

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.14
(75 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
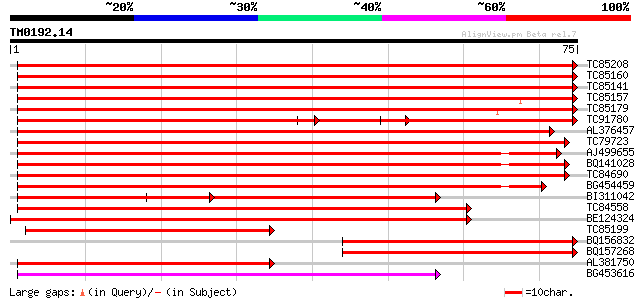
Score E
Sequences producing significant alignments: (bits) Value
TC85208 homologue to PIR|F86214|F86214 protein T6D22.2 [imported... 131 6e-32
TC85160 homologue to PIR|F86214|F86214 protein T6D22.2 [imported... 131 6e-32
TC85141 homologue to PIR|F86214|F86214 protein T6D22.2 [imported... 128 4e-31
TC85157 homologue to PIR|F86214|F86214 protein T6D22.2 [imported... 126 2e-30
TC85179 similar to GP|2662343|dbj|BAA23658.1 EF-1 alpha {Oryza s... 112 2e-26
TC91780 homologue to SP|P17786|EF1A_LYCES Elongation factor 1-al... 76 1e-23
AL376457 similar to SP|P13549|EF10 Elongation factor 1-alpha so... 101 6e-23
TC79723 homologue to SP|Q01520|EF1A_PODAN Elongation factor 1-al... 100 8e-23
AJ499655 homologue to SP|P28295|EF1A_ Elongation factor 1-alpha ... 100 8e-23
BQ141028 homologue to SP|P40911|EF1A Elongation factor 1-alpha (... 97 2e-21
TC84690 similar to GP|21540004|gb|AAM54368.1 elongation factor 1... 96 3e-21
BG454459 similar to SP|P34823|EF12 Elongation factor 1-alpha (EF... 95 4e-21
BI311042 homologue to SP|P40590|RL34_ 60S ribosomal protein L34.... 55 3e-19
TC84558 weakly similar to SP|Q04634|EF1A_TETPY Elongation factor... 73 2e-14
BE124324 weakly similar to SP|Q41011|EF1A Elongation factor 1-al... 67 2e-12
TC85199 homologue to SP|O24534|EF1A_VICFA Elongation factor 1-al... 67 2e-12
BQ156832 homologue to PIR|F86214|F862 protein T6D22.2 [imported]... 53 2e-08
BQ157268 homologue to PIR|F86214|F862 protein T6D22.2 [imported]... 53 2e-08
AL381750 similar to GP|438910|gb|AA elongation factor 1-alpha {T... 51 7e-08
BG453616 homologue to GP|21539549|gb| putative guanine nucleotid... 40 2e-04
>TC85208 homologue to PIR|F86214|F86214 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (46%)
Length = 2287
Score = 131 bits (329), Expect = 6e-32
Identities = 63/74 (85%), Positives = 69/74 (93%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG +KM+PTKPMVVETF+EYP LGRFAVRDMRQTV GVIK+VEKK
Sbjct: 1589 KEIEKEPKFLKNGDAGIIKMVPTKPMVVETFAEYPPLGRFAVRDMRQTVAVGVIKAVEKK 1768
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA KK
Sbjct: 1769 DPTGAKVTKAAAKK 1810
>TC85160 homologue to PIR|F86214|F86214 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (24%)
Length = 1198
Score = 131 bits (329), Expect = 6e-32
Identities = 63/74 (85%), Positives = 69/74 (93%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG +KM+PTKPMVVETF+EYP LGRFAVRDMRQTV GVIK+VEKK
Sbjct: 453 KEIEKEPKFLKNGDAGIIKMVPTKPMVVETFAEYPPLGRFAVRDMRQTVAVGVIKAVEKK 632
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA KK
Sbjct: 633 DPTGAKVTKAAAKK 674
>TC85141 homologue to PIR|F86214|F86214 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (47%)
Length = 1881
Score = 128 bits (322), Expect = 4e-31
