

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.1
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
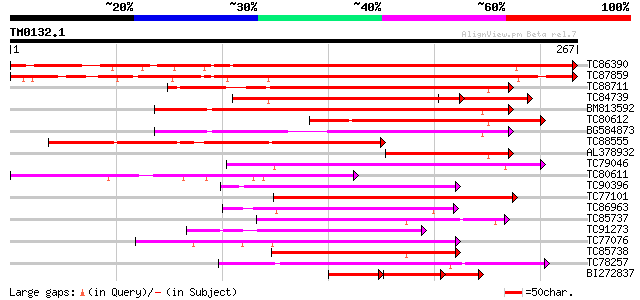
TC86390 similar to PIR|T52371|T52371 homeobox protein HAT9 [impo... 342 1e-94
TC87859 similar to SP|P46604|HT22_ARATH Homeobox-leucine zipper ... 322 9e-89
TC88711 similar to PIR|T07614|T07614 homeobox-leucine zipper pro... 192 9e-50
TC84739 homologue to PIR|T52367|T52367 homeobox protein HAT14 [i... 161 2e-40
BM813592 similar to GP|20161555|db putative homeodomain-leucine ... 160 6e-40
TC80612 similar to PIR|T52373|T52373 homeobox protein THOM1 [imp... 126 8e-30
BG584873 similar to GP|20161555|dbj putative homeodomain-leucine... 124 5e-29
TC88555 similar to PIR|T07614|T07614 homeobox-leucine zipper pro... 114 4e-26
AL378932 homologue to PIR|T07614|T076 homeobox-leucine zipper pr... 97 6e-21
TC79046 similar to PIR|T12634|T12634 homeotic protein - common s... 88 3e-18
TC80611 similar to PIR|T07614|T07614 homeobox-leucine zipper pro... 86 2e-17
TC90396 similar to PIR|H96719|H96719 homeobox gene 13 protein 1... 82 2e-16
TC77101 similar to GP|15148920|gb|AAK84887.1 homeodomain leucine... 82 3e-16
TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine... 79 2e-15
TC85737 similar to GP|6091551|gb|AAF01764.2| homeodomain-leucine... 76 1e-14
TC91273 similar to GP|1435022|dbj|BAA05625.1 DNA-binding protein... 75 2e-14
TC77076 similar to PIR|T07734|T07734 homeotic protein VAHOX1 - t... 75 3e-14
TC85738 similar to GP|6091551|gb|AAF01764.2| homeodomain-leucine... 74 8e-14
TC78257 similar to PIR|T47981|T47981 homeobox-leucine zipper pro... 74 8e-14
BI272837 similar to PIR|T52367|T523 homeobox protein HAT14 [impo... 50 1e-09
>TC86390 similar to PIR|T52371|T52371 homeobox protein HAT9 [imported] -
Arabidopsis thaliana, partial (58%)
Length = 1288
Score = 342 bits (876), Expect = 1e-94
Identities = 202/286 (70%), Positives = 217/286 (75%), Gaps = 19/286 (6%)
Frame = +2
Query: 1 MGLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYS-SKEPSLTLGLSG 59
MGL+ S LHL LGLSL T+ T KE TTTT M PYS S EPSLTLGLSG
Sbjct: 245 MGLNDQDS---LHLVLGLSLNTSTTPKEITTTTP--------MNPYSTSNEPSLTLGLSG 391
Query: 60 NN----------KVYCEDPLELSRQTS-PHSDVVSSFSTARV--VKRERVDLSCEEIEAE 106
+ K Y E EL RQTS PHS V SSFS+ RV VKRER D EE+E E
Sbjct: 392 ESYNLISHKQATKGYGE---ELCRQTSSPHSVVNSSFSSGRVLQVKRER-DEEEEEVE-E 556
Query: 107 ERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQV 166
ER+SSRVSDED+D TNARKKLRLTKEQS LLEESFK HSTLNPKQKQALARQLNLR RQV
Sbjct: 557 ERVSSRVSDEDEDATNARKKLRLTKEQSLLLEESFKLHSTLNPKQKQALARQLNLRPRQV 736
