

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.1
(458 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
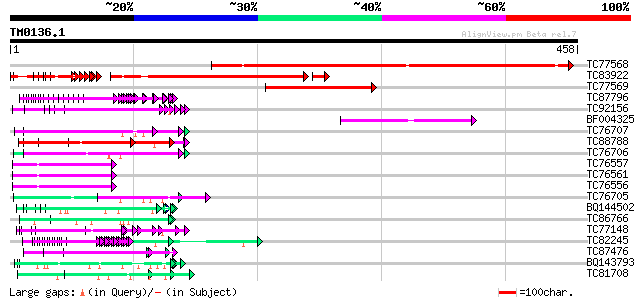
TC77568 weakly similar to GP|9857516|gb|AAG00871.1| Unknown prot... 286 2e-77
TC83922 similar to GP|15809990|gb|AAL06922.1 AT5g60530/muf9_180 ... 242 9e-68
TC77569 weakly similar to GP|9757754|dbj|BAB08235.1 contains sim... 121 5e-28
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 84 9e-17
TC92156 similar to GP|15724296|gb|AAL06541.1 AT5g61660/k11j9_180... 83 2e-16
BF004325 weakly similar to GP|11994465|dbj contains similarity t... 82 3e-16
TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 82 3e-16
TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy cha... 79 3e-15
TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 75 4e-14
TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 75 5e-14
TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 75 5e-14
TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 75 5e-14
TC76705 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 71 8e-13
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 69 4e-12
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 66 2e-11
TC77148 ENOD20 64 1e-10
TC82245 similar to GP|16648720|gb|AAL25552.1 AT4g21620/F17L22_80... 64 2e-10
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 62 6e-10
BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 61 8e-10
TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protei... 61 8e-10
>TC77568 weakly similar to GP|9857516|gb|AAG00871.1| Unknown protein
{Arabidopsis thaliana}, partial (63%)
Length = 1206
Score = 286 bits (731), Expect = 2e-77
Identities = 132/293 (45%), Positives = 191/293 (65%), Gaps = 1/293 (0%)
Frame = +1
Query: 164 KNTCQFKTLSCPAECQKRKPKNNKKAKGCFIDCSSK-CEATCKFRKSKCDGYGALCYDPR 222
K+ C K L CPA+C R P N ++ K C++DC S C+ CK RK+ C+G G+ C DPR
Sbjct: 55 KSRCFLKKLPCPAQCPSRSPANPRE-KVCYLDCDSPMCKTQCKSRKANCNGRGSACLDPR 231
Query: 223 FVGGDGVMFYFHGAKGGNFAIASDDEFQINAHFIGTRPEGRTRDYTWVQALAVMFDTHTL 282
FVG DG++FYFHG + +F++ SD QINA FIG RP+GR RDYTW+QAL V+FD+H
Sbjct: 232 FVGADGIVFYFHGRRNEHFSLVSDTNLQINARFIGLRPQGRPRDYTWIQALGVLFDSHNF 411
Query: 283 VIAANRVSKWDDNVDSLIVKWDGEEITVPIDGETEWRANGDEREVVVERTEETNNVRVTV 342
I A S WDD VD L + +DG+++ +P + W++ +E ++ VERT N V VT+
Sbjct: 412 SIEATPSSIWDDEVDHLKLSYDGKDLVIPEGHLSTWQS--EENQLRVERTSNENGVMVTI 585
Query: 343 SGLLEMDIRVRPIGEKENKVHNYQLPSGDAFAHLETQFRFKKHTDSIEGILGQTYRPSYV 402
+ E+ + V P+ ++++++HNYQ+P D FAHL+ QF+F + +EG+LG+TY+P +
Sbjct: 586 PEVAEISVNVVPVTKEDSRIHNYQIPDDDCFAHLDVQFKFYGLSSKVEGVLGRTYQPDFQ 765
Query: 403 SPVKRGVPMPMMGGEDKYQTPSLFSTTCKHCRFQRPSGSATSEGLMLNIDTMN 455
+P K GV MP++GGEDKY+T SL S C C F S EG+ + M+
Sbjct: 766 NPAKPGVAMPVVGGEDKYRTKSLVSADCLACIFSSAKDS-VQEGMEIEYGMMD 921
>TC83922 similar to GP|15809990|gb|AAL06922.1 AT5g60530/muf9_180
{Arabidopsis thaliana}, partial (21%)
Length = 619
Score = 242 bits (617), Expect(2) = 9e-68
Identities = 112/160 (70%), Positives = 123/160 (76%)
Frame = +1
Query: 82 GNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKND 141
G GNGNGNGN NGNG GNGN NGN+ GNGKGN + GK G+ K + +
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGNN-----GNGKGNDKEEGKEKGKEKEKAPKKKKAPKE 264
Query: 142 EASEYEVLSALPSGQERAFCKAKNTCQFKTLSCPAECQKRKPKNNKKAKGCFIDCSSKCE 201
E+S Y+ L LPSGQERAFCKA NTC F TL CPAEC+ RKPK NKK KGCFIDCSSKCE
Sbjct: 265 ESSNYQQLQTLPSGQERAFCKANNTCHFATLVCPAECKTRKPKKNKKDKGCFIDCSSKCE 444
Query: 202 ATCKFRKSKCDGYGALCYDPRFVGGDGVMFYFHGAKGGNF 241
ATCKFR++ CDGYG+LCYDPRFVGGDGVMFYFHGAKGGNF
Sbjct: 445 ATCKFRRANCDGYGSLCYDPRFVGGDGVMFYFHGAKGGNF 564
Score = 62.8 bits (151), Expect = 3e-10
Identities = 34/54 (62%), Positives = 34/54 (62%)
Frame = +1
Query: 1 MEVVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 54
ME IAQ G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 82 MEAAIAQ------GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 62.0 bits (149), Expect = 5e-10
Identities = 30/53 (56%), Positives = 33/53 (61%)
Frame = +1
Query: 4 VIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 56
++ + E G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 67 ILLISMEAAIAQGQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/41 (70%), Positives = 29/41 (70%)
Frame = +1
Query: 20 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 60
G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/41 (70%), Positives = 29/41 (70%)
Frame = +1
Query: 30 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 70
G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/41 (70%), Positives = 29/41 (70%)
Frame = +1
Query: 34 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 74
G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/41 (70%), Positives = 29/41 (70%)
Frame = +1
Query: 28 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 68
G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/41 (70%), Positives = 29/41 (70%)
Frame = +1
Query: 24 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 64
G GNGNGNGN NGNG GNGNGNGN NGNG GN G G
Sbjct: 103 GQGNGNGNGN-NGNGKGNGNGNGN-NGNGKGNDKEEGKEKG 219
Score = 33.5 bits (75), Expect(2) = 9e-68
Identities = 14/14 (100%), Positives = 14/14 (100%)
Frame = +3
Query: 245 SDDEFQINAHFIGT 258
SDDEFQINAHFIGT
Sbjct: 576 SDDEFQINAHFIGT 617
>TC77569 weakly similar to GP|9757754|dbj|BAB08235.1 contains similarity to
root cap protein~gene_id:MUF9.15 {Arabidopsis thaliana},
partial (26%)
Length = 835
Score = 121 bits (304), Expect = 5e-28
Identities = 52/90 (57%), Positives = 65/90 (71%)
Frame = +3
Query: 207 RKSKCDGYGALCYDPRFVGGDGVMFYFHGAKGGNFAIASDDEFQINAHFIGTRPEGRTRD 266
