

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0181.13
(507 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
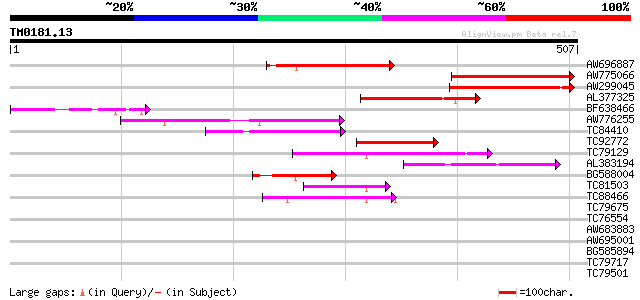
Score E
Sequences producing significant alignments: (bits) Value
AW696887 similar to PIR|T07669|T07 cyclin a1-type mitosis-speci... 121 6e-28
AW775066 similar to PIR|T07669|T07 cyclin a1-type mitosis-speci... 101 8e-22
AW299045 weakly similar to GP|18175785|gb putative cyclin {Arabi... 96 3e-20
AL377325 homologue to GP|3608181|dbj cyclin B {Pisum sativum}, p... 94 2e-19
BF638466 similar to PIR|T07675|T076 cyclin a2-type mitosis-spec... 88 9e-18
AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosi... 82 5e-16
TC84410 similar to GP|15419009|gb|AAK81695.1 cyclin A2 {Medicago... 81 1e-15
TC92772 similar to GP|19347837|gb|AAL86330.1 putative prolyl end... 77 2e-14
TC79129 similar to PIR|A96725|A96725 hypothetical protein F20P5.... 75 5e-14
AL383194 similar to GP|6031209|gb| cyclin {Pisum sativum}, parti... 73 2e-13
BG588004 similar to GP|7242793|emb| cyclin A3.1 {Pisum sativum},... 70 1e-12
TC81503 similar to GP|6448480|emb|CAB61221.1 cyclin D1 {Antirrhi... 50 3e-06
TC88466 similar to GP|6434199|emb|CAB60837.1 CycD3;2 {Lycopersic... 43 3e-04
TC79675 similar to GP|16604386|gb|AAL24199.1 At1g10700/F20B24.13... 37 0.023
TC76554 similar to GP|7295688|gb|AAF50994.1| CG11931 gene produc... 34 0.12
AW683883 similar to EGAD|126998|135 cyclin {Lupinus luteus}, par... 34 0.12
AW695001 PIR|T14004|T1 trfA protein - slime mold (Dictyostelium ... 34 0.12
BG585894 weakly similar to GP|19683051|gb| HDCKB03P (FRAGMENT). ... 34 0.15
TC79717 similar to GP|10086499|gb|AAG12559.1 Unknown Protein {Ar... 33 0.26
TC79501 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabido... 33 0.34
>AW696887 similar to PIR|T07669|T07 cyclin a1-type mitosis-specific -
soybean, partial (33%)
Length = 648
Score = 121 bits (304), Expect = 6e-28
Identities = 61/117 (52%), Positives = 85/117 (72%), Gaps = 2/117 (1%)
Frame = +3
Query: 230 DNDHMDPQLCASFACDIYKHLRASEA--KKRPSTDFMEKVQKDINTSMRAILIDWLVEVA 287
DN H + + DI +LR E K+RP ++E VQ+ I+++MR L+DWLVEVA
Sbjct: 309 DNQH----IIQPYVSDISDYLRTMEMQKKRRPMVGYLENVQRGISSNMRGTLVDWLVEVA 476
Query: 288 EEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFC 344
+EY+L+P+TL+L+V+YIDR+LS +SR KLQLLGV+SM+IASKYEEI P+ +FC
Sbjct: 477 DEYKLLPETLHLSVSYIDRFLSIEPVSRSKLQLLGVSSMLIASKYEEITPPKAVDFC 647
>AW775066 similar to PIR|T07669|T07 cyclin a1-type mitosis-specific -
soybean, partial (31%)
Length = 696
Score = 101 bits (251), Expect = 8e-22
Identities = 50/110 (45%), Positives = 73/110 (65%)
Frame = +2
Query: 396 LQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPS 455
LQ+E L NY+AELSL++Y + + PS+VAAS IFLA+FI+ P + P +S+L Y+ +
Sbjct: 5 LQMEFLCNYLAELSLIDYECIRFLPSMVAASVIFLARFIICPGVHPLTSSLSECLFYKSA 184
Query: 456 DLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFF 505
+L CV LH L+ ++L A++EKY QHK+K VA P IP+ +F
Sbjct: 185 ELEECVLILHDLYLVRRAASLKAVREKYKQHKFKNVANLPSSPEIPNHYF 334
>AW299045 weakly similar to GP|18175785|gb putative cyclin {Arabidopsis