Identities = 63/74 (85%), Positives = 66/74 (89%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG +KMIPTKPMVVETFSEYP LGRFAVRDMRQTV GVIK VEKK
Sbjct: 1194 KEIEKEPKFLKNGDAGIIKMIPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKGVEKK 1373
Query: 62 EPSGAKVTKAALKK 75
EP AK+TK+A KK
Sbjct: 1374 EPGTAKITKSAAKK 1415
>TC85157 homologue to PIR|F86214|F86214 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (46%)
Length = 1738
Score = 126 bits (316), Expect = 2e-30
Identities = 64/75 (85%), Positives = 68/75 (90%), Gaps = 1/75 (1%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG +KMIPTKPMVVETFSEYP LGRFAVRDMRQTV GVIK+VEKK
Sbjct: 1198 KEIEKEPKFLKNGDAGIIKMIPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKAVEKK 1377
Query: 62 EPSGA-KVTKAALKK 75
+PSG K TK+ALKK
Sbjct: 1378 DPSGGLKQTKSALKK 1422
>TC85179 similar to GP|2662343|dbj|BAA23658.1 EF-1 alpha {Oryza sativa},
partial (96%)
Length = 1789
Score = 112 bits (281), Expect = 2e-26
Identities = 55/75 (73%), Positives = 65/75 (86%), Gaps = 1/75 (1%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K IE EPKFLKNGDAG +KMIPTKPMVVETFS YP LGRFAVRD+R+TV GVIK+V+KK
Sbjct: 1274 KVIENEPKFLKNGDAGIIKMIPTKPMVVETFSNYPSLGRFAVRDIRRTVAVGVIKAVKKK 1453
Query: 62 EP-SGAKVTKAALKK 75
+P +G +TK+AL+K
Sbjct: 1454 DPGAGGSITKSALQK 1498
>TC91780 homologue to SP|P17786|EF1A_LYCES Elongation factor 1-alpha
(EF-1-alpha). [Tomato] {Lycopersicon esculentum},
partial (82%)
Length = 1165
Score = 75.9 bits (185), Expect(3) = 1e-23
Identities = 35/40 (87%), Positives = 37/40 (92%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRF 41
KE+EKEPKFLKNGDAG VKMIPTKPMVVETF+EYP LG F
Sbjct: 910 KELEKEPKFLKNGDAGMVKMIPTKPMVVETFAEYPPLGSF 1029
Score = 38.1 bits (87), Expect(3) = 1e-23
Identities = 16/26 (61%), Positives = 24/26 (91%)
Frame = +3
Query: 50 VVTGVIKSVEKKEPSGAKVTKAALKK 75
++ GV+K+V++K+P+GAKVTKAA KK
Sbjct: 1056 LLLGVVKNVDQKDPTGAKVTKAAQKK 1133
Score = 30.0 bits (66), Expect(3) = 1e-23
Identities = 13/15 (86%), Positives = 13/15 (86%)
Frame = +2
Query: 39 GRFAVRDMRQTVVTG 53
GRFAVRDMRQTV G
Sbjct: 1022 GRFAVRDMRQTVAVG 1066
>AL376457 similar to SP|P13549|EF10 Elongation factor 1-alpha somatic form
(EF-1-alpha-S). [African clawed frog] {Xenopus laevis},
partial (25%)
Length = 479
Score = 101 bits (251), Expect = 6e-23
Identities = 49/71 (69%), Positives = 58/71 (81%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K +E+ PKFLK+G+A VKMIP+KPMVVE F++YP LGRFAVRDMRQTV GVIK VEKK
Sbjct: 128 KTLEEAPKFLKSGEAAMVKMIPSKPMVVEKFADYPPLGRFAVRDMRQTVAVGVIKDVEKK 307
Query: 62 EPSGAKVTKAA 72
+ KVTK+A
Sbjct: 308 AATSGKVTKSA 340
>TC79723 homologue to SP|Q01520|EF1A_PODAN Elongation factor 1-alpha
(EF-1-alpha). {Podospora anserina}, complete
Length = 1537
Score = 100 bits (250), Expect = 8e-23
Identities = 47/73 (64%), Positives = 57/73 (77%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K +E PKF+K+GDA VKM+P+KPM VE F++YP LGRFAVRDMRQTV GVIKSV+K