Query: 167 EVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAA 226
EVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELK+LK+AQPLYMPM AA
Sbjct: 737 EVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKSLKVAQPLYMPMPAA 916
Query: 227 TLTMCPSCERL-----GDGGSNIKSPFTITPKPHFFNPFTHPSAAC 267
TL++CPSCERL G GGSN + FT+ P HF+NPF +PSAAC
Sbjct: 917 TLSICPSCERLGRVADGGGGSNKITAFTMAPNTHFYNPFNNPSAAC 1054
>TC87859 similar to SP|P46604|HT22_ARATH Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22). [Mouse-ear cress]
{Arabidopsis thaliana}, partial (48%)
Length = 1109
Score = 322 bits (825), Expect = 9e-89
Identities = 193/286 (67%), Positives = 220/286 (76%), Gaps = 19/286 (6%)
Frame = +2
Query: 1 MGLDH-DASN-PGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSK-------EP 51
MGL++ DASN GL L LGL+LT T K+ T + + V KPYSS EP
Sbjct: 53 MGLNNQDASNNSGLQLILGLALTLT---KQDTPPSNNN-----VTKPYSSSNHNNYEAEP 208
Query: 52 SLTLGLSGNNKVYCEDPLELSRQT-SPHSDVVSSFSTARVVKRERVDLSCE---EIEAEE 107
SLTLGLS K Y E+PL+ S QT SPH VVSSFS+ RV KRER D+S E E E
Sbjct: 209 SLTLGLS--TKSYYEEPLDFSTQTNSPHHSVVSSFSSGRV-KRER-DVSSELEMETTEME 376
Query: 108 RLSSRVSDEDDDG-TNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQV 166
R+SSR+SDED+DG T ARKKLRLTK+QSA+LEESFK+HSTLNPKQKQALAR+LNL RQV
Sbjct: 377 RVSSRISDEDEDGATAARKKLRLTKDQSAMLEESFKEHSTLNPKQKQALARELNLTPRQV 556
Query: 167 EVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAA 226
EVWFQNRRARTKLKQTEVDCEFLKKCCETLT+ENRRL+KE+QELKALKL +PLYMPM AA
Sbjct: 557 EVWFQNRRARTKLKQTEVDCEFLKKCCETLTEENRRLQKEVQELKALKLVKPLYMPMPAA 736
Query: 227 TLTMCPSCERLG-----DGGSNIKSPFTITPKPHFFNPFTHPSAAC 267
TLTMCPSCERLG +GGS+ K+ F+ KPHF+NPFT+ SAAC
Sbjct: 737 TLTMCPSCERLGGGGGVNGGSSNKTNFS---KPHFYNPFTNSSAAC 865
>TC88711 similar to PIR|T07614|T07614 homeobox-leucine zipper protein
homolog h1 - soybean, partial (73%)
Length = 1238
Score = 192 bits (489), Expect = 9e-50
Identities = 105/164 (64%), Positives = 127/164 (77%), Gaps = 1/164 (0%)
Frame = +1
Query: 75 TSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQS 134
+SP+S VSS S R ++ E D+ E +SDE+D T ARKKLRL+K+QS
Sbjct: 385 SSPNS-TVSSVSGKRSLREEDHDVENRE---------NISDEEDAET-ARKKLRLSKDQS 531
Query: 135 ALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCE 194
A+LEE+FK+H+TLNPKQK ALA+QL LR RQVEVWFQNRRARTKLKQTEVDCE LK+CCE
Sbjct: 532 AILEETFKEHNTLNPKQKLALAKQLGLRPRQVEVWFQNRRARTKLKQTEVDCEVLKRCCE 711
Query: 195 TLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERL 237
LT+ENRRL+KE+QEL+ALKL+ YM M+ TLTMCPSCER+
Sbjct: 712 NLTEENRRLQKEVQELRALKLSPQFYMQMTPPTTLTMCPSCERV 843
