

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
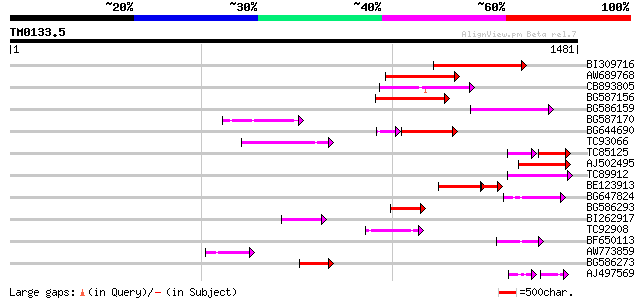
Query= TM0133.5
(1481 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 269 5e-72
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 185 1e-46
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 171 3e-42
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 164 3e-40
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 160 3e-39
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 149 8e-36
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 112 2e-33
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 129 1e-29
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 71 6e-25
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 111 2e-24
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 107 4e-23
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 100 5e-21
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 100 7e-21
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 90 6e-18
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 88 3e-17
TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol po... 86 1e-16
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 83 7e-16
AW773859 82 2e-15
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 81 3e-15
AJ497569 weakly similar to PIR|T04833|T04 hypothetical protein F... 48 1e-11
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 269 bits (688), Expect = 5e-72
Identities = 140/243 (57%), Positives = 180/243 (73%)
Frame = +2
Query: 1108 QVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDD 1167
+VC LQKS+YGLKQASRQW+S L ++L G+ QS +D +L+ K + SFT LL+YVDD
Sbjct: 17 KVCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDS-SFTTLLVYVDD 193
Query: 1168 VLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDS 1227
++LAGND+ EI+ VK L +F+IKDLG ++FLGL +ARS++GI+LNQRKY LELL DS
Sbjct: 194 IVLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDS 373
Query: 1228 GLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFL 1287
G L KS TP D S KL S +D + YRRLIG+L+YLTTTRPDI++ V QLSQF+
Sbjct: 374 GNLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFV 553
Query: 1288 SAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMF 1347
S P +H AA RVL+Y+K +P GLFY A S+ L++F+DSD A C TRKS+TGY +F
Sbjct: 554 SKPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVF 733
Query: 1348 LGS 1350
LGS
Sbjct: 734 LGS 742
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 185 bits (470), Expect = 1e-46
Identities = 91/191 (47%), Positives = 125/191 (64%)
Frame = +1
Query: 983 WRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQ 1042
W AM E +AL N TW LV PP K IGCKWVYR+K DG+++++KARLV KG++Q
Sbjct: 97 WLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQ 276
Query: 1043 IEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMN 1102
G D+ +TFSPV K T+R++L++ T W + Q+D++NAFL+ L EE+YMS PQG
Sbjct: 277 TLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGFE 456
Query: 1103 SDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALL 1162
+ + VC L KSLYGLKQA R W+ L A GFT+S D +L + + G+ L
Sbjct: 457 AANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLI-YNQNGACIYLX 633
Query: 1163 LYVDDVLLAGN 1173
+YVDD+L+ G+
Sbjct: 634 IYVDDILITGS 666
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 171 bits (432), Expect = 3e-42
Identities = 93/257 (36%), Positives = 150/257 (58%), Gaps = 7/257 (2%)
Frame = +3
Query: 965 ITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQ 1024
+T T +PT + EAVK E WR +MN E+EA ERNNTW L D IG KW+++ K +
Sbjct: 24 LTMTSDPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNE 203