thaliana}, partial (24%)
Length = 646
Score = 95.9 bits (237), Expect = 3e-20
Identities = 51/112 (45%), Positives = 70/112 (61%)
Frame = -1
Query: 394 PSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQ 453
PS++LE L N++AEL+LM Y L + PS++AASA+FLA++ L S PW+ TL+HY Y+
Sbjct: 637 PSVELEYLANFLAELTLMSYGFLNFFPSMIAASAVFLARWTLDQSRHPWNPTLEHYASYK 458
Query: 454 PSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFF 505
SDL V L L NS + PAI+ KY Q K+ VA P++P F
Sbjct: 457 ASDLKATVLALQNLQLNSDDCPYPAIRTKYRQSKFHGVA-VLSSPTLPETMF 305
>AL377325 homologue to GP|3608181|dbj cyclin B {Pisum sativum}, partial (45%)
Length = 329
Score = 93.6 bits (231), Expect = 2e-19
Identities = 49/110 (44%), Positives = 76/110 (68%), Gaps = 2/110 (1%)
Frame = +2
Query: 314 SRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTA 373
SR++LQL+G+++M++ASKYEEI P+V +F ++D Y E++L ME +L L++ +T
Sbjct: 2 SRRELQLVGISAMLMASKYEEIWPPEVNDFVCLSDRAYSHEQILIMEKTILGKLEWTLTV 181
Query: 374 PTIKCFLRRFVRAAQGVEEVPSLQ--LESLTNYIAELSLMEYSMLCYAPS 421
PT FL RF++AA V VPS Q LE + ++++EL +M Y+ L Y PS
Sbjct: 182 PTPFVFLVRFIKAA-SVSAVPSDQGDLEMMAHFLSELGMMHYATLRYCPS 328
>BF638466 similar to PIR|T07675|T076 cyclin a2-type mitosis-specific -
soybean, partial (9%)
Length = 566
Score = 87.8 bits (216), Expect = 9e-18
Identities = 65/141 (46%), Positives = 77/141 (54%), Gaps = 15/141 (10%)
Frame = +3
Query: 1 MSSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPL 60
MS+ N+RRSSFSSST SS+AKR + NN V+ A SAKKRPPL
Sbjct: 156 MSNXNNRRSSFSSSTTSSMAKRXPPVTXLTSENNAGKVLAA-----------SAKKRPPL 302
Query: 61 SNLTNHIASRNSSSQSLVPCVSKFAKTKKEAPP-----------KVAPALPNVKSAAAAV 109
+NLTN +SRN+SS V+K AKTKKE P P L NVKS A V
Sbjct: 303 TNLTNQNSSRNNSS----TLVTKVAKTKKEPSPCNSSSSSVIXVNKKPTLSNVKS--ATV 464
Query: 110 VFPKVAT----ATASFTERNE 126
VFPK T T+SF+ +NE
Sbjct: 465 VFPKAITTTTXTTSSFSGKNE 527
>AW776255 weakly similar to PIR|T07672|T07 cyclin a2-type mitosis-specific -
soybean, partial (22%)
Length = 704
Score = 82.0 bits (201), Expect = 5e-16
Identities = 61/208 (29%), Positives = 98/208 (46%), Gaps = 8/208 (3%)
Frame = +1
Query: 100 PNVKSAAAAVVFPKVATATASFTERNEDDAAAAASGAR-----VSVPVSTSMDVSPCKSD 154
P+ A V V +A +R G++ V+ S+ V K D
Sbjct: 127 PSRNEQAVVVGLKDVTNISAKSHKRTHTSNFQKKEGSKKRKTNVASEDDVSLQVWTIKED 306
Query: 155 GCSVSMDESMSSCDSFKSPAEIE-YVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSR 213
+S +S + K +++ V+NS +S D++ I
Sbjct: 307 AKKELAKDSSTSTMTKKESLQVQPSVENSLLSMQDTLNSPNTEI---------------- 438
Query: 214 DILVEELEK--GEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDI 271
+++ E+L G IV+ID+ D + S+A DIY ++ E ++RP ++ME +Q+DI
Sbjct: 439 NLICEKLSASVGLGIVDIDSKLRDSPIWTSYAPDIYTNIHVRECERRPLANYMETLQQDI 618
Query: 272 NTSMRAILIDWLVEVAEEYRLVPDTLYL 299
MR IL+DWLVEVA+E++LV DTLYL
Sbjct: 619 TPGMRGILVDWLVEVADEFKLVSDTLYL 702
>TC84410 similar to GP|15419009|gb|AAK81695.1 cyclin A2 {Medicago sativa},
partial (23%)
Length = 771
Score = 80.9 bits (198), Expect = 1e-15
Identities = 48/125 (38%), Positives = 67/125 (53%)
Frame = +1
Query: 176 IEYVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHMD 235