Sbjct: 1226 KSVENNPKFIKSGDAAIVKMVPSKPMCVEAFTDYPPLGRFAVRDMRQTVAVGVIKSVDKS 1405
Query: 62 EPSGAKVTKAALK 74
+ KVTK+A K
Sbjct: 1406 AATSGKVTKSAAK 1444
>AJ499655 homologue to SP|P28295|EF1A_ Elongation factor 1-alpha
(EF-1-alpha). [Pin mould] {Absidia glauca}, partial
(18%)
Length = 275
Score = 100 bits (250), Expect = 8e-23
Identities = 51/72 (70%), Positives = 59/72 (81%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K++E PKF+K+GDA VKMIP+KPM VE FS+YP LGRFAVRDMRQTV GVIKSVEK
Sbjct: 63 KKLEDNPKFVKSGDAAIVKMIPSKPMCVEAFSDYPPLGRFAVRDMRQTVAVGVIKSVEKV 242
Query: 62 EPSGAKVTKAAL 73
+ SG K TKAA+
Sbjct: 243 DKSG-KTTKAAM 275
>BQ141028 homologue to SP|P40911|EF1A Elongation factor 1-alpha (EF-1-alpha).
[Histoplasma capsulatum] {Ajellomyces capsulata},
partial (31%)
Length = 616
Score = 96.7 bits (239), Expect = 2e-21
Identities = 49/73 (67%), Positives = 57/73 (77%)
Frame = -2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K E PKF+K+GDA VKM+P+KPM VE F++YP LGRFAVRDMRQTV GVIKSV K
Sbjct: 402 KSTEASPKFIKSGDAAIVKMVPSKPMCVEAFTDYPPLGRFAVRDMRQTVAVGVIKSVVKA 223
Query: 62 EPSGAKVTKAALK 74
+ +G KVTKAA K
Sbjct: 222 DKAG-KVTKAAQK 187
>TC84690 similar to GP|21540004|gb|AAM54368.1 elongation factor 1-alpha
{Trichophyton rubrum}, partial (24%)
Length = 546
Score = 95.9 bits (237), Expect = 3e-21
Identities = 46/73 (63%), Positives = 55/73 (75%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K E PKF+K+GD+ VKM+P+KPM VE F++YP LGRFAVRDMRQTV GVIK+VEK
Sbjct: 111 KATEAAPKFIKSGDSAIVKMVPSKPMCVEAFTDYPPLGRFAVRDMRQTVAVGVIKAVEKS 290
Query: 62 EPSGAKVTKAALK 74
KVTK+A K
Sbjct: 291 VGGSGKVTKSAAK 329
>BG454459 similar to SP|P34823|EF12 Elongation factor 1-alpha (EF-1-alpha).
[Carrot] {Daucus carota}, partial (46%)
Length = 642
Score = 95.1 bits (235), Expect = 4e-21
Identities = 46/70 (65%), Positives = 55/70 (77%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KE+E PKF+KNGDA +VKM+P KPM VE F+ YP LGRF VRDMR TV G+IK+V KK
Sbjct: 430 KELEASPKFIKNGDAAYVKMVPXKPMCVEPFTTYPPLGRFXVRDMRXTVXVGIIKAVXKK 609
Query: 62 EPSGAKVTKA 71
+ +G KVTKA
Sbjct: 610 DTTG-KVTKA 636
>BI311042 homologue to SP|P40590|RL34_ 60S ribosomal protein L34. [Garden
pea] {Pisum sativum}, partial (75%)
Length = 754
Score = 55.5 bits (132), Expect(2) = 3e-19
Identities = 26/39 (66%), Positives = 30/39 (76%)
Frame = +2
Query: 19 VKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKS 57
++ P P VVETF+EYP LGRFAVRDMRQT GVIK+
Sbjct: 638 LRWFPPSPWVVETFAEYPPLGRFAVRDMRQTGAVGVIKA 754
Score = 54.3 bits (129), Expect(2) = 3e-19
Identities = 23/26 (88%), Positives = 25/26 (95%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPM 27
KEIEKEPKFLKNGDAG +KM+PTKPM
Sbjct: 586 KEIEKEPKFLKNGDAGIIKMVPTKPM 663
>TC84558 weakly similar to SP|Q04634|EF1A_TETPY Elongation factor 1-alpha
(EF-1-alpha) (14-nm filament-associated protein).