RK+ C+G G+ C DPRFVG DG++FYFHG + +F++ SD QINA FIG RP+GR RD
Sbjct: 564 RKANCNGRGSACLDPRFVGADGIVFYFHGRRNEHFSLVSDTNLQINARFIGLRPQGRPRD 743
Query: 267 YTWVQALAVMFDTHTLVIAANRVSKWDDNV 296
YTW+QAL V+FD+H I A S WDD V
Sbjct: 744 YTWIQALGVLFDSHNFSIEATPSSIWDDEV 833
Score = 38.1 bits (87), Expect = 0.007
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = +3
Query: 164 KNTCQFKTLSCPAECQKRKPKNNKKAKGCFIDCSS-KCEATCK 205
K+ C K L CPA+C R P N ++ K ++DC S C+ CK
Sbjct: 111 KSRCFLKKLPCPAQCPSRSPANPRE-KVRYLDCDSPMCKTQCK 236
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 84.3 bits (207), Expect = 9e-17
Identities = 36/81 (44%), Positives = 46/81 (56%)
Frame = +1
Query: 40 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 99
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 100 GNSNGNSNGNGEGNGKGNGEG 120
G ++G + G G G G+G G G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 83.6 bits (205), Expect = 1e-16
Identities = 36/81 (44%), Positives = 44/81 (53%)
Frame = +1
Query: 18 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 77
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 78 GNGNGNGNGNGNGNGNGNGNG 98
G +G G G G G G G+G
Sbjct: 181GGAHGGAYGGGIGGGEGGGHG 243
Score = 83.6 bits (205), Expect = 1e-16
Identities = 37/84 (44%), Positives = 45/84 (53%)
Frame = +1
Query: 44 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSN 103
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 104 GNSNGNGEGNGKGNGEGNGKGNGE 127
G ++G G G G GEG G G E
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHGGYE 252
Score = 83.6 bits (205), Expect = 1e-16
Identities = 37/84 (44%), Positives = 45/84 (53%)
Frame = +1
Query: 36 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 95
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 96 GNGNGNSNGNSNGNGEGNGKGNGE 119
G +G + G G GEG G G E
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHGGYE 252
Score = 83.6 bits (205), Expect = 1e-16
Identities = 36/81 (44%), Positives = 44/81 (53%)
Frame = +1
Query: 20 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 79
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 80 GNGNGNGNGNGNGNGNGNGNG 100
G +G G G G G G G+G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 83.6 bits (205), Expect = 1e-16
Identities = 36/81 (44%), Positives = 44/81 (53%)
Frame = +1
Query: 14 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 73
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 74 GNGNGNGNGNGNGNGNGNGNG 94
G +G G G G G G G+G
Sbjct: 181GGAHGGAYGGGIGGGEGGGHG 243
Score = 83.6 bits (205), Expect = 1e-16
Identities = 36/81 (44%), Positives = 44/81 (53%)
Frame = +1
Query: 16 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 75
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 76 GNGNGNGNGNGNGNGNGNGNG 96
G +G G G G G G G+G
Sbjct: 181GGAHGGAYGGGIGGGEGGGHG 243
Score = 83.2 bits (204), Expect = 2e-16
Identities = 36/81 (44%), Positives = 44/81 (53%)
Frame = +1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 72 GNGNGNGNGNGNGNGNGNGNG 92
G +G G G G G G G+G
Sbjct: 181GGAHGGAYGGGIGGGEGGGHG 243
Score = 82.0 bits (201), Expect = 4e-16
Identities = 35/80 (43%), Positives = 44/80 (54%)
Frame = +1
Query: 9 QEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 68
++ G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G G
Sbjct: 4 RDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGGG 183
Query: 69 NGNGNGNGNGNGNGNGNGNG 88
+G G G G G G G+G
Sbjct: 184GAHGGAYGGGIGGGEGGGHG 243
Score = 81.3 bits (199), Expect = 7e-16
Identities = 35/80 (43%), Positives = 43/80 (53%)
Frame = +1
Query: 22 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 81
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 82 GNGNGNGNGNGNGNGNGNGN 101
G +G G G G G G G+
Sbjct: 181 GGAHGGAYGGGIGGGEGGGH 240
Score = 81.3 bits (199), Expect = 7e-16
Identities = 35/81 (43%), Positives = 43/81 (52%)
Frame = +1
Query: 24 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 83
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 84 GNGNGNGNGNGNGNGNGNSNG 104
G +G G G G G G +G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 79.7 bits (195), Expect = 2e-15
Identities = 35/84 (41%), Positives = 43/84 (50%)
Frame = +1
Query: 28 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 87
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 88 GNGNGNGNGNGNGNSNGNSNGNGE 111
G +G G G G G +G E
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHGGYE 252
Score = 79.0 bits (193), Expect = 4e-15
Identities = 34/80 (42%), Positives = 42/80 (52%)
Frame = +1
Query: 26 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 85
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 86 GNGNGNGNGNGNGNGNSNGN 105
G +G G G G G G+
Sbjct: 181 GGAHGGAYGGGIGGGEGGGH 240
Score = 78.6 bits (192), Expect = 5e-15
Identities = 34/81 (41%), Positives = 41/81 (49%)
Frame = +1
Query: 48 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSN 107
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+ G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 108 GNGEGNGKGNGEGNGKGNGEG 128
G G G G G G+G G G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 78.6 bits (192), Expect = 5e-15
Identities = 34/81 (41%), Positives = 41/81 (49%)
Frame = +1
Query: 32 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 91
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 92 GNGNGNGNGNSNGNSNGNGEG 112
G +G G G G G G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 78.6 bits (192), Expect = 5e-15
Identities = 35/81 (43%), Positives = 41/81 (50%)
Frame = +1
Query: 52 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGE 111