Query: 1025 DGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAF 1084
+G I++YKARLV KGY+Q G+D+ + F+PVA+ T+R++++L + Q+ D
Sbjct: 204 NGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAA-------QIKRDGVC 362
Query: 1085 L------HAKLDEEIYMSLPQGMNSDKPNQVC-LLQKSLYGLKQASRQWFSTLCQALQGL 1137
+ H+ ++ + L + ++++LYGLKQA R W+S +
Sbjct: 363 IS*M*KAHSCMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKE 542
Query: 1138 GFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEA 1197
GF + +HTL+VK S G + LYVDD++ GND + + K S+ ++F + DLG+
Sbjct: 543 GFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKM 722
Query: 1198 KFFLGLAIARSQKGIIL 1214
+FLG+ + +++KGI +
Sbjct: 723 HYFLGVEVTQNEKGIYI 773
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 164 bits (414), Expect = 3e-40
Identities = 81/196 (41%), Positives = 125/196 (63%), Gaps = 2/196 (1%)
Frame = -1
Query: 956 PSYH-AFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGC 1014
P H AF++N+ P Y EA++ + W+ ++ E A+ +N+TW + P K +
Sbjct: 606 PEAHCAFMVNLNENHIPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSS 427
Query: 1015 KWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWF 1074
+W++ IKYK DG+I+R K RLV +G+T G D+++TF+PVAK+ T+R++LSL W
Sbjct: 426 RWIFTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWG 247
Query: 1075 LHQLDVDNAFLHAKLDEEIYMSLPQGM-NSDKPNQVCLLQKSLYGLKQASRQWFSTLCQA 1133
L Q+DV NAFL +L++E+YM P G+ + K V L+K++YGLKQ+ R W++ L
Sbjct: 246 LWQMDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTT 67
Query: 1134 LQGLGFTQSFADHTLY 1149
L G GF +S DHTL+
Sbjct: 66 LNGRGFRKSELDHTLF 19
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 160 bits (405), Expect = 3e-39
Identities = 79/215 (36%), Positives = 127/215 (58%)
Frame = +1
Query: 1205 IARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIG 1264
+ ++++GI + QRKY +LL G+ + P+ KL D + Y++++G
Sbjct: 7 VIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQIVG 186
Query: 1265 RLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLT 1324
L+YL TRPD+ YV++ +S+F++ PT++H A RVLRY+ G+ G+ Y S L
Sbjct: 187 CLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEKLE 366
Query: 1325 AFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQW 1384
A++DSD AG +D RKS +GY L S VSW SKKQ + S+ +AE+ A A C+ W
Sbjct: 367 AYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQSVW 546
Query: 1385 LIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYH 1419
+ +L+ L +Q+ ++M+CDN S + ++ NP H
Sbjct: 547 MRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 149 bits (376), Expect = 8e-36
Identities = 83/220 (37%), Positives = 128/220 (57%), Gaps = 7/220 (3%)
Frame = -3
Query: 555 NLVCEFSPSVQ----NLWHYRLGHPSFVKGQSIKDLFPYVQYTQDHVCEVCPIAKQKRLK 610
N C F+ S LWH RLGHP G+++ + P V + + CE C + K +
Sbjct: 632 NYKCSFTSSSSLNKDALWHARLGHPH---GRALNLMLPGVVFENKN-CEACILGKHCKNV 465
Query: 611 FPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLV 670
FP +++ + F +I+ D+W S LS + YF+T +D+ S+YTW+ L+ SKD V
Sbjct: 464 FPRTSTVYENCFDLIYTDLWTAPS-LSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAF 288
Query: 671 KDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYA---EKGIIHHTSCVETPQQNSIVERKH 727
K+F A+VTN + +KI+RSDNG E+ F + GI+H TSC TPQQN + +RK+
Sbjct: 287 KNFQAYVTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKN 108
Query: 728 QHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPIL 767
+H++ VAR+L+FQA+ + + +LIN +PT +L
Sbjct: 107 KHLMEVARSLMFQAN---------VSTACYLINWIPTKVL 15
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 112 bits (279), Expect(2) = 2e-33