{Tetrahymena pyriformis}, partial (42%)
Length = 611
Score = 73.2 bits (178), Expect = 2e-14
Identities = 33/60 (55%), Positives = 42/60 (70%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K +E+ P +K GDA + + PTKPMVVE + + LGRFAVRDM+QTV GVIK + KK
Sbjct: 386 KVVEENPAHIKTGDAAMIVLTPTKPMVVEVYQNFAPLGRFAVRDMKQTVAVGVIKEITKK 565
>BE124324 weakly similar to SP|Q41011|EF1A Elongation factor 1-alpha
(EF-1-alpha). [Garden pea] {Pisum sativum}, partial
(32%)
Length = 625
Score = 66.6 bits (161), Expect = 2e-12
Identities = 34/61 (55%), Positives = 42/61 (68%)
Frame = +3
Query: 1 NKEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEK 60
+K IEK P FLKNGD +KMIPT+ MVVE FS P L RF V+D+ Q V GV +V++
Sbjct: 288 HKIIEKRPPFLKNGDNCLIKMIPTESMVVEPFSLSPPLSRFVVQDLHQIVAVGVAINVKR 467
Query: 61 K 61
K
Sbjct: 468 K 470
>TC85199 homologue to SP|O24534|EF1A_VICFA Elongation factor 1-alpha
(EF-1-alpha). [Broad bean] {Vicia faba}, partial (28%)
Length = 887
Score = 66.6 bits (161), Expect = 2e-12
Identities = 29/33 (87%), Positives = 32/33 (96%)
Frame = +3
Query: 3 EIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEY 35
EIEKEPKFLKNGDAG +KM+PTKPMVVETF+EY
Sbjct: 789 EIEKEPKFLKNGDAGIIKMVPTKPMVVETFAEY 887
>BQ156832 homologue to PIR|F86214|F862 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 662
Score = 52.8 bits (125), Expect = 2e-08
Identities = 25/31 (80%), Positives = 28/31 (89%)
Frame = +3
Query: 45 DMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
DMRQTV GVIK+VEKK+P+GAKVTKAA KK
Sbjct: 3 DMRQTVAVGVIKAVEKKDPTGAKVTKAAAKK 95
>BQ157268 homologue to PIR|F86214|F862 protein T6D22.2 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 656
Score = 52.8 bits (125), Expect = 2e-08
Identities = 25/31 (80%), Positives = 28/31 (89%)
Frame = +3
Query: 45 DMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
DMRQTV GVIK+VEKK+P+GAKVTKAA KK
Sbjct: 3 DMRQTVAVGVIKAVEKKDPTGAKVTKAAAKK 95
>AL381750 similar to GP|438910|gb|AA elongation factor 1-alpha {Trypanosoma
brucei}, partial (17%)
Length = 212
Score = 51.2 bits (121), Expect = 7e-08
Identities = 20/34 (58%), Positives = 30/34 (87%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEY 35
KE+EK+PK +K+GDA V+++P+KPM VE+FS+Y
Sbjct: 109 KELEKQPKNIKSGDAAIVRLVPSKPMCVESFSDY 210
>BG453616 homologue to GP|21539549|gb| putative guanine nucleotide
regulatory protein {Arabidopsis thaliana}, partial
(12%)
Length = 667
Score = 40.0 bits (92), Expect = 2e-04
Identities = 20/56 (35%), Positives = 32/56 (56%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKS 57
K ++K+ F+KNG ++ + +E FS++P LGRF +R +TV G I S
Sbjct: 41 KPMKKKVLFVKNGAGVVCRIQVNNSICIEKFSDFPQLGRFTLRTEGKTVAVGRINS 208
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.133 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,379,537
Number of Sequences: 36976
Number of extensions: 10438
Number of successful extensions: 50
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 50
length of query: 75
length of database: 9,014,727
effective HSP length: 51
effective length of query: 24
effective length of database: 7,128,951
effective search space: 171094824
effective search space used: 171094824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0192.14