>TC84739 homologue to PIR|T52367|T52367 homeobox protein HAT14 [imported] -
Arabidopsis thaliana (fragment), partial (70%)
Length = 737
Score = 161 bits (408), Expect = 2e-40
Identities = 81/111 (72%), Positives = 94/111 (83%), Gaps = 2/111 (1%)
Frame = +1
Query: 106 EERLSSRVSDEDDDG--TNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRA 163
++R SSRVSDEDD+ N RKKLRL+K+QSA LEESFK+H TLNPKQK ALA+QLNLR
Sbjct: 13 DQRNSSRVSDEDDNCGVRNTRKKLRLSKDQSAFLEESFKEHHTLNPKQKLALAKQLNLRP 192
Query: 164 RQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALK 214
RQVEVWFQNRRARTKLKQTEVDCE+LK+CCETLT+ENRR + +K+ K
Sbjct: 193 RQVEVWFQNRRARTKLKQTEVDCEYLKRCCETLTEENRRPTQGTSRIKSFK 345
Score = 50.8 bits (120), Expect = 5e-07
Identities = 24/44 (54%), Positives = 29/44 (65%)
Frame = +2
Query: 203 LKKELQELKALKLAQPLYMPMSAATLTMCPSCERLGDGGSNIKS 246
L KELQEL+ALK + P M + A TLTMCPSCER+ + S
Sbjct: 311 LHKELQELRALKTSNPFNMQLPATTLTMCPSCERVATNSTATSS 442
>BM813592 similar to GP|20161555|db putative homeodomain-leucine zipper
{Oryza sativa (japonica cultivar-group)}, partial (42%)
Length = 683
Score = 160 bits (404), Expect = 6e-40
Identities = 87/175 (49%), Positives = 120/175 (67%), Gaps = 6/175 (3%)
Frame = +2
Query: 69 LELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLR 128
LEL+ + SF + VK+ +D++ IE E + +G RKKLR
Sbjct: 20 LELTISIPGFASSPISFLPSSSVKK--LDVNRVTIEEEWMALEEEEESSVNGDTPRKKLR 193
Query: 129 LTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEF 188
LTKEQS LLEESF+++ TLNPKQK+ LA QL LR RQVEVWFQNRRAR+KLKQTE++CE+
Sbjct: 194 LTKEQSHLLEESFRKNHTLNPKQKECLAMQLKLRPRQVEVWFQNRRARSKLKQTEMECEY 373
Query: 189 LKKCCETLTDENRRLKKELQELKALKLAQPLYM------PMSAATLTMCPSCERL 237
LK+ +LT++NRRL++E++EL+A+K+ P + P+ A+TL+MCP CER+
Sbjct: 374 LKRWFGSLTEQNRRLQREVEELRAMKVGPPTVLSPHSSEPLPASTLSMCPRCERV 538
>TC80612 similar to PIR|T52373|T52373 homeobox protein THOM1 [imported] -
tomato, partial (35%)
Length = 622
Score = 126 bits (317), Expect = 8e-30
Identities = 67/112 (59%), Positives = 82/112 (72%), Gaps = 1/112 (0%)
Frame = +1
Query: 142 KQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENR 201
K+H+TLNP +KQAL + A+ FQNRRARTKLKQ EVDCE+LK+CCETLTDENR
Sbjct: 1 KEHNTLNPNKKQALQSS*S-EAKTS*GVFQNRRARTKLKQPEVDCEYLKRCCETLTDENR 177
Query: 202 RLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNIKSPFTITP 252
RL+KE+QEL+ALKL+ LYM M+ TLTMCPSCER+ ++ S P
Sbjct: 178 RLQKEVQELRALKLSPQLYMQMNPPTTLTMCPSCERVAVSSASSSSAAAAPP 333
>BG584873 similar to GP|20161555|dbj putative homeodomain-leucine zipper
{Oryza sativa (japonica cultivar-group)}, partial (31%)