G +G G G G G+G G G G G +G G G+G G G G G G+ G G G+ G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 112 GNGKGNGEGNGKGNGEGNGKG 132
G G G G G GEG G G
Sbjct: 181 GGAHGGAYGGGIGGGEGGGHG 243
Score = 74.7 bits (182), Expect = 7e-14
Identities = 33/79 (41%), Positives = 40/79 (49%)
Frame = +1
Query: 56 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGK 115
G +G G G G G+G G G G G +G G G+G G G G G G+ G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 116 GNGEGNGKGNGEGNGKGNG 134
G G G G G G+G G
Sbjct: 181 GGAHGGAYGGGIGGGEGGG 237
Score = 69.3 bits (168), Expect = 3e-12
Identities = 31/76 (40%), Positives = 38/76 (49%)
Frame = +1
Query: 60 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGE 119
G +G G G G G+G G G G G +G G G+G G G G G + G G G+G G G
Sbjct: 1 GRDHGGGYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGGGGGGGSGGGGGG 180
Query: 120 GNGKGNGEGNGKGNGD 135
G G G G G G+
Sbjct: 181 GGAHGGAYGGGIGGGE 228
>TC92156 similar to GP|15724296|gb|AAL06541.1 AT5g61660/k11j9_180
{Arabidopsis thaliana}, partial (67%)
Length = 650
Score = 83.2 bits (204), Expect = 2e-16
Identities = 43/124 (34%), Positives = 66/124 (52%), Gaps = 1/124 (0%)
Frame = +2
Query: 3 VVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG-N 61
++ + A G +G +G+G+G G G+ G+G G G+G+G+ +
Sbjct: 113 LITSMAAASSGGRKLDMASGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPS 292
Query: 62 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGN 121
G G G G G G G+G+G+G+G G G+G +G G G+G+GNS G +G G N + E
Sbjct: 293 GFGRGYGYGFGTGSGSGSGSGYGYGSGGAHGGGYGSGSGNSGGGGHGGGRTNNRSPSESK 472
Query: 122 GKGN 125
GK N
Sbjct: 473 GKTN 484
Score = 78.6 bits (192), Expect = 5e-15
Identities = 41/117 (35%), Positives = 63/117 (53%)
Frame = +2
Query: 13 SGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 72
S G +G +G+G+G G G+ G+G G G+G+G+ + +G G G G
Sbjct: 137 SSGGRKLDMASGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGH-SPSGFGRGYG 313
Query: 73 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGN 129
G G G+G+G+G+G G G+G +G G G+ +GNS G G G G+ N + G+ N
Sbjct: 314 YGFGTGSGSGSGSGYGYGSGGAHGGGYGSGSGNSGGGGHGGGRTNNRSPSESKGKTN 484
Score = 76.3 bits (186), Expect = 2e-14
Identities = 39/106 (36%), Positives = 58/106 (53%), Gaps = 1/106 (0%)
Frame = +2
Query: 29 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGN 87
+G +G+G+G G G+ G+G G G+G+G+ +G G G G G G G+G+G+
Sbjct: 167 SGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGS 346
Query: 88 GNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGN 133
G+G G G+G +G G+ +GN G G G G N + E GK N
Sbjct: 347 GSGYGYGSGGAHGGGYGSGSGNSGGGGHGGGRTNNRSPSESKGKTN 484
Score = 75.5 bits (184), Expect = 4e-14
Identities = 40/108 (37%), Positives = 60/108 (55%), Gaps = 3/108 (2%)
Frame = +2
Query: 33 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGN 91
+G +G+G+G G G+ G+G G G+G+G+ +G G G G G G G+G+G+
Sbjct: 167 SGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGS 346
Query: 92 GNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGE--GNGKGNGDQP 137
G+G G G+G ++G G+G GN G G G G+ N KG +QP
Sbjct: 347 GSGYGYGSGGAHGGGYGSGSGNSGGGGHGGGRTNNRSPSESKGKTNQP 490
Score = 73.6 bits (179), Expect = 2e-13
Identities = 37/107 (34%), Positives = 59/107 (54%), Gaps = 1/107 (0%)
Frame = +2
Query: 37 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGN 95
+G +G+G+G G G+ G+G G G+G+G+ +G G G G G G G+G+G+
Sbjct: 167 SGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGS 346
Query: 96 GNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDE 142
G+G G +G ++G G G+G GN G G G G N + + K ++
Sbjct: 347 GSGYGYGSGGAHGGGYGSGSGNSGGGGHGGGRTNNRSPSESKGKTNQ 487
Score = 66.2 bits (160), Expect = 2e-11
Identities = 38/102 (37%), Positives = 53/102 (51%), Gaps = 1/102 (0%)
Frame = +2
Query: 45 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNSN 103
+G +G+G+G G G+ G+G G G+G+G+ +G G G G G G G+G+ +
Sbjct: 167 SGGSPTGASGSGHGPNWDYNWGWGSAPGSGWGYGSGSGHSPSGFGRGYGYGFGTGSGSGS 346
Query: 104 GNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEASE 145
G+ G G G G G G+G GN G G G G T N SE
Sbjct: 347 GSGYGYGSGGAHGGGYGSGSGNSGGGGHGGG--RTNNRSPSE 466
>BF004325 weakly similar to GP|11994465|dbj contains similarity to late
embryogenesis abundant protein~gene_id:MLD14.16
{Arabidopsis thaliana}, partial (8%)
Length = 357
Score = 82.4 bits (202), Expect = 3e-16
Identities = 39/110 (35%), Positives = 62/110 (55%)
Frame = +1
Query: 268 TWVQALAVMFDTHTLVIAANRVSKWDDNVDSLIVKWDGEEITVPIDGETEWRANGDEREV 327
TWVQ++ ++ D H L + A + + WDD+VD L + +DGE IT+ +W ++G V
Sbjct: 1 TWVQSIVILLDNHQLFLGAQKTATWDDSVDRLAISFDGEPITLHESEGAKWESSG----V 168
Query: 328 VVERTEETNNVRVTVSGLLEMDIRVRPIGEKENKVHNYQLPSGDAFAHLE 377
R TNN+ V G ++ + PI E+ ++VHNY + D AHL+
Sbjct: 169 SFGRETSTNNIIAEVEGNCKITAKGVPITEEHSRVHNYGITKDDCCAHLD 318
>TC76707 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (87%)
Length = 780
Score = 82.4 bits (202), Expect = 3e-16
Identities = 43/115 (37%), Positives = 52/115 (44%)
Frame = +1
Query: 5 IAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 64
+ A+ G G G +G G NG G +G G N G G +G G N G G +G G
Sbjct: 139 VNDAKYGGYGGGYNHGGGGYNGGGYNHGGGGYNNGGGGYNHGGGGYNNGGGGYNHGGGGY 318
Query: 65 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGE 119
NG G +G G N G G +G G NG G G+G G NG G G G E
Sbjct: 319 NGGGYNHGGGGYNHGGGGYNHGGGGYNGGGGHGGHG--GGGYNGGGGHGGHGAAE 477
Score = 81.6 bits (200), Expect = 6e-16
Identities = 49/159 (30%), Positives = 59/159 (36%), Gaps = 25/159 (15%)
Frame = +1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G G +G G N G G +G G N G G +G G NG G +G G N G G +
Sbjct: 202 GGGYNHGGGGYNNGGGGYNHGGGGYNNGGGGYNHGGGGYNGGGYNHGGGGYNHGGGGYNH 381
