + I + R+
Sbjct: 584 SDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEIQAVLRA 733
>TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 788
Score = 33.9 bits (76), Expect = 0.34
Identities = 15/52 (28%), Positives = 31/52 (58%)
Frame = +3
Query: 517 DEEKVEKNILIPTQRPQEEVEEEAENPGEPTSPPQQQEQQQDLSPESTPRRV 568
+EEK + + ++P++ +++E E E PP ++E++ + S E TPR +
Sbjct: 132 EEEKKPEETKVEEKKPEQPLKKEEEKK-ETEKPPTEEEKKPEESKEVTPREI 284
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,731,226
Number of Sequences: 36976
Number of extensions: 508928
Number of successful extensions: 3063
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 2276
Number of HSP's successfully gapped in prelim test: 90
Number of HSP's that attempted gapping in prelim test: 712
Number of HSP's gapped (non-prelim): 2443
length of query: 1070
length of database: 9,014,727
effective HSP length: 106
effective length of query: 964
effective length of database: 5,095,271
effective search space: 4911841244
effective search space used: 4911841244
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0192.15