Length = 737
Score = 124 bits (310), Expect = 5e-29
Identities = 74/175 (42%), Positives = 104/175 (59%), Gaps = 6/175 (3%)
Frame = +1
Query: 69 LELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLR 128
LEL+ + SF + VK+ +D++ IE E + +G RKKLR
Sbjct: 115 LELTISIPGFASSPISFLPSSSVKK--LDVNRVTIEEEWMALEEEEESSVNGDTPRKKLR 288
Query: 129 LTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEF 188
LTK KQK+ LA QL LR RQVEVWFQNRRAR+KLKQTE++CE+
Sbjct: 289 LTK------------------KQKECLAMQLKLRPRQVEVWFQNRRARSKLKQTEMECEY 414
Query: 189 LKKCCETLTDENRRLKKELQELKALKLAQPLYM------PMSAATLTMCPSCERL 237
LK+ +LT++NRRL++E++EL+A+K+ P + P+ A+TL+MCP CER+
Sbjct: 415 LKRWFGSLTEQNRRLQREVEELRAMKVGPPTVLSPHSSEPLPASTLSMCPRCERV 579
>TC88555 similar to PIR|T07614|T07614 homeobox-leucine zipper protein
homolog h1 - soybean, partial (47%)
Length = 704
Score = 114 bits (285), Expect = 4e-26
Identities = 75/159 (47%), Positives = 99/159 (62%)
Frame = +3
Query: 19 SLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNNKVYCEDPLELSRQTSPH 78
S +T N K + TSS + + + P + G+ N D E + +SP+
Sbjct: 249 SPSTFNPQKPSWNDVFTSSDRDSETCRIEER-PLILRGIDVNRLPSGADCEEEAGVSSPN 425
Query: 79 SDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLE 138
S VSS S KR +++ E+++ E S +SDE+D T+ RKKLRLTK+QS +LE
Sbjct: 426 S-TVSSVSG----KRSEREVTGEDLDMERDCSRGISDEEDAETS-RKKLRLTKDQSIILE 587
Query: 139 ESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRART 177
ESFK+H+TLNPKQK ALA+QL LRARQVEVWFQNRRART
Sbjct: 588 ESFKEHNTLNPKQKLALAKQLGLRARQVEVWFQNRRART 704
>AL378932 homologue to PIR|T07614|T076 homeobox-leucine zipper protein
homolog h1 - soybean, partial (23%)
Length = 423
Score = 97.1 bits (240), Expect = 6e-21
Identities = 46/61 (75%), Positives = 53/61 (86%), Gaps = 1/61 (1%)
Frame = +2
Query: 178 KLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCER 236
KLKQTEVDCEFLK+CCE LTDENRRL+KE+QEL+ALKL+ YM M+ TLTMCPSCER
Sbjct: 2 KLKQTEVDCEFLKRCCENLTDENRRLQKEVQELRALKLSPQFYMQMTPPTTLTMCPSCER 181
Query: 237 L 237
+
Sbjct: 182 V 184
>TC79046 similar to PIR|T12634|T12634 homeotic protein - common sunflower,
partial (56%)
Length = 1136
Score = 88.2 bits (217), Expect = 3e-18
Identities = 60/153 (39%), Positives = 81/153 (52%), Gaps = 3/153 (1%)
Frame = +2
Query: 103 IEAEERLSSRVSDEDDDGTNA-RKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNL 161
IE E + D DDG+ A KK RL EQ LE+SF+ + L P++K LAR LNL
Sbjct: 332 IELGEEANIPEEDLSDDGSQAGEKKRRLNMEQVKTLEKSFELGNKLEPERKMQLARALNL 511
Query: 162 RARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYM 221
+ RQV +WFQNRRAR K KQ E D + LK+ + + +N L+ + Q+L+A LA
