

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.8
(488 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
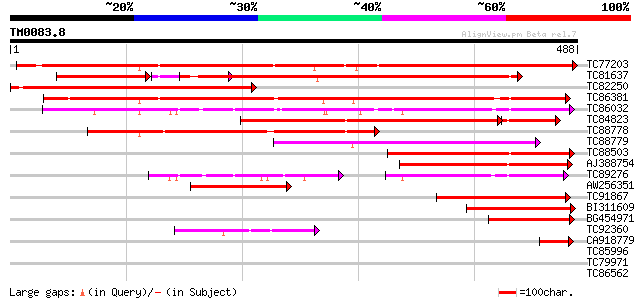
Score E
Sequences producing significant alignments: (bits) Value
TC77203 similar to GP|22208353|emb|CAC94006. endo-beta-1 4-gluca... 447 e-126
TC81637 homologue to PIR|JC7226|JC7226 endo-1 3(4)-beta-glucanas... 322 e-115
TC82250 homologue to PIR|T06770|T06770 cellulase (EC 3.2.1.4) pr... 365 e-101
TC86381 similar to PIR|E96786|E96786 protein F10A5.13 [imported]... 360 e-100
TC86032 similar to PIR|T07612|T07612 cellulase (EC 3.2.1.4) Cel3... 320 6e-88
TC84823 weakly similar to GP|3549291|gb|AAC95009.1| endo-1 4-bet... 240 8e-76
TC88778 similar to GP|1039431|gb|AAC78504.1| cellulase {Phaseolu... 244 5e-65
TC88779 similar to GP|1039431|gb|AAC78504.1| cellulase {Phaseolu... 210 9e-55
TC88503 similar to GP|1657380|emb|CAA65600.1 endo-beta-1 4-gluca... 202 2e-52
AJ388754 similar to GP|16648911|gb putative endo-1 4-beta-gluca... 201 4e-52
TC89276 weakly similar to GP|21740771|emb|CAD41248. OSJNBa0067K0... 121 1e-50
AW256351 similar to GP|12957206|dbj beta-1 4-glucanase {Atriplex... 123 1e-28
TC91867 weakly similar to GP|11094813|gb|AAG29742.1 endo-beta-1 ... 121 7e-28
BI311609 similar to PIR|T01108|T011 cellulase (EC 3.2.1.4) T21L1... 96 2e-20
BG454971 similar to GP|1039431|gb|A cellulase {Phaseolus vulgari... 95 5e-20
TC92360 similar to PIR|A96668|A96668 probable endo-beta-1 4-gluc... 59 5e-09
CA918779 similar to GP|6009979|dbj| endo-1 4-beta-glucanase {Pis... 44 1e-04
TC85996 weakly similar to GP|16648871|gb|AAL24287.1 Unknown prot... 30 1.6
TC79971 similar to GP|11994319|dbj|BAB02278. casein kinase {Arab... 30 1.6
TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regu... 30 2.1
>TC77203 similar to GP|22208353|emb|CAC94006. endo-beta-1 4-glucanase
{Fragaria x ananassa}, partial (92%)
Length = 2243
Score = 447 bits (1151), Expect = e-126
Identities = 237/495 (47%), Positives = 319/495 (63%), Gaps = 13/495 (2%)
Frame = +1
Query: 7 SFMSLLFLI-PMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFD 65
S + L FL+ P L F G +Y +AL+KS++F + QRSG LP +Q++ WRS SGL D
Sbjct: 181 SMLPLFFLLLPSLSFAGG----HDYGQALSKSIMFFEAQRSGYLPHNQRVSWRSHSGLQD 348
Query: 66 GRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGT 123
G+ + VDL GGYYDAGDNVKF PMAFT TM+SWS IEYGK+M +L A A++WGT
Sbjct: 349 GKASGVDLVGGYYDAGDNVKFGLPMAFTVTMMSWSIIEYGKQMASSGELGHAMEAVKWGT 528
Query: 124 DYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAAL 183
DY +K A P LY VGD N DHKCW+RPEDM T R Y V P+NPGSD+A ETAAA+
Sbjct: 529 DYFIK-AHPQPNLLYGEVGDGNTDHKCWQRPEDMTTDRRAYKVDPSNPGSDLAGETAAAM 705
Query: 184 AAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLW 243
AAAS+VF++ +P YS+ L+R A +++ FA +Y+G Y S+ A +Y S SG+ DELLW
Sbjct: 706 AAASIVFRRSNPAYSRELLRHAYELFDFADKYRGKYDSSITVAQ-KYYRSISGYNDELLW 882
Query: 244 GAAWLFRATNAVKYYNLI-----KTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKN- 297