+ + DN+ + + + + IS AK + + D + K I NID D D
Sbjct: 409 VSFEDNAPSNLIGGQTSEPSAQPLISQAKAKKDS----DSKLMNAPKDPDIPNIDADIED 576
Query: 236 PQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPD 295
PQLC+ +A DIY +LR +E +FME VQ+DI MRAILIDWLVEV+ + L +
Sbjct: 577 PQLCSFYAADIYDNLRVAELSXXXHPNFMETVQRDITXXMRAILIDWLVEVSXXFNLQAN 756
Query: 296 TLYLT 300
YLT
Sbjct: 757 XXYLT 771
>TC92772 similar to GP|19347837|gb|AAL86330.1 putative prolyl endopeptidase
{Arabidopsis thaliana}, partial (14%)
Length = 531
Score = 77.0 bits (188), Expect = 2e-14
Identities = 37/73 (50%), Positives = 54/73 (73%)
Frame = +3
Query: 311 NAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFE 370
N + R++LQL+G++SM+IASKYEEI AP+V +F I+DN Y +E+VL ME +L L++
Sbjct: 312 NQVPRKELQLVGISSMLIASKYEEIWAPEVNDFVCISDNAYVREQVLVMEKTILRNLEWY 491
Query: 371 MTAPTIKCFLRRF 383
+T PT FL R+
Sbjct: 492 LTVPTPYVFLVRY 530
>TC79129 similar to PIR|A96725|A96725 hypothetical protein F20P5.7
[imported] - Arabidopsis thaliana, partial (64%)
Length = 1328
Score = 75.5 bits (184), Expect = 5e-14
Identities = 57/183 (31%), Positives = 87/183 (47%), Gaps = 5/183 (2%)
Frame = +2
Query: 254 EAKKRPSTDFMEKVQ-KDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNA 312
E K P D++ + Q + + +S R I W+++V E Y P T YL+VNY+DR+L
Sbjct: 293 EFKFVPGFDYVSRFQSRSLESSTREEAIAWILKVHEYYGFQPLTAYLSVNYMDRFLDSRP 472
Query: 313 MSRQK---LQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFK-EEVLQMESEVLNFLK 368
+ LQLL VA + +A+K EE P + +F F+ +L+ME VL L
Sbjct: 473 LPESNGWPLQLLSVACLSLAAKMEEPLVPSLLDFQIEGAKYIFQPRTILRMELLVLTILD 652
Query: 369 FEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAI 428
+ + + T FL F + T I ++ + S L Y PS +AA+AI
Sbjct: 653 WRLRSITPLSFLSFFACKLDSTGTFTHFIISRATEIILS-NIQDASFLTYRPSCIAAAAI 829
Query: 429 FLA 431
A
Sbjct: 830 LSA 838
>AL383194 similar to GP|6031209|gb| cyclin {Pisum sativum}, partial (55%)
Length = 435
Score = 73.2 bits (178), Expect = 2e-13
Identities = 46/141 (32%), Positives = 76/141 (53%), Gaps = 1/141 (0%)
Frame = +2
Query: 353 KEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLME 412
++++L ME +L LKF + APT FL RF++AAQ ++LE + ++ +L L+E
Sbjct: 2 RDQMLGMEKLILRKLKFRLNAPTPYVFLIRFLKAAQS-----DMKLEHMAFFLIDLCLVE 166
Query: 413 YSMLCYAPSLVAASAIFLAKFILFPSIKP-WSSTLQHYTLYQPSDLCVCVKELHRLFCNS 471
Y L + PSL+ AS +++A+ L I P W+ LQ + Y+ S + C + + +
Sbjct: 167 YEALAFKPSLLCASXLYVARCTL--XITPSWTPLLQKHARYEVSQIXDCADMILKFHKAA 340
Query: 472 PNSNLPAIKEKYSQHKYKYVA 492
L EKYS+ + VA
Sbjct: 341 GKGKLTVAYEKYSRKELGGVA 403
>BG588004 similar to GP|7242793|emb| cyclin A3.1 {Pisum sativum}, partial
(22%)
Length = 713
Score = 70.5 bits (171), Expect = 1e-12
Identities = 37/77 (48%), Positives = 50/77 (64%), Gaps = 2/77 (2%)
Frame = +2
Query: 218 EELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASE--AKKRPSTDFMEKVQKDINTSM 275
E+LEK E DPQLC ++ DIY +LR E + KRP D+++KVQ+D+N SM
Sbjct: 497 EKLEKFE----------DPQLCETYVSDIYDYLRNMEVDSSKRPLCDYIQKVQRDVNASM 646
Query: 276 RAILIDWLVEVAEEYRL 292
R +L+DWL EVAE+ L
Sbjct: 647 RGVLVDWLGEVAEDTSL 697