Sbjct: 512 QPRQVAIWFQNRRARWKTKQLEKDYDVLKRQYDAIKLDNDALQAQNQKLQAEILALKNRE 691
Query: 222 PMSAATLTMCP--SCERLGDGGSNIKSPFTITP 252
P + L S + S IK + TP
Sbjct: 692 PTESINLNKETEGSSSNRSENSSEIKLDMSRTP 790
>TC80611 similar to PIR|T07614|T07614 homeobox-leucine zipper protein
homolog h1 - soybean, partial (32%)
Length = 816
Score = 85.5 bits (210), Expect = 2e-17
Identities = 71/191 (37%), Positives = 95/191 (49%), Gaps = 27/191 (14%)
Frame = +2
Query: 1 MGLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKP------------YSS 48
MG D + GL L+LG N ++ + P TV S+
Sbjct: 260 MGEKDDEFSLGLSLSLGFGEYANNNNQTPLKVSHMHKPPQTVTNQRVSFNNLFHFHDLST 439
Query: 49 KEPSLTLGLSGNNKVYCEDPLELSRQTSPHSD----VVSSFSTARVV--KRERVDLSCEE 102
+ S G+ GN PL + T+ D V S ST + KR + + +E
Sbjct: 440 ETRSFLRGIDGNT------PLPSTAATAAAFDDENGVSSPNSTVSSISGKRSEREGNGDE 601
Query: 103 IEAEERLSSRV---SDEDD------DGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQ 153
+A ER S SD+DD DG ++RKKLRL+KEQS LLEE+FK+H+TLNPKQKQ
Sbjct: 602 NDAVERGSCSRGGGSDDDDGGGCGGDGDSSRKKLRLSKEQSMLLEETFKEHNTLNPKQKQ 781
Query: 154 ALARQLNLRAR 164
ALA+QLNL+ R
Sbjct: 782 ALAKQLNLKPR 814
>TC90396 similar to PIR|H96719|H96719 homeobox gene 13 protein 11736-10437
[imported] - Arabidopsis thaliana, partial (41%)
Length = 997
Score = 82.0 bits (201), Expect = 2e-16
Identities = 47/113 (41%), Positives = 68/113 (59%)
Frame = +1
Query: 100 CEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQL 159
C+E+ + ++SDE+ KK RL+ EQ LE+SF+ + L P++K LA+ L
Sbjct: 193 CDELVHGDE--DQLSDEEGYSQMGEKKKRLSLEQVKALEKSFEIGNKLEPERKMQLAKAL 366
Query: 160 NLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKA 212
L+ RQV +WFQNRRAR K KQ E + E LKK ++L +N LK + +L A
Sbjct: 367 GLQPRQVAIWFQNRRARWKTKQLEKEYEVLKKQFDSLKADNNTLKAQNNKLHA 525
>TC77101 similar to GP|15148920|gb|AAK84887.1 homeodomain leucine zipper
protein HDZ3 {Phaseolus vulgaris}, complete
Length = 1532
Score = 81.6 bits (200), Expect = 3e-16
Identities = 46/115 (40%), Positives = 69/115 (60%)
Frame = +1
Query: 125 KKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEV 184
KK RLT EQ +LE+SF++ + L P++K LA++L L+ RQV VWFQNRRAR K KQ E
Sbjct: 631 KKRRLTSEQVHMLEKSFEEENKLEPERKTQLAKKLGLQPRQVAVWFQNRRARWKTKQLER 810
Query: 185 DCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERLGD 239
D + LK ++L + KE ++LK+ ++ + + A + P E+ D
Sbjct: 811 DYDVLKSSYDSLLSTYDSINKENEKLKSEVVSLNEKLQVQAKDMLEEPLSEKKAD 975
>TC86963 similar to GP|15148918|gb|AAK84886.1 homeodomain leucine zipper
protein HDZ2 {Phaseolus vulgaris}, partial (87%)
Length = 1519