AAWL++A+N Y + + G Q FSWD KY G L+++ L+ G
Sbjct: 883 AAAWLYQASNNQYYLDYLGRNGDSMGGTGWQMTEFSWDVKYPGVQTLVAK-FLMQGKAGV 1059
Query: 298 ----FDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKY 353
F++Y Q +E FMC + + + Q T GGL+++ +NLQ+ TS +FL T Y+ Y
Sbjct: 1060HAPVFEKYLQRSEYFMCSCIGKG-TRNIQKTPGGLIYRQRWNNLQFATSSSFLATVYSDY 1236
Query: 354 MSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSL 413
+++ C + V + L S AK QVDYILG+NP SYMVGYG N+P+R+HHRGSS+
Sbjct: 1237LASSGRNLKCASGNVPSSQLLSFAKSQVDYILGDNPRATSYMVGYGNNYPQRVHHRGSSI 1416
Query: 414 PSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYI 473
S+ A+P + C+GG+ +F S N NIL GAIVGGP+ D F D+R +Y +EP TY
Sbjct: 1417VSIKANPNFVSCNGGYATWFSSKKSNTNILTGAIVGGPDAYDDFADERKNYEQTEPATYN 1596
Query: 474 NGAVVGPLAYFAGIH 488
N ++G LA +G H
Sbjct: 1597NAPLIGVLARLSGGH 1641
>TC81637 homologue to PIR|JC7226|JC7226 endo-1 3(4)-beta-glucanase (EC
3.2.1.6) - garden pea, partial (82%)
Length = 1251
Score = 322 bits (826), Expect(2) = e-115
Identities = 158/299 (52%), Positives = 207/299 (68%), Gaps = 4/299 (1%)
Frame = +2
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
D K W E + +PGS+VAAETAAALAAASLVF++ DPTYSK+LVR A
Sbjct: 374 DQKTWTHQE------VFLRLMQTHPGSEVAAETAAALAAASLVFRRTDPTYSKILVRRAI 535
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT--- 263
+V+QFA Q++G YS+ L VCP YC YSG++DELLWGAAW+ +AT Y N I+
Sbjct: 536 RVFQFADQHRGPYSNVLKPFVCPLYCDYSGYQDELLWGAAWVHKATKNPMYLNYIQVNGQ 715
Query: 264 -LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQ 322
LGA + + F WD+K+ GA +LLS+ L+ ++ Y+ ++NF+C ++P + SSS Q
Sbjct: 716 ILGAAEFDNTFGWDNKHVGARILLSKEFLVQRVRSLHDYKGHSDNFVCSLIPGAGSSSAQ 895
Query: 323 YTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVD 382
YT GGL+FK+ DSN+QYVTS +FLL YAKY++ CG +VTP LR++AK+QVD
Sbjct: 896 YTPGGLLFKMSDSNMQYVTSTSFLLGAYAKYLTKSHSVVRCGGTIVTPKRLRTLAKKQVD 1075
Query: 383 YILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPN 441
Y+LG+NPL+MSYMVGYGP +P+RIHHRGSSL S+A HP I C GF S +PNPN
Sbjct: 1076YLLGDNPLKMSYMVGYGPRYPQRIHHRGSSLASMAVHPGKIQCSAGFS-VMSSQSPNPN 1249
Score = 110 bits (276), Expect(2) = e-115
Identities = 52/81 (64%), Positives = 60/81 (73%)
Frame = +3
Query: 41 LQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWS 100
L+ + G LP +Q++ WR DSGL DG +VDL GGYYDAGDNVKF FPMAFTTTMLSWS
Sbjct: 39 LKAKDQGSLPSNQRMSWRKDSGLSDGSAMHVDLVGGYYDAGDNVKFGFPMAFTTTMLSWS 218
Query: 101 AIEYGKRMGPQLKEARAAIRW 121
IE+G M +L AR AIRW
Sbjct: 219 VIEFGGLMKGELPNAREAIRW 281
Score = 65.5 bits (158), Expect = 4e-11
Identities = 32/70 (45%), Positives = 42/70 (59%)
Frame = +1
Query: 123 TDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAA 182
TDYLLK AT+ P +YV VGD DH CWERPEDMDT R+V+ + N P +
Sbjct: 286 TDYLLK-ATAHPNTIYVQVGDAKKDHACWERPEDMDTPRSVFKIDANAPWF*SCCRNCCS 462
Query: 183 LAAASLVFKK 192
+++ F+K
Sbjct: 463 ISSCFSCFQK 492
Score = 37.0 bits (84), Expect = 0.017
Identities = 18/32 (56%), Positives = 24/32 (74%), Gaps = 2/32 (6%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLP--PDQQLKWR 58