Query: 72 GNGNGNGNGNGNGNGNGNGNGNGNGNGNG-------------------------NSNGNS 106
G G NG G G+G G NG G G+G N G S
Sbjct: 382 GGGGYNGGGGHGGHGGGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGYNHGGGS 561
Query: 107 NGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEASE 145
+G G+ G G G G G G G E +E
Sbjct: 562 YNHGGGSYHHGGGGYNHGGGGHGGHGGGGHGGHGAEQTE 678
Score = 81.3 bits (199), Expect = 7e-16
Identities = 48/150 (32%), Positives = 61/150 (40%), Gaps = 21/150 (14%)
Frame = +1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G G +G G N G G +G G NG G +G G N G G +G G NG G G+
Sbjct: 244 GGGYNHGGGGYNNGGGGYNHGGGGYNGGGYNHGGGGYNHGGGGYNHGGGGYNGGGGHGGH 423
Query: 72 GNGNGNGNGNGNGNGNGN------------------GNGNGNGNGNGNSNGNS---NGNG 110
G G NG G G+G G G +G G+ N G S G G
Sbjct: 424 GGGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGYNHGGGSYNHGGGSYHHGGGG 603
Query: 111 EGNGKGNGEGNGKGNGEGNGKGNGDQPTKN 140
+G G G+G G G+G + T+N
Sbjct: 604 YNHGGGGHGGHGGGGHGGHGAEQTEDKTQN 693
Score = 38.1 bits (87), Expect = 0.007
Identities = 19/63 (30%), Positives = 28/63 (44%)
Frame = +1
Query: 2 EVVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 61
E V Q +EK + + G N G +G G+ + G G +G G G+G G
Sbjct: 475 ESVAVQTEEKTNEVNDAKYGGGYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGGHG 654
Query: 62 GNG 64
G+G
Sbjct: 655 GHG 663
>TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy chain
precursor (Fib-H) (H-fibroin). [Silk moth] {Bombyx
mori}, partial (2%)
Length = 782
Score = 79.3 bits (194), Expect = 3e-15
Identities = 47/127 (37%), Positives = 77/127 (60%), Gaps = 5/127 (3%)
Frame = +3
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN-GNGNGNGNGNGNGNG 70
G G G G G G G G G G+G G G+G+G+G+ + +G+G+G+G+ G G+ G+ G+ G
Sbjct: 171 GIGLGAGLGLGLGGGGGAGSGAGAGSGSGSGSASTSGSGSGSGSSGAGSEAGSHAGSRAG 350
Query: 71 NGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNSNGNSNGNGEGNGK---GNGEGNGKGNG 126
+G G+G G+ G+ G+ G+G +G G+ G+ G+ G+ G+G G+ G+ G+
Sbjct: 351 SGAGSGAGSEAGSYAGSHAGSGSSGAGSEAGSEAGSYAGSHAGSGSSRAGSEAGSYAGSH 530
Query: 127 EGNGKGN 133
G+G GN
Sbjct: 531 AGSGSGN 551
Score = 73.2 bits (178), Expect = 2e-13
Identities = 42/121 (34%), Positives = 71/121 (57%)
Frame = +3
Query: 24 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 83
G G G G G G G G G G G+G G G+G+G+G+ + +G+G+G+G+ +G G+ G+
Sbjct: 165 GVGIGLGAGLGLGLGGGGGAGSGAGAGSGSGSGSASTSGSGSGSGS---SGAGSEAGSHA 335
Query: 84 GNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEA 143
G+ G+G G+G G+ G+ G+ G+G +G G+ G+ G+ G+ G+G ++
Sbjct: 336 GSRAGSGAGSGAGSEAGSYAGSHAGSG-SSGAGSEAGSEAGSYAGSHAGSGSSRAGSEAG 512
Query: 144 S 144
S
Sbjct: 513 S 515
Score = 69.3 bits (168), Expect = 3e-12
Identities = 36/98 (36%), Positives = 59/98 (59%)
Frame = +3
Query: 48 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSN 107
G G G G G G G G G G G+G G G+G+G+G+ + +G+G+G+G+ +G G+ G+
Sbjct: 165 GVGIGLGAGLGLGLGGGGGAGSGAGAGSGSGSGSASTSGSGSGSGS---SGAGSEAGSHA 335
Query: 108 GNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEASE 145
G+ G+G G+G G+ G+ G+ G+G ++ SE
Sbjct: 336 GSRAGSGAGSGAGSEAGSYAGSHAGSGSSGAGSEAGSE 449
Score = 50.4 bits (119), Expect = 1e-06
Identities = 33/96 (34%), Positives = 58/96 (60%), Gaps = 2/96 (2%)
Frame = +3
Query: 8 AQEKGSGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 66
A GSG+G+G +G G+ G+ G+ G+G G+G G+ G+ G+ G+G+ +G G+ G
Sbjct: 267 ASTSGSGSGSGSSGAGSEAGSHAGSRAGSGAGSGAGSEAGSYAGSHAGSGS-SGAGSEAG 443
Query: 67 NGNGNGNGNGNGNGNGN-GNGNGNGNGNGNGNGNGN 101
+ G+ G+ G+G+ G+ G+ G+ G+G+GN
Sbjct: 444 SEAGSYAGSHAGSGSSRAGSEAGSYAGSHAGSGSGN 551
>TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (94%)
Length = 1034
Score = 75.5 bits (184), Expect = 4e-14
Identities = 46/149 (30%), Positives = 62/149 (40%), Gaps = 20/149 (13%)
Frame = +1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G NG G +G G NG G + G G +G G NG G G+G NG G G+G NG
Sbjct: 220 GGYNGGGYNHGGGYHNGGGGYHNGGGGYNHGGGGYNGGGGHGGHGGYNGGG-GHGGYNGG 396
Query: 72 GNGNGNGN--------------------GNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGE 111
G G+G G G+ N G+ N G +G + + G G
Sbjct: 397 GGHGGHGAAESVAVQTEEKTNEVNDAKYGGGSYNHGGSYNHGGGSYNHGGGSYHHGGGGY 576
Query: 112 GNGKGNGEGNGKGNGEGNGKGNGDQPTKN 140
+G G G+G G G+G + T+N
Sbjct: 577 NHGGGGHGGHGGGGHGGHGAEQTEDKTQN 663
Score = 73.2 bits (178), Expect = 2e-13
Identities = 49/167 (29%), Positives = 62/167 (36%), Gaps = 25/167 (14%)
Frame = +1
Query: 4 VIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 63
V+ + E G +G N G NG G +G G NG G + G G +G G N
Sbjct: 148 VVEKTNEVNDAKYGGYNHGGYNHGGGYNGGGYNHGGGYHNGGGGYHNGGGGYNHGGGGYN 327
Query: 64 GNGNGNGNGNGNGNG-----NGNGNGNGNG--------------------NGNGNGNGNG 98
G G G+G NG G NG G G+G G G+ N G
Sbjct: 328 GGGGHGGHGGYNGGGGHGGYNGGGGHGGHGAAESVAVQTEEKTNEVNDAKYGGGSYNHGG 507
Query: 99 NGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEASE 145
+ N G S +G G+ G G G G G G G E +E
Sbjct: 508 SYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGGHGGGGHGGHGAEQTE 648
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/71 (30%), Positives = 32/71 (44%)
Frame = +1
Query: 2 EVVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 61
E V Q +EK + + G +G +G G+ N G +G G N G G+G
Sbjct: 424 ESVAVQTEEKTNEVNDAKYGGGSYNHGGSYNHGGGSYNHGGGSYHHGGGGYNHGGGGHGG 603
Query: 62 GNGNGNGNGNG 72
G G+G G+G
Sbjct: 604 HGGGGHG-GHG 633
>TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 912
Score = 75.1 bits (183), Expect = 5e-14
Identities = 36/84 (42%), Positives = 40/84 (46%)
Frame = +1
Query: 3 VVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 62
+ + QAQ +GSG G G G G G G G G G G G G G G G G G G G G G