Identities = 60/147 (40%), Positives = 93/147 (62%), Gaps = 1/147 (0%)
Frame = -2
Query: 1023 KQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDN 1082
KQ I R K++LVV+GY Q EGID+ + FSPVA+M +R+L++ + + L+Q+DV +
Sbjct: 439 KQA*VITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKS 260
Query: 1083 AFLHAKLDEEIYMSLPQGM-NSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQ 1141
AF++ L EE+++ P G +++ PN V L K+LYGLKQA R W+ L + L GF +
Sbjct: 259 AFINGDLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKR 80
Query: 1142 SFADHTLYVKKSATGSFTALLLYVDDV 1168
D+TL++ K + +YVDD+
Sbjct: 79 GKIDNTLFLLK-RE*ELLIIQVYVDDI 2
Score = 50.4 bits (119), Expect(2) = 2e-33
Identities = 25/65 (38%), Positives = 37/65 (56%)
Frame = -3
Query: 957 SYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKW 1016
S+ AFI +++EP EA++ W +M E+ ER+ W LV +P KT IG +W
Sbjct: 615 SFSAFI----SSIEPKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRW 448
Query: 1017 VYRIK 1021
V+R K
Sbjct: 447 VFRNK 433
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 129 bits (323), Expect = 1e-29
Identities = 82/247 (33%), Positives = 125/247 (50%), Gaps = 7/247 (2%)
Frame = +1
Query: 606 QKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDE 665
+K++ F + + I IH D+WGP V S G Y +TI+DD+ R WVY L+ K+E
Sbjct: 49 RKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNE 228
Query: 666 VQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLS---QFYAEKGIIHHTSCVETPQQNSI 722
K + V Q G +VK + +DN EF S +F GI H + PQQN +
Sbjct: 229 TFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGV 408
Query: 723 VERKHQHILNVARALLFQAHL--PKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQL 780
ER + +L AR +L A L + W A + L+NR P LD + P + L
Sbjct: 409 AERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGNL 588
Query: 781 PDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVY--DLNTRDISISR 838
D + L++FG +A + K RA C+FL + S +KGY ++ D ++ + +SR
Sbjct: 589 VDYSNLRIFGCPAYALV---NDGKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLILSR 759
Query: 839 NVIFHEN 845
+V F+E+
Sbjct: 760 DVTFNED 780
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 70.9 bits (172), Expect(2) = 6e-25
Identities = 33/85 (38%), Positives = 54/85 (62%)
Frame = +3
Query: 1381 EVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGL 1440
E W+ L+++L Q ++++CD+QSA+HIA NP++H RTKHI + H VRE + +G
Sbjct: 249 EAIWMQRLMEELGHKQEQ-ITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVREVVEEGS 425
Query: 1441 VHLLPVTSSLQLADIFTKPLSPAPF 1465
V + + ++ LAD TK ++ F
Sbjct: 426 VDMQKIHTNDNLADAMTKSINTDKF 500
Score = 63.2 bits (152), Expect(2) = 6e-25
Identities = 35/75 (46%), Positives = 44/75 (58%)
Frame = +1
Query: 1300 RVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKK 1359
R++RYIKG+ G + + + TV + DSD AG D RKS TGY L VSW SK
Sbjct: 7 RIMRYIKGTSGVAVCFGGSELTV-RGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLSKL 183
Query: 1360 QTTTSRSSCEAEYRA 1374
QT + S+ EAEY A
Sbjct: 184 QTVVALSTTEAEYMA 228
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 111 bits (278), Expect = 2e-24
Identities = 55/137 (40%), Positives = 83/137 (60%)
Frame = +2
Query: 1329 SDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYL 1388
SD AG +TRKS +GY LG+ +SW SKKQ + S+ EAEY A + + WL +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1389 LQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTS 1448
L+ + Q +P ++CDN+SA+ ++ NP +H R+KHI++ H +RE I + V + +
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1449 SLQLADIFTKPLSPAPF 1465
++ADIFTKPL F
Sbjct: 362 EEKIADIFTKPLKIESF 412
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 107 bits (266), Expect = 4e-23