Score = 78.6 bits (192), Expect = 2e-15
Identities = 50/120 (41%), Positives = 69/120 (56%), Gaps = 9/120 (7%)
Frame = +1
Query: 101 EEIEAEERLSSRVSDEDDDGTNAR--KKLRLTKEQSALLEESFKQHSTLNPKQKQALARQ 158
+++E EE DED + + KK RL+ EQ LE+SF+ + L P +K LA++
Sbjct: 667 QQLEKEENCG----DEDYEACYHQQGKKRRLSSEQVQFLEKSFEVENKLEPDRKVQLAKE 834
Query: 159 LNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTD-------ENRRLKKELQELK 211
L L+ RQV +WFQNRRAR K KQ E D LK ++L D EN +LK+E+ LK
Sbjct: 835 LGLQPRQVAIWFQNRRARFKTKQLEKDYGTLKASFDSLKDDYDNLLQENDKLKEEVNSLK 1014
>TC85737 similar to GP|6091551|gb|AAF01764.2| homeodomain-leucine zipper
protein 56 {Glycine max}, partial (63%)
Length = 1622
Score = 76.3 bits (186), Expect = 1e-14
Identities = 48/129 (37%), Positives = 74/129 (57%), Gaps = 10/129 (7%)
Frame = +2
Query: 117 DDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRAR 176
++ G ++ KK RL +Q LE++F+ + L P++K+ LA +L L+ RQV VWFQNRRAR
Sbjct: 530 EETGHHSEKKRRLRVDQVKALEKNFEVENKLEPERKEKLAIELGLQPRQVAVWFQNRRAR 709
Query: 177 TKLKQTEVD-------CEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAAT-- 227
K KQ E D + LK + + +N+ KE++ELK+ KL + ++
Sbjct: 710 WKTKQLERDYGVLKANYDALKLKFDAIAQDNKAFHKEIKELKS-KLGEEEKSTINVLVKE 886
Query: 228 -LTMCPSCE 235
LTM SC+
Sbjct: 887 ELTMLESCD 913
>TC91273 similar to GP|1435022|dbj|BAA05625.1 DNA-binding protein {Daucus
carota}, partial (33%)
Length = 722
Score = 75.5 bits (184), Expect = 2e-14
Identities = 47/113 (41%), Positives = 62/113 (54%)
Frame = +1
Query: 84 SFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQ 143
SF R + ++L EE EE LS DD KK RL EQ LE+SF+
Sbjct: 400 SFLGKRSMSFSGIELG-EEANVEEELS------DDGSQLGEKKRRLNMEQVKTLEKSFEL 558
Query: 144 HSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETL 196
+ L P++K LAR L L+ RQ+ +WFQNRRAR K KQ E D + L + +T+
Sbjct: 559 GNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDVLXRQYDTI 717
>TC77076 similar to PIR|T07734|T07734 homeotic protein VAHOX1 - tomato,
partial (43%)
Length = 1664
Score = 75.1 bits (183), Expect = 3e-14
Identities = 54/165 (32%), Positives = 84/165 (50%), Gaps = 12/165 (7%)
Frame = +1
Query: 60 NNKVYCEDPLELSRQTSPHSDVVSSF-STARVVKRERVDLSCEEIEAEER---------- 108
N + ++ + R S + S F S++ R +S E+++ +R
Sbjct: 409 NMNILLQNQQQTPRGNSSQQPLDSLFLSSSASFFGSRSMVSFEDVQGRKRRNRSFFGGFD 588
Query: 109 LSSRVSDEDDDGTN-ARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVE 167
L DE D+ + + KK RL+ +Q LE+SF++ + L P++K LA+ L L+ RQV
Sbjct: 589 LDENGEDEMDEYFHQSEKKRRLSVDQVQFLEKSFEEDNKLEPERKTKLAKDLGLQPRQVA 768
Query: 168 VWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKA 212