Sbjct: 472 ITVNQAQSRGSGGGGGGGRG-GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGG 648
Query: 63 NGNGNGNGNGNGNGNGNGNGNGNG 86
G G G G G G G G+G
Sbjct: 649 GGYSRGGGGGGYGGGGYSRGGGDG 720
>TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 1054
Score = 75.1 bits (183), Expect = 5e-14
Identities = 36/84 (42%), Positives = 40/84 (46%)
Frame = +1
Query: 3 VVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 62
+ + QAQ +GSG G G G G G G G G G G G G G G G G G G G G G
Sbjct: 568 ITVNQAQSRGSGGGGGGGRG-GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGG 744
Query: 63 NGNGNGNGNGNGNGNGNGNGNGNG 86
G G G G G G G G+G
Sbjct: 745 GGYSRGGGGGGYGGGGYSRGGGDG 816
>TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 754
Score = 75.1 bits (183), Expect = 5e-14
Identities = 36/84 (42%), Positives = 40/84 (46%)
Frame = +2
Query: 3 VVIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 62
+ + QAQ +GSG G G G G G G G G G G G G G G G G G G G G G
Sbjct: 314 ITVNQAQSRGSGGGGGGGRG-GGGYGGGGGGYGGERRGYGGGGGYGGGGGGGYGERRGGG 490
Query: 63 NGNGNGNGNGNGNGNGNGNGNGNG 86
G G G G G G G G+G
Sbjct: 491 GGYSRGGGGGGYGGGGYSRGGGDG 562
>TC76705 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (80%)
Length = 854
Score = 71.2 bits (173), Expect = 8e-13
Identities = 48/152 (31%), Positives = 61/152 (39%), Gaps = 15/152 (9%)
Frame = +2
Query: 4 VIAQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 63
V+ + E G +G N G NG G +G G NG G NG G G NG G N
Sbjct: 158 VVEKTNEVNDAKYGGYNHGGYNHGGGYNGGGYNHGGGYHNGGGYHNG-GGGYHNGGGGYN 334
Query: 64 GNGNGNGNGNGNGNGNGNGNGNGNG------------NGNGNGNGNGNGNSNGNSNGNGE 111
G G G+G N G G G N NG G G+ N G S +G
Sbjct: 335 GGGGHGGHGGYNRGGGHGGRGAAESVAVQTEEKTNEVNDARNGGGGGSFNKGGASYNHGR 514
Query: 112 G---NGKGNGEGNGKGNGEGNGKGNGDQPTKN 140
G +G G G G+G G +G + T++
Sbjct: 515 GSYHHG*GGYNHGGGGHGGGGHGSHGAEQTED 610
Score = 47.0 bits (110), Expect = 2e-05
Identities = 34/97 (35%), Positives = 41/97 (42%), Gaps = 6/97 (6%)
Frame = +2
Query: 72 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNS---NGNGEGNGKGNGEGNG---KGN 125
G +G N G NG G +G G NG G NG NG G NG G G+G +G
Sbjct: 200 GYNHGGYNHGGGYNGGGYNHGGGYHNGGGYHNGGGGYHNGGGGYNGGGGHGGHGGYNRGG 379
Query: 126 GEGNGKGNGDQPTKNDEASEYEVLSALPSGQERAFCK 162
G G G+G + E EV A G +F K
Sbjct: 380 GHG-GRGAAESVAVQTEEKTNEVNDARNGGGGGSFNK 487
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 68.9 bits (167), Expect = 4e-12
Identities = 42/145 (28%), Positives = 47/145 (31%), Gaps = 23/145 (15%)
Frame = +3
Query: 14 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 73
G G G G G G G G G G G G G G G+G G G G + G
Sbjct: 111 GGGGGGRCGGGGGGGGGRGGAEAAGPGGGGGGGGDALGRRRLAKGSGAGGGGGARSRGGG 290
Query: 74 GNGNGNGNGNGNG-----------------------NGNGNGNGNGNGNGNSNGNSNGNG 110
G G G G G G G G G G +
Sbjct: 291 GPGGWGRAGQGTGLWTPVCRPWGVS*PWPGHHGDALTGGARVAGGVGGAGGPEGGARAGA 470
Query: 111 EGNGKGNGEGNGKGNGEGNGKGNGD 135
G G+G G G G G G G+G GD
Sbjct: 471 AGGGRGGGRGLGLWRGSGGGRGRGD 545
Score = 61.6 bits (148), Expect = 6e-10
Identities = 41/154 (26%), Positives = 48/154 (30%), Gaps = 31/154 (20%)
Frame = +3
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN--------GNGNGNGNGNGNGNGNGN 63
G G G G G G G G G G G G G G G+G G G G + G
Sbjct: 111 GGGGGGRCGGGGGGGGGRGGAEAAGPGGGGGGGGDALGRRRLAKGSGAGGGGGARSRGGG 290
Query: 64 GNGNGNGNGNGNG-----------------------NGNGNGNGNGNGNGNGNGNGNGNG 100
G G G G G G G G G G
Sbjct: 291 GPGGWGRAGQGTGLWTPVCRPWGVS*PWPGHHGDALTGGARVAGGVGGAGGPEGGARAGA 470
Query: 101 NSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNG 134
G G G G +G+G G G+G+ G + G
Sbjct: 471 AGGGRGGGRGLGLWRGSGGGRGRGDWRGGARWAG 572
Score = 52.8 bits (125), Expect = 3e-07
Identities = 30/97 (30%), Positives = 34/97 (34%)
Frame = +3
Query: 26 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 85
G G G G G+G G G G G G G G G G G G G G G
Sbjct: 39 GGARGAGGWRRGGEVGGSGWCVWGGGGGGGRCGGGGGGGGGRGGAEAAGPGGGGGGGGDA 218
Query: 86 GNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNG 122
G+G G G + G G G G+G G
Sbjct: 219 LGRRRLAKGSGAGGGGGARSRGGGGPGGWGRAGQGTG 329
Score = 49.7 bits (117), Expect = 2e-06
Identities = 41/145 (28%), Positives = 48/145 (32%), Gaps = 22/145 (15%)
Frame = +1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGN-----------GNGN-----GNGNGNGNGNGN 55
G+G G G G G G G G G G+G G G G G+G
Sbjct: 256 GAGGGPGPGAAGGRVGGGGRGRGRGSGPRCAALGEFPDLGPGTTATP*RGARGWPAGSGA 435
Query: 56 GNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNG 114
G G G G G G + G G G G G G G G + G G G
Sbjct: 436 LGGRRGARGRAPRGGGGGGGAASDCGGALGGDGGGGTGGGGRGGRGYVATRTRGTSAGRG 615
Query: 115 -----KGNGEGNGKGNGEGNGKGNG 134
G + G G G G+G G
Sbjct: 616 CPPIFLARGTRSHSGGG-GGGRGGG 687
Score = 47.8 bits (112), Expect = 9e-06
Identities = 30/101 (29%), Positives = 33/101 (31%)
Frame = +1
Query: 22 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 81
G G G G G G G G G G G G G G G G
Sbjct: 37 GGGRG-GRGAGGGGGRWGAQAGACGGVGAGGAAAGGVGGGAGAEGGRRRRARGGGAVAGG 213
Query: 82 GNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNG 122
G G G G G G G + G G G+G G G+G
Sbjct: 214 MPSAAGGLLRGVGPGAGGGPGPGAAGGRVGGGGRGRGRGSG 336
Score = 44.7 bits (104), Expect = 8e-05
Identities = 29/101 (28%), Positives = 31/101 (29%)
Frame = +1
Query: 30 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 89
G G G G G G G G G G G G G G G G
Sbjct: 37 GGGRG-GRGAGGGGGRWGAQAGACGGVGAGGAAAGGVGGGAGAEGGRRRRARGGGAVAGG 213
Query: 90 GNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNG 130
G G G G G G G G G+G G G+G
Sbjct: 214 MPSAAGGLLRGVGPGAGGGPGPGAAGGRVGGGGRGRGRGSG 336
Score = 41.6 bits (96), Expect = 6e-04