>TC81503 similar to GP|6448480|emb|CAB61221.1 cyclin D1 {Antirrhinum majus},
partial (36%)
Length = 1000
Score = 49.7 bits (117), Expect = 3e-06
Identities = 27/81 (33%), Positives = 43/81 (52%), Gaps = 3/81 (3%)
Frame = +3
Query: 263 FMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQK---LQ 319
F++ ++ R I W+++V Y P T YL VNY+DR+L+ + + LQ
Sbjct: 345 FVKMKSSSFDSDARDESIRWILKVQGYYGFQPVTAYLAVNYMDRFLNSRRLPQTNGWPLQ 524
Query: 320 LLGVASMMIASKYEEICAPQV 340
LL VA + +A+K EE P +
Sbjct: 525 LLSVACLSLAAKMEETLVPSL 587
>TC88466 similar to GP|6434199|emb|CAB60837.1 CycD3;2 {Lycopersicon
esculentum}, partial (39%)
Length = 692
Score = 42.7 bits (99), Expect = 3e-04
Identities = 31/128 (24%), Positives = 62/128 (48%), Gaps = 8/128 (6%)
Frame = +2
Query: 227 VNIDNDHMDPQLCASFACDIY---KHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWL 283
V+++N ++ C+ D++ + L++ K++ + ++ + + R I+W+
Sbjct: 41 VSLNNTTINTTTCSLLETDMFWEDEELKSLLNKEQQNPLYIFLQTNPVLETARRESIEWI 220
Query: 284 VEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQK---LQLLGVASMMIASKYEEICAPQV 340
++V Y T L VNY+DR+L +K QL VA + +A+K EE P +
Sbjct: 221 LKVNAHYSFSALTSVLAVNYLDRFLFSFRFQNEKPWMTQLAAVACLSLAAKMEETHVPLL 400
Query: 341 EEF--CYI 346
+ C+I
Sbjct: 401 LDLQVCFI 424
>TC79675 similar to GP|16604386|gb|AAL24199.1 At1g10700/F20B24.13
{Arabidopsis thaliana}, partial (45%)
Length = 907
Score = 36.6 bits (83), Expect = 0.023
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +3
Query: 26 DNNNNNNNNNNYVVGAAAVKAPAAASHSAK 55
+NNNNNNNNNN ++ V A +A+ H+ K
Sbjct: 258 NNNNNNNNNNNSSFSSSLVSASSASQHAMK 347
>TC76554 similar to GP|7295688|gb|AAF50994.1| CG11931 gene product
{Drosophila melanogaster}, partial (42%)
Length = 861
Score = 34.3 bits (77), Expect = 0.12
Identities = 15/25 (60%), Positives = 18/25 (72%)
Frame = -2
Query: 12 SSSTASSLAKRHASDNNNNNNNNNN 36
SSS S + ++S NNNNNNNNNN
Sbjct: 587 SSSNGGSGSSSNSSTNNNNNNNNNN 513
Score = 33.5 bits (75), Expect = 0.20
Identities = 15/31 (48%), Positives = 21/31 (67%)
Frame = -2
Query: 6 HRRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
+ SS S +SS + + ++NNNNNNNNNN
Sbjct: 593 YNSSSNGGSGSSSNSSTNNNNNNNNNNNNNN 501
Score = 33.5 bits (75), Expect = 0.20
Identities = 14/32 (43%), Positives = 21/32 (64%)
Frame = -2
Query: 5 NHRRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
N + S S+++S + ++NNNNNNNNNN
Sbjct: 590 NSSSNGGSGSSSNSSTNNNNNNNNNNNNNNNN 495
Score = 33.1 bits (74), Expect = 0.26
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = -2
Query: 4 NNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVV 39
N+ SS+ SS + ++NNNNNNNNN + +
Sbjct: 590 NSSSNGGSGSSSNSSTNNNNNNNNNNNNNNNNTHFI 483
Score = 29.3 bits (64), Expect = 3.7
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -2
Query: 10 SFSSSTASSLAKRHASDNNNNNNNNNN 36
+++SS+ S NNNNNNNNN
Sbjct: 596 TYNSSSNGGSGSSSNSSTNNNNNNNNN 516
>AW683883 similar to EGAD|126998|135 cyclin {Lupinus luteus}, partial (12%)
Length = 217
Score = 34.3 bits (77), Expect = 0.12
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = +2
Query: 442 WSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKY 483