Identities = 60/170 (35%), Positives = 98/170 (57%), Gaps = 2/170 (1%)
Frame = +1
Query: 1301 VLRYIKGSPGCGLFYPAASST--VLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSK 1358
VL+Y+ S L Y A+ L + D+D AG VDTRKS++G+ L + +SW++
Sbjct: 16 VLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTISWKAN 195
Query: 1359 KQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSY 1418
+Q+ + S+ +AEY A V + WL ++ +L ++Q V + CD+QSA+H+A++ Y
Sbjct: 196 QQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEY-VKIHCDSQSAIHLANHQVY 372
Query: 1419 HERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHI 1468
HERTKHI++ H +R+ I + + + S AD+FTK A H+
Sbjct: 373 HERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTKLKKHARATHV 522
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 100 bits (248), Expect = 5e-21
Identities = 47/122 (38%), Positives = 76/122 (61%)
Frame = +1
Query: 1121 QASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKL 1180
Q+ R WF ++ G+ Q DH +++K S+T L++YVDD+ L G+ IK
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1181 VKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMD 1240
+K+ L ++F IKDLG K+FLG+ +AR +KG ++QRKY L+LL ++ ++G K+ P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDPYG 360
Query: 1241 CS 1242
C+
Sbjct: 361 CN 366
Score = 55.5 bits (132), Expect = 2e-07
Identities = 26/55 (47%), Positives = 37/55 (67%)
Frame = +2
Query: 1233 KSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFL 1287
K + TPMD + KL L D Y+RL+G+L+YL+ TRPDI++VV +SQF+
Sbjct: 338 KPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 99.8 bits (247), Expect = 7e-21
Identities = 65/161 (40%), Positives = 91/161 (56%), Gaps = 1/161 (0%)
Frame = -3
Query: 1291 TDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGS 1350
T H AA ++L Y+K SP +F+P S + AF DSD ++ + S
Sbjct: 599 TAQHPQAAIQIL-YLKISPSQ*IFFP--S*IQIKAFCDSD*ID--QAA*TLENQSVIFAS 435
Query: 1351 SLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAM 1410
S + R + + YR++ +T+CE++WL YLL DL+ + P ++CDNQSA
Sbjct: 434 S*ATHSYAGNLKKKRYNFKI-YRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAA 258
Query: 1411 -HIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSL 1450
HIA N S+ ERTKHIELDCHIVR K+ L H+L + SSL
Sbjct: 257 RHIAANSSFLERTKHIELDCHIVRVKLQLKLFHILHILSSL 135
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 90.1 bits (222), Expect = 6e-18
Identities = 42/91 (46%), Positives = 62/91 (67%)
Frame = +2
Query: 996 RNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPV 1055
+ T LV KP PIG +W+Y+IK +DGT+ +YKARLV KGY + +GIDF + F+PV
Sbjct: 47 QKQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPV 226
Query: 1056 AKMTTLRVLLSLVSTKNWFLHQLDVDNAFLH 1086
++ T+ +LL+L +T +H +DV AFL+
Sbjct: 227 VRIETI*LLLALAATNGC*IHHIDVKIAFLN 319
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 87.8 bits (216), Expect = 3e-17
Identities = 40/119 (33%), Positives = 70/119 (58%)
Frame = +1
Query: 710 HTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN 769
++ TPQQN + ER ++ +L RA+L A + K FWA A+ + ++INR P+ ++D
Sbjct: 67 NSXVAHTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDL 246
Query: 770 QCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYD 828
+ P ++ + D + L VFG + + RTK D ++++C+FLG+ KGY ++D
Sbjct: 247 KTPMEMWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol polyprotein
{Oryza sativa}, partial (1%)
Length = 638
Score = 85.9 bits (211), Expect = 1e-16
Identities = 56/152 (36%), Positives = 82/152 (53%)
Frame = +2
Query: 929 STVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN 988
S +V NSSSK S+ + + Y + A + IT+ +EP Y +A + + W AM