Identities = 34/130 (26%), Positives = 41/130 (31%), Gaps = 7/130 (5%)
Frame = +1
Query: 6 AQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 65
A+ GSG G G G G G + G G+G G G G G G G
Sbjct: 406 ARGWPAGSGALGGRRGARGRAPRGGGGGGGAASDCGGALGGDGGGGTGGGGRG-GRGYVA 582
Query: 66 GNGNGNGNGNG-------NGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNG 118
G G G G + +G G G G G G + G G G
Sbjct: 583 TRTRGTSAGRGCPPIFLARGTRSHSGGGGGGRGGGRRGVAAGCARGMLCGARAQGAPGGV 762
Query: 119 EGNGKGNGEG 128
G G +G
Sbjct: 763 ALGGAGRADG 792
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 66.2 bits (160), Expect = 2e-11
Identities = 50/158 (31%), Positives = 61/158 (37%), Gaps = 34/158 (21%)
Frame = -1
Query: 9 QEKGSGNGNG-------------NGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNG-- 52
Q KG G G+ G G G+ G G G G+G G G G G
Sbjct: 594 Q*KGGGGGDAYEIPKTGGIGPSYTGGGGGDEYTGGGGEGYAYTGDGGGEVYTGGGGEG*A 415
Query: 53 -NGNGNGNGNGNG----------NGNGNGNGNGNGNGNGNGNGNGNGNG---NGNG---- 94
G+G G+G+G G +G G G G G G+ G G+G G G G
Sbjct: 414 YTGDGGGDGDGGG**GV*CGAGDDGGGGSTQGGGGGGGGSTQGGGDGGGGE*TGGGGECT 235
Query: 95 NGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKG 132
G G G+ G + G GE G G G G G G G G
Sbjct: 234 GGGGGGDKYGYTGGVGE*GGGGGEXGGGGGE*AGGGGG 121
Score = 44.7 bits (104), Expect = 8e-05
Identities = 30/104 (28%), Positives = 37/104 (34%), Gaps = 3/104 (2%)
Frame = -1
Query: 34 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG---NG 90
G G G+ G G G G G G G+G G G G G G
Sbjct: 585 GGGGGDAYEIPKTGGIGPSYTGGGGGDEYTGGGGEGYAYTGDGGGEVYTGGGGEG*AYTG 406
Query: 91 NGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNG 134
+G G+G+G G +G G G G G G G G+G
Sbjct: 405 DGGGDGDGGG**GV*CGAGDDGGGGSTQGGGGGGGGSTQGGGDG 274
Score = 30.0 bits (66), Expect = 1.9
Identities = 23/67 (34%), Positives = 28/67 (41%), Gaps = 11/67 (16%)
Frame = -1
Query: 80 GNGNGNGNGNGNGNGNG---NGNGNSNGNSNGNGEG---NGKGNGEGNGKGNGEG----- 128
G G G+ G G G G + + G GEG G G GE G GEG
Sbjct: 585 GGGGGDAYEIPKTGGIGPSYTGGGGGDEYTGGGGEGYAYTGDGGGEVYTGGGGEG*AYTG 406
Query: 129 NGKGNGD 135
+G G+GD
Sbjct: 405 DGGGDGD 385
>TC77148 ENOD20
Length = 1108
Score = 63.9 bits (154), Expect = 1e-10
Identities = 34/107 (31%), Positives = 54/107 (49%)
Frame = -1
Query: 18 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 77
G+G G G+ + + N G+G +G+ G G+ + +G+G+G G G G
Sbjct: 742 GDGAGEGDSSEGEDDGANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGV 563
Query: 78 GNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKG 124
G+G+G+G+G G+G+G G G G+ G G + GEG G G
Sbjct: 562 DRGDGDGDGDGEGDGDGEG-GEGSERLGG*GIGVDDRGVGGEGGGGG 425
Score = 63.9 bits (154), Expect = 1e-10
Identities = 38/120 (31%), Positives = 61/120 (50%), Gaps = 3/120 (2%)
Frame = -1
Query: 22 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 81
G+G G G+ + + N G+G +G+ G G+ + +G+G+G G G G
Sbjct: 742 GDGAGEGDSSEGEDDGANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGV 563
Query: 82 GNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNG---KGNGEGNGKGNGDQPT 138
G+G+G+G+G G+G+G G GEG+ + G G G +G G G G G G++ T
Sbjct: 562 DRGDGDGDGDGEGDGDGEG---------GEGSERLGG*GIGVDDRGVG-GEGGGGGEEST 413
Score = 63.9 bits (154), Expect = 1e-10
Identities = 35/110 (31%), Positives = 54/110 (48%)
Frame = -1
Query: 10 EKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 69
E G G G G+ + + N G+G +G+ G G+ + +G+G+G G G
Sbjct: 748 ELGDGAGEGDSSEGEDDGANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GI 569
Query: 70 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGE 119
G G+G+G+G+G G+G+G G G G+ G G + G G G GE
Sbjct: 568 GVDRGDGDGDGDGEGDGDGEG-GEGSERLGG*GIGVDDRGVGGEGGGGGE 422
Score = 57.4 bits (137), Expect = 1e-08
Identities = 30/91 (32%), Positives = 47/91 (50%)
Frame = -1
Query: 6 AQAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 65
A + G+G +G+ G G+ + +G+G+G G G G G+G+G+G+G G+G+
Sbjct: 694 ANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGVDRGDGDGDGDGEGDGD 515
Query: 66 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 96
G G G G+ G G G + G G G G
Sbjct: 514 GEG-GEGSERLGG*GIGVDDRGVGGEGGGGG 425
Score = 57.4 bits (137), Expect = 1e-08
Identities = 29/86 (33%), Positives = 46/86 (52%)
Frame = -1
Query: 9 QEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 68
+ +G G +G+ G G+ + +G+G+G G G G G+G+G+G+G G+G+G G G
Sbjct: 679 ESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGVDRGDGDGDGDGEGDGDGEG-G 503
Query: 69 NGNGNGNGNGNGNGNGNGNGNGNGNG 94
G+ G G G + G G G G
Sbjct: 502 EGSERLGG*GIGVDDRGVGGEGGGGG 425
Score = 57.0 bits (136), Expect = 1e-08
Identities = 29/86 (33%), Positives = 46/86 (52%)
Frame = -1
Query: 18 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 77
G+G +G+ G G+ + +G+G+G G G G G+G+G+G+G G+G+G G G G+
Sbjct: 670 GDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGVDRGDGDGDGDGEGDGDGEG-GEGS 494
Query: 78 GNGNGNGNGNGNGNGNGNGNGNGNSN 103
G G G + G G G G +
Sbjct: 493 ERLGG*GIGVDDRGVGGEGGGGGEES 416
Score = 55.8 bits (133), Expect = 3e-08
Identities = 30/98 (30%), Positives = 47/98 (47%)
Frame = -1
Query: 10 EKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 69
E N G+G +G+ G G+ + +G+G+G G G G G+G+G+G+G
Sbjct: 706 EDDGANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGVDRGDGDGDGDGE 527
Query: 70 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSN 107
G+G+G G G G+ G G G + G G G +
Sbjct: 526 GDGDGEG-GEGSERLGG*GIGVDDRGVGGEGGGGGEES 416
Score = 53.9 bits (128), Expect = 1e-07
Identities = 26/84 (30%), Positives = 44/84 (51%)
Frame = -1
Query: 62 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGN 121
G+G G G+ + + N G+G +G+ G G+ +S+G+ +G +G G G