NYR+AL KS+LF +GQRSG+ P P+ L+ R
Sbjct: 2 NYRDALTKSILFFEGQRSGKPPF*PENVLEKR 97
>TC82250 homologue to PIR|T06770|T06770 cellulase (EC 3.2.1.4) precursor -
garden pea, partial (41%)
Length = 719
Score = 365 bits (937), Expect = e-101
Identities = 181/212 (85%), Positives = 191/212 (89%)
Frame = +1
Query: 1 MMRTSSSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSD 60
+MRTS F L L+ LL V NVQC PNYREALAKSLLF QGQRSGRLPPDQQ+KWRS+
Sbjct: 91 IMRTSLVF---LLLLTCLLVVENVQCKPNYREALAKSLLFFQGQRSGRLPPDQQIKWRSN 261
Query: 61 SGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIR 120
SG+ DGRLANVDL+GGYYDAGDNVKFNFPMAFTTTMLSWS IEYGKRMGPQ+KEARAAIR
Sbjct: 262 SGMSDGRLANVDLTGGYYDAGDNVKFNFPMAFTTTMLSWSTIEYGKRMGPQMKEARAAIR 441
Query: 121 WGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETA 180
TDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDT RTVY+VS NPGSDVAAETA
Sbjct: 442 HATDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTVRTVYFVSSKNPGSDVAAETA 621
Query: 181 AALAAASLVFKKVDPTYSKLLVRTAQKVYQFA 212
AALAAAS+VF KVDPTYSKLL+RTAQKVY FA
Sbjct: 622 AALAAASIVFXKVDPTYSKLLLRTAQKVYXFA 717
>TC86381 similar to PIR|E96786|E96786 protein F10A5.13 [imported] -
Arabidopsis thaliana, partial (92%)
Length = 2291
Score = 360 bits (924), Expect = e-100
Identities = 195/468 (41%), Positives = 288/468 (60%), Gaps = 15/468 (3%)
Frame = +2
Query: 30 YREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNFP 89
Y AL ++ F Q+SG+L + ++ WR DS L DG+ A++DL+ G YDAGD++KF FP
Sbjct: 590 YASALKTAMQFFDVQKSGKLE-NNKISWRGDSALKDGKQADLDLTKGMYDAGDHMKFGFP 766
Query: 90 MAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVD 147
MAFT ++LSW+ +EYG +M QL+ A+ ++RW TD+L+ A + LY+ VGDP D
Sbjct: 767 MAFTASVLSWAILEYGDQMDAVGQLEPAQDSLRWITDFLVN-AHPSENVLYIQVGDPVAD 943
Query: 148 HKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQK 207
HKCW RPE + R + V+ + PGSDVAAETAAA+A+ASLVFKK D TYS L++ A++
Sbjct: 944 HKCWNRPELITEERPLLQVNVSCPGSDVAAETAAAMASASLVFKKSDATYSSTLLKHAKQ 1123
Query: 208 VYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGAD 267
++ FA +++G YS+++ Y + +G+ DELLW A WL+ AT Y + +
Sbjct: 1124 LFTFADKHRGIYSENIPEVAT--YYNSTGYGDELLWAATWLYHATGDDSYLQYVTGQNGE 1297
Query: 268 D-----QPDIFSWDDKYAGAHVLLSRRALLNG-------DKNFDQYRQEAENFMCKILPN 315
D P FSWD+K AG VLLSR + YR+ AE MC +LP+
Sbjct: 1298 DYAQFGSPTWFSWDNKLAGTQVLLSRVSFFKAKGLSSSFGSGLQNYRKSAEAVMCGLLPD 1477
Query: 316 SPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYM-SAKKYTFNCGNVLVTPNTLR 374
SP+++ T GL++ + LQ+ + FL + Y+ YM + + C + P+ LR
Sbjct: 1478 SPTATKSRTDNGLIWVSEWNALQHPVASAFLASVYSDYMLTTQTPNIKCDSDSFKPSDLR 1657
Query: 375 SVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFH 434
A+ Q DY+LG+NP MS++VGYG NFP+ +HHRG+S+P+ A GC G+Q +
Sbjct: 1658 DFARSQADYVLGKNPQHMSFLVGYGKNFPQFVHHRGASIPANA----KTGCKDGWQ-WLD 1822
Query: 435 SSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
SS+PNPN+ GA+VGGP ++ + D R++ EP+TY + +VG L+