Sbjct: 164 SHLVTLNSSSK--SYHIFHYVGY-SFSAKHRASLAAITSNIEPKNYVQAAQ*QEWLAAME 334
Query: 989 *EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDF 1048
EI+ LE NNT L K + C+ VY+I +K +G I++YKA+LV K + Q+EG DF
Sbjct: 335 QEIQVLEENNTSTLEPLREGKKWVDCRPVYKIIHKANGEIEKYKAQLVAKDFVQVEGEDF 514
Query: 1049 MDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDV 1080
D K R LL++ + K LH +DV
Sbjct: 515 *D-LCLSNKDDNCRCLLTIAAAKG*QLHLMDV 607
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 83.2 bits (204), Expect = 7e-16
Identities = 49/124 (39%), Positives = 72/124 (57%), Gaps = 3/124 (2%)
Frame = +1
Query: 1273 RPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYP-AASSTV--LTAFSDS 1329
RPDI Y V+ +S+F+ P H AA+R+LRY++G+ GL +P A S V L +SDS
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 1330 DGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLL 1389
D G R+S +GY + +SW +KKQ T+ SS EAEY A + WL ++
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 1390 QDLQ 1393
++L+
Sbjct: 472 KELK 483
>AW773859
Length = 538
Score = 82.0 bits (201), Expect = 2e-15
Identities = 44/128 (34%), Positives = 66/128 (51%)
Frame = -3
Query: 512 LIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYR 571
LIQ+ + KMIG+ LY L PS S+ F P + LWH+R
Sbjct: 383 LIQEKRSLKMIGSGELFEGLYFLTTQDT---PSATIASSQVQPQPQPTFLPQ-EALWHFR 216
Query: 572 LGHPSFVKGQSIKDLFPYVQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWG 631
LGH S K S+ FP++ Q+ VC++C ++ K+L F LS + + +++ H DIWG
Sbjct: 215 LGHLSNRKLLSLHSNFPFITIDQNSVCDICHYSRHKKLPFQLSTNRASKCYELFHFDIWG 36
Query: 632 PVSVLSLN 639
P S S++
Sbjct: 35 PFSTQSIH 12
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 80.9 bits (198), Expect = 3e-15
Identities = 34/88 (38%), Positives = 59/88 (66%)
Frame = -2
Query: 757 FLINRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLG 816
+LINR+PT +L +Q PF++L+++ P +T+++VFG LC+ R K + R+++ +F+G
Sbjct: 698 YLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMFIG 519
Query: 817 FKSGTKGYLVYDLNTRDISISRNVIFHE 844
+ + KGY YD R + +SR+V F E
Sbjct: 518 YSTTQKGYKCYDPEARRVLVSRDVKFIE 435
>AJ497569 weakly similar to PIR|T04833|T04 hypothetical protein F21P8.50 -
Arabidopsis thaliana, partial (4%)
Length = 723
Score = 48.1 bits (113), Expect(2) = 1e-11
Identities = 28/74 (37%), Positives = 41/74 (54%)
Frame = +2
Query: 1387 YLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPV 1446
++ + ++ V +CDN SA+HIA N +HERT H E D +IV+ ++ L+P
Sbjct: 242 FIFYMISAKHST*VLQYCDNISALHIAANMVFHERT*HRETDPYIVQ---GSRMLQLMPS 412
Query: 1447 TSSLQLADIFTKPL 1460
S Q A TKPL
Sbjct: 413 ASKDQPAYSLTKPL 454
Score = 40.8 bits (94), Expect(2) = 1e-11
Identities = 31/75 (41%), Positives = 41/75 (54%)
Frame = +3
Query: 1302 LRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQT 1361
L Y+K +PG +F AS+ S C + + T +C FL SSL+SW+SKKQ
Sbjct: 15 LHYLK-TPGKCIFVSNASNPHFNRGS------CPYSIR*TTEFC-FLSSSLISWKSKKQC 170
Query: 1362 TTSRSSCEAEYRAMA 1376
SRS EA RA+A
Sbjct: 171 VVSRSFSEA**RALA 215
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,241,436
Number of Sequences: 36976
Number of extensions: 803197
Number of successful extensions: 5852
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 4364
Number of HSP's successfully gapped in prelim test: 202
Number of HSP's that attempted gapping in prelim test: 1312
Number of HSP's gapped (non-prelim): 4781
length of query: 1481
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1373
effective length of database: 5,021,319
effective search space: 6894270987
effective search space used: 6894270987
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0133.5