Sbjct: 742 GDGAGEGDSSEGEDDGANEATESEGDGASDGDSEGEGDFDSDGDGDGEAGERFRG*GIGV 563
Query: 122 GKGNGEGNGKGNGDQPTKNDEASE 145
+G+G+G+G G GD + E SE
Sbjct: 562 DRGDGDGDGDGEGDGDGEGGEGSE 491
>TC82245 similar to GP|16648720|gb|AAL25552.1 AT4g21620/F17L22_80
{Arabidopsis thaliana}, partial (67%)
Length = 604
Score = 63.5 bits (153), Expect = 2e-10
Identities = 47/137 (34%), Positives = 54/137 (39%), Gaps = 7/137 (5%)
Frame = +3
Query: 75 NGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNG 134
N G G G+ G G G G G G N GNG GNG G G G+G G G NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGG-FNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Query: 135 -DQPTKNDEASEYEVLSALPSGQERAFCKAKNTCQFKTLSCPAECQKRKPKNNK------ 187
+PT CK K C K L+CPA+C ++ K
Sbjct: 381 IVRPT--------------------VVCKDKGPCYQKKLTCPAKCFSSFSRSGKGYGGGG 500
Query: 188 KAKGCFIDCSSKCEATC 204
GC IDC KC A C
Sbjct: 501 GGGGCTIDCKKKCSAYC 551
Score = 46.2 bits (108), Expect = 3e-05
Identities = 36/102 (35%), Positives = 41/102 (39%), Gaps = 16/102 (15%)
Frame = +3
Query: 47 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGN 105
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNGI 383
Query: 106 SNGNGEGNGKG---------------NGEGNGKGNGEGNGKG 132
KG + +GKG G G G G
Sbjct: 384 VRPTVVCKDKGPCYQKKLTCPAKCFSSFSRSGKGYGGGGGGG 509
Score = 45.4 bits (106), Expect = 4e-05
Identities = 28/64 (43%), Positives = 31/64 (47%)
Frame = +3
Query: 11 KGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 70
K N G G G+ G G G G G G N G GNG GNG G G G+G G NG
Sbjct: 192 KDKNNNQGGGGGSIPGVGGIPGGYFGPGGGF-NIPGFGNGFGNGVGGGYGSGYGGPNGGH 368
Query: 71 NGNG 74
+ NG
Sbjct: 369 SKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 21 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 78
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 27 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 84
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 29 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 86
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 25 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 82
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 33 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 90
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 35 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 92
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 37 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 94
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 39 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 96
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 41 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 98
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 43 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 100
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 31 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 88
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 23 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 80
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/59 (45%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 19 NGNGNGNGNGNGNGNGNGNGNGNGNG-NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 76
N G G G+ G G G G G G N G GNG GNG G G G+G G NG + NG
Sbjct: 204 NNQGGGGGSIPGVGGIPGGYFGPGGGFNIPGFGNGFGNGVGGGYGSGYGGPNGGHSKNG 380
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 61.6 bits (148), Expect = 6e-10
Identities = 36/105 (34%), Positives = 44/105 (41%)
Frame = -1
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G G G+G G G G+G+G G G G G+G G G G G G G
Sbjct: 342 GGGEGDGGGGDL*TYGGGGDGDGGGGDL*TYGGGGEGDGGGGDL*I*GGGGEGVGGGGDL 163
Query: 72 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKG 116
G G+G G G G G+G+G G G GEG+G G
Sbjct: 162 *TYGGGGDGEGGGGLL*TYGGGGDGDGGGGDLYTYGGGGEGDGGG 28
Score = 61.6 bits (148), Expect = 6e-10
Identities = 38/108 (35%), Positives = 46/108 (42%)
Frame = -1
Query: 28 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 87
G G G+G G G G+G+G G G G G+G G G G G G G
Sbjct: 342 GGGEGDGGGGDL*TYGGGGDGDGGGGDL*TYGGGGEGDGGGGDL*I*GGGGEGVGGGGDL 163
Query: 88 GNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGD 135
G G+G G G G G+G+G G G GEG+G G GD
Sbjct: 162 *TYGGGGDGEGGGGLL*TYGGGGDGDGGGGDLYTYGGGGEGDG-GGGD 22
Score = 56.2 bits (134), Expect = 3e-08
Identities = 35/103 (33%), Positives = 40/103 (37%)
Frame = -3
Query: 28 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 87
G G+G+G G G G G G G G G+G G G G G G G
Sbjct: 679 GGGDGDGGGGDL*I*GGGGEGEGGGGDL*PYGGGGDGEGGGGDL**YGGGGEAEGGGGDL 500
Query: 88 GNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGEGNG 130
G G G+G G G GEG G G G GEG+G
Sbjct: 499 *T*GGGGEGDGGGGDL*TYGGGGEGEGGGGDL*TYGGGGEGDG 371
Score = 55.8 bits (133), Expect = 3e-08
Identities = 34/103 (33%), Positives = 39/103 (37%)
Frame = -3
Query: 12 GSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 71
G G+G+G G G G G G G G G+G G G G G G G
Sbjct: 679 GGGDGDGGGGDL*I*GGGGEGEGGGGDL*PYGGGGDGEGGGGDL**YGGGGEAEGGGGDL 500
Query: 72 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNG 114
G G G+G G G G G G G G GEG+G
Sbjct: 499 *T*GGGGEGDGGGGDL*TYGGGGEGEGGGGDL*TYGGGGEGDG 371
Score = 50.4 bits (119), Expect = 1e-06
Identities = 33/92 (35%), Positives = 37/92 (39%)
Frame = -3
Query: 44 GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSN 103
G G+G+G G G G G G G G G+G G G G G G G
Sbjct: 679 GGGDGDGGGGDL*I*GGGGEGEGGGGDL*PYGGGGDGEGGGGDL**YGGGGEAEGGGGDL 500
Query: 104 GNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGD 135
G GEG+G G G GEG G G GD