W+ TL+HYT Y L C K + +P S L AI +K+
Sbjct: 26 WTDTLKHYTGYSEEQLRDCAKLMASFHSAAPESRLRAIYKKF 151
>AW695001 PIR|T14004|T1 trfA protein - slime mold (Dictyostelium discoideum),
partial (1%)
Length = 609
Score = 34.3 bits (77), Expect = 0.12
Identities = 15/34 (44%), Positives = 23/34 (67%)
Frame = +3
Query: 2 SSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNN 35
SS +++ + TA++ K+H + NNNNNNNNN
Sbjct: 318 SSQKQKKTKTNPKTATT--KKHQNKNNNNNNNNN 413
>BG585894 weakly similar to GP|19683051|gb| HDCKB03P (FRAGMENT). 10/100
{Dictyostelium discoideum}, partial (7%)
Length = 816
Score = 33.9 bits (76), Expect = 0.15
Identities = 32/105 (30%), Positives = 49/105 (46%), Gaps = 2/105 (1%)
Frame = +1
Query: 10 SFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNLTNHIAS 69
+ S S +++ + N+N NNNNN VV A + +++S PL+ S
Sbjct: 76 NLSKSPPNNINNSNCGSNSNFNNNNN--VVSTAPKQTQQKSANSRPVPKPLN------LS 231
Query: 70 RNSSSQSLVPCVSKFAKTKKEA--PPKVAPALPNVKSAAAAVVFP 112
++SSSQ VS + + A PPKV AL N+ + FP
Sbjct: 232 KSSSSQ-----VSNLGRLPQTATFPPKVNRAL-NLPNPPKTATFP 348
>TC79717 similar to GP|10086499|gb|AAG12559.1 Unknown Protein {Arabidopsis
thaliana}, partial (5%)
Length = 1579
Score = 33.1 bits (74), Expect = 0.26
Identities = 18/69 (26%), Positives = 32/69 (46%)
Frame = +2
Query: 4 NNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNL 63
N RR+SF S + DN+++NN ++ + KA A+ S + P + +
Sbjct: 1133 NGKRRNSFGSIKPEHTDQEPTKDNSSSNNTLPRFMQATQSAKAKINANSSPRSSPDVHDT 1312
Query: 64 TNHIASRNS 72
+I R+S
Sbjct: 1313 DINIKKRHS 1339
>TC79501 similar to GP|9294059|dbj|BAB02016.1 MAP kinase {Arabidopsis
thaliana}, partial (18%)
Length = 1093
Score = 32.7 bits (73), Expect = 0.34
Identities = 13/29 (44%), Positives = 20/29 (68%)
Frame = +2
Query: 7 RRSSFSSSTASSLAKRHASDNNNNNNNNN 35
+R S ++ST+ ++H N+NNNNNNN
Sbjct: 164 QRRSSTTSTSKQQQQQHQHSNDNNNNNNN 250
Score = 32.3 bits (72), Expect = 0.44
Identities = 14/29 (48%), Positives = 19/29 (65%)
Frame = +2
Query: 7 RRSSFSSSTASSLAKRHASDNNNNNNNNN 35
R S+ S+S +H++DNNNNNNN N
Sbjct: 170 RSSTTSTSKQQQQQHQHSNDNNNNNNNIN 256
Score = 31.2 bits (69), Expect = 0.98
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = +2
Query: 7 RRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
RRSS +S++ + S++NNNNNNN N
Sbjct: 167 RRSSTTSTSKQQQQQHQHSNDNNNNNNNIN 256
Score = 28.9 bits (63), Expect = 4.9
Identities = 11/27 (40%), Positives = 20/27 (73%)
Frame = +2
Query: 12 SSSTASSLAKRHASDNNNNNNNNNNYV 38
SS+T++S ++ ++N+NNNNNN +
Sbjct: 173 SSTTSTSKQQQQQHQHSNDNNNNNNNI 253
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,565,320
Number of Sequences: 36976
Number of extensions: 224513
Number of successful extensions: 3258
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 1961
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2602
length of query: 507
length of database: 9,014,727
effective HSP length: 100
effective length of query: 407
effective length of database: 5,317,127
effective search space: 2164070689
effective search space used: 2164070689
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0181.13