+WFQNRRAR K KQ E D + L E+L E L KE L++
Sbjct: 769 IWFQNRRARWKTKQLEKDYDSLNDGYESLKTEYDNLLKEKDRLQS 903
>TC85738 similar to GP|6091551|gb|AAF01764.2| homeodomain-leucine zipper
protein 56 {Glycine max}, partial (77%)
Length = 2144
Score = 73.6 bits (179), Expect = 8e-14
Identities = 43/96 (44%), Positives = 59/96 (60%), Gaps = 7/96 (7%)
Frame = +3
Query: 124 RKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTE 183
+KK RL+ +Q LE++F+ + L P +K LA++L L+ RQV VWFQNRRAR K KQ E
Sbjct: 1047 QKKRRLSVDQVKALEKNFEVENKLEPDRKVKLAQELGLQPRQVAVWFQNRRARWKTKQLE 1226
Query: 184 VD-------CEFLKKCCETLTDENRRLKKELQELKA 212
D + LK CE L + L KE++ELK+
Sbjct: 1227 RDYGVLKANYDALKLNCEDLQRDKETLLKEVKELKS 1334
>TC78257 similar to PIR|T47981|T47981 homeobox-leucine zipper protein
ATHB-12 - Arabidopsis thaliana, partial (45%)
Length = 994
Score = 73.6 bits (179), Expect = 8e-14
Identities = 53/163 (32%), Positives = 76/163 (46%), Gaps = 7/163 (4%)
Frame = +2
Query: 99 SCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQ 158
S E E E +S +S K R T EQ LE F+ + L P++K LAR+
Sbjct: 173 SAEAGEEETYTTSSISSMRKKKNKNTK--RFTDEQIKSLETMFETETRLEPRKKLQLARE 346
Query: 159 LNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKE-------LQELK 211
L L+ RQV +WFQN+RAR K KQ E + L+ L + +KKE LQ+L
Sbjct: 347 LGLQPRQVAIWFQNKRARWKSKQLEREYNKLQNSYNNLASKFESMKKERQTLLIQLQKLN 526
Query: 212 ALKLAQPLYMPMSAATLTMCPSCERLGDGGSNIKSPFTITPKP 254
L + +P+ S++ + S E + G K + P P
Sbjct: 527 DL-IQKPIEQSQSSSQVKEAKSMESASENGGRNKCEAEVKPSP 652
>BI272837 similar to PIR|T52367|T523 homeobox protein HAT14 [imported] -
Arabidopsis thaliana (fragment), partial (44%)
Length = 650
Score = 50.4 bits (119), Expect(2) = 1e-09
Identities = 21/29 (72%), Positives = 25/29 (85%)
Frame = +1
Query: 177 TKLKQTEVDCEFLKKCCETLTDENRRLKK 205
TKLKQTEVDCE+L +CCET T+ N RL+K
Sbjct: 490 TKLKQTEVDCEYLXRCCETXTEXNXRLQK 576
Score = 46.6 bits (109), Expect = 1e-05
Identities = 22/26 (84%), Positives = 23/26 (87%)
Frame = +1
Query: 151 QKQALARQLNLRARQVEVWFQNRRAR 176
QK ALA+QLNL RQVEVWFQNRRAR
Sbjct: 1 QKLALAKQLNLSPRQVEVWFQNRRAR 78
Score = 28.9 bits (63), Expect(2) = 1e-09
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +2
Query: 204 KKELQELKALKLAQPLYMPM 223
K++LQEL ALK QP YM +
Sbjct: 572 KRDLQELXALKTTQPFYMQL 631
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,785,774
Number of Sequences: 36976
Number of extensions: 98238
Number of successful extensions: 1114
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1000
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0132.1