Sbjct: 499 *T*GGGGEGDGGGGDL*TYGGGGEGEG-GGGD 407
>BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (19%)
Length = 1100
Score = 61.2 bits (147), Expect = 8e-10
Identities = 47/151 (31%), Positives = 54/151 (35%), Gaps = 15/151 (9%)
Frame = -1
Query: 7 QAQEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN--- 63
QA+E+ G G G G GNG G G G G G
Sbjct: 494 QAKERKRGRKRRKGENAKREKGGKERGKKERKKGAGNGIYEGEYKGGGGGGGKRKKKREG 315
Query: 64 GNGNGNGN-----GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNS-------NGNSNGNGE 111
G G G G G G GNG G G G G G G G G G+ G+
Sbjct: 314 GRGKGKDEII*EKGGGMGRGNG-GEGGGGGEGGGREWGGGGGDK*SLKRKEKEKDGGDKI 138
Query: 112 GNGKGNGEGNGKGNGEGNGKGNGDQPTKNDE 142
G GKG G G G + GNG G + K +E
Sbjct: 137 G*GKGRGGGEGGMDVMGNGGGRRGEGRKKEE 45
Score = 53.9 bits (128), Expect = 1e-07
Identities = 46/140 (32%), Positives = 53/140 (37%), Gaps = 10/140 (7%)
Frame = -1
Query: 5 IAQAQEKGSGNGNGNGN-----GNGNGNGN-----GNGNGNGNGNGNGNGNGNGNGNGNG 54
I + + KG G G G G G G G G G GNG G G G G G G
Sbjct: 380 IYEGEYKGGGGGGGKRKKKREGGRGKGKDEII*EKGGGMGRGNGG-EGGGGGEGGGREWG 204
Query: 55 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGEGNG 114
G G+ +G G+ G G G G G G + GNG G GEG
Sbjct: 203 GGGGDK*SLKRKEKEKDG-GDKIG*GKGRGGGEGGMDVMGNGGGR---------RGEGRK 54
Query: 115 KGNGEGNGKGNGEGNGKGNG 134
K E G+ EG GK G
Sbjct: 53 KEEREKRGE---EGEGKEGG 3
Score = 49.7 bits (117), Expect = 2e-06
Identities = 44/150 (29%), Positives = 52/150 (34%), Gaps = 23/150 (15%)
Frame = -3
Query: 9 QEKGSGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN----- 63
+E+G NGN G G G G G G+G +
Sbjct: 543 RERGRVNGNRGKGRGGTSERKEEGKEKEERGKRKEGKGGERKGEEGEEEGSGKWDI*RGV 364
Query: 64 -GNGNGNGNGNGNGNG-NGNGNG-NGNGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNG-- 118
G G G G G G G G N G G G+G G G G GEG G+G G
Sbjct: 363 *GGGRGGGKKEKKKRGWEGKGEG*NNIGEGRGDGKRKWGGGGGGRRRG-GEGMGRGGGG* 187
Query: 119 -----EGNGK--------GNGEGNGKGNGD 135
+G GK G EG G+G GD
Sbjct: 186 MIFKEKGKGKRWGG*DRVGEREG-GRGRGD 100
Score = 46.6 bits (109), Expect = 2e-05
Identities = 40/155 (25%), Positives = 51/155 (32%), Gaps = 23/155 (14%)
Frame = -3
Query: 5 IAQAQEKGSGNGNGNGNGNGNG--------------NGNGNGNGNGNGNGNGNGNGNGNG 50
+ Q +++G G G G G G G+G N G G NGN G
Sbjct: 672 LKQKKKRGGGGGRGGG-GRGDGIVVREVRVKRRKMDGANEERWGRERGRVNGNRGKGRGG 496
Query: 51 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN------GNGNGNGNGNGNSNG 104
G G G G G+G + G G G G G
Sbjct: 495 TSERKEEGKEKEERGKRKEGKGGERKGEEGEEEGSGKWDI*RGV*GGGRGGGKKEKKKRG 316
Query: 105 NSNGNGEG---NGKGNGEGNGKGNGEGNGKGNGDQ 136
G GEG G+G G+G K G G G+ G +
Sbjct: 315 -WEGKGEG*NNIGEGRGDGKRKWGGGGGGRRRGGE 214
Score = 33.9 bits (76), Expect = 0.13
Identities = 27/121 (22%), Positives = 36/121 (29%), Gaps = 8/121 (6%)
Frame = -3
Query: 27 NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNG 86
+G G G G G G G+G N G G NGN
Sbjct: 687 HGGGRLKQKKKRGGGGGRGGGGRGDGIVVREVRVKRRKMDGANEERWGRERGRVNGNRGK 508
Query: 87 NGNGNGNGNGNGNGNSNGNSNGNGEGNGKGNGEGNGKGNGE--------GNGKGNGDQPT 138
G G G+G + EG +G+G+ G G+G G +
Sbjct: 507 GRGGTSERKEEGKEKEERGKRKEGKGGERKGEEGEEEGSGKWDI*RGV*GGGRGGGKKEK 328
Query: 139 K 139
K
Sbjct: 327 K 325
>TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protein {Oryza
sativa}, partial (6%)
Length = 605
Score = 61.2 bits (147), Expect = 8e-10
Identities = 52/163 (31%), Positives = 58/163 (34%), Gaps = 35/163 (21%)
Frame = +1
Query: 7 QAQEKG-SGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGN 65
Q KG SGN NGNG G G+ G N G G G+ N + NG G
Sbjct: 25 QMHMKGKSGNYGEVKNGNGGGKKGGDDGGKKN-KGEKQKGGGGDWGDEKNSSKKKNGKGK 201
Query: 66 GNGNG-----NGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSNGNSNGNGE--------- 111
G G G G + G G + NG G + NS G GE
Sbjct: 202 NGGGGFLVKFLGLGKKSKKGGGAADTTNKKKNNGGGGNSNNSKGKDGKKGEKLDKVEFDF 381
Query: 112 -----------GNGKGNGEGNGKG--------NGEG-NGKGNG 134
G G GNG GNGKG NG G N GNG
Sbjct: 382 QDFDITPHGKSGKG-GNGNGNGKGAPAKGNANNGHGSNNNGNG 507
Score = 55.8 bits (133), Expect = 3e-08
Identities = 37/131 (28%), Positives = 46/131 (34%), Gaps = 21/131 (16%)
Frame = +1
Query: 7 QAQEKGSGNGNGNGNGNGNGNGNGNGNGNGN-----GNGNGNGNGNGNGNGNGNGNGNGN 61
+ Q+ G G+ N + NG G G G G G + G G + NG
Sbjct: 127 EKQKGGGGDWGDEKNSSKKKNGKGKNGGGGFLVKFLGLGKKSKKGGGAADTTNKKKNNGG 306
Query: 62 GNGNGNGNGNGNGNGNG-------------NGNGNGNGNGNGNGNGNG---NGNGNSNGN 105
G + N G G +G GNGNGNG G GN N+
Sbjct: 307 GGNSNNSKGKDGKKGEKLDKVEFDFQDFDITPHGKSGKGGNGNGNGKGAPAKGNANNGHG 486
Query: 106 SNGNGEGNGKG 116
SN NG G G
Sbjct: 487 SNNNGNGGHMG 519
Score = 45.1 bits (105), Expect = 6e-05
Identities = 31/106 (29%), Positives = 42/106 (39%), Gaps = 1/106 (0%)
Frame = +1
Query: 45 NGNGNGNGNGNGNGNGNGNGNGNGN-GNGNGNGNGNGNGNGNGNGNGNGNGNGNGNGNSN 103
+GN NGNG G G+ G N G G G+ N + NG G NG G
Sbjct: 46 SGNYGEVKNGNGGGKKGGDDGGKKNKGEKQKGGGGDWGDEKNSSKKKNGKGK-NGGGGFL 222
Query: 104 GNSNGNGEGNGKGNGEGNGKGNGEGNGKGNGDQPTKNDEASEYEVL 149
G G+ + KG G + + NG G +K + + E L
Sbjct: 223 VKFLGLGKKSKKGGGAADTTNKKKNNGGGGNSNNSKGKDGKKGEKL 360
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.309 0.134 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,780,514
Number of Sequences: 36976
Number of extensions: 280656
Number of successful extensions: 12030
Number of sequences better than 10.0: 242
Number of HSP's better than 10.0 without gapping: 1220
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7513
length of query: 458
length of database: 9,014,727
effective HSP length: 99
effective length of query: 359
effective length of database: 5,354,103
effective search space: 1922122977
effective search space used: 1922122977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0136.1