Sbjct: 1823 SSDPNPNVATGALVGGPFLNETYIDSRNNSMQGEPSTYNSAVIVGLLS 1966
>TC86032 similar to PIR|T07612|T07612 cellulase (EC 3.2.1.4) Cel3
membrane-anchored - tomato, complete
Length = 2536
Score = 320 bits (821), Expect = 6e-88
Identities = 207/495 (41%), Positives = 279/495 (55%), Gaps = 37/495 (7%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANV-----DLSGGYYDAGDN 83
NY AL K+L+F Q+SG+LP + WR +SG+ DG+ V DL GGYYDAGD
Sbjct: 530 NYTLALHKALMFFNAQKSGKLPKHNNVSWRGNSGMQDGKGDGVSAAIKDLVGGYYDAGDA 709
Query: 84 VKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCATSTPGRL---- 137
+KFNFP AF+ TMLSWS IEY + +L + I+WGTDY LK ST +
Sbjct: 710 IKFNFPQAFSITMLSWSVIEYSGKYEATGELNHVKELIKWGTDYFLKTFNSTADTITTLA 889
Query: 138 -YVGVG-------DPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLV 189
VG G PN DH CW RPED+D R V + + SD+AAE AAALAAAS+V
Sbjct: 890 AQVGSGVTGEDSTTPN-DHYCWMRPEDIDYDRPV---TECHSCSDLAAEMAAALAAASIV 1057
Query: 190 FKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLF 249
FK + YSK LV A +++F+ +G YS A FY S +G+ DE +WG AW++
Sbjct: 1058 FKD-NKAYSKKLVHGATTLFKFSRDQRGRYSPGRSEAAT-FYNS-TGYFDEFVWGGAWMY 1228
Query: 250 RATNAVKYYNLIKTLGADDQ-------PD--IFSWDDKYAGAHVLLSR-RALLNGDKNFD 299
AT Y L T G PD +FSWD+K AGA VLLSR R L+ ++
Sbjct: 1229 FATGNNSYLKLATTPGIAKHAGAFWGGPDYGVFSWDNKLAGAQVLLSRLRLFLSPGYPYE 1408
Query: 300 Q----YRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSN---LQYVTSITFLLTTYAK 352
+ + + MC LP +S T+G L+ +L LQYV + FL T Y+
Sbjct: 1409 EILRTFHNQTSIVMCSYLP--VFTSFNRTKGSLI-QLNHGRPQPLQYVVNAAFLATLYSD 1579
Query: 353 YMSAKKYT-FNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGS 411
Y+ A + CG + LR A+ Q+DYILG+NP +MSY+VG+G ++PK +HHR +
Sbjct: 1580 YLDAADTPGWYCGPNFFSTEKLREFARTQIDYILGKNPRKMSYVVGFGNHYPKHVHHRAA 1759
Query: 412 SLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTT 471
S+P + C GG++ + SS PNPNILVGA+V GP++ DGF D R +Y+++EPT
Sbjct: 1760 SIPK---NKVKYNCKGGWK-WRESSKPNPNILVGAMVAGPDRHDGFHDVRKNYNYTEPTL 1927
Query: 472 YINGAVVGPLAYFAG 486
N +V L +G
Sbjct: 1928 AGNAGLVAALVALSG 1972
>TC84823 weakly similar to GP|3549291|gb|AAC95009.1| endo-1 4-beta-glucanase
precursor {Fragaria x ananassa}, partial (31%)
Length = 955
Score = 240 bits (612), Expect(2) = 8e-76
Identities = 110/226 (48%), Positives = 158/226 (69%)
Frame = +2
Query: 199 KLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYY 258
++L+ A++V+ FA ++GSY++++GS CPFYC +G+ DEL+WGAAWL++A+N Y
Sbjct: 2 RILLARAKRVFDFANYHRGSYNNAIGSGACPFYCDINGYIDELIWGAAWLYKASNNQYYM 181
Query: 259 NLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPS 318
N +K+ Q + F WD K+AG +VL+S ++N N + A+N +C +LPNSP+
Sbjct: 182 NFVKSNIEYIQSNEFGWDSKHAGINVLVSHW-MMNDASNQSPFISNADNLICSLLPNSPT 358
Query: 319 SSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAK 378
S Y++GGL+FK SNLQ+VT+++FLL Y YM A NCGNV+ P L ++AK
Sbjct: 359 KSVTYSKGGLLFKAGPSNLQHVTALSFLLVVYGHYMEANNKIVNCGNVVAKPCDLINLAK 538
Query: 379 RQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIG 424
QVDY+LG NPL+MSYMVGYG +P++IHHRGS+LPSL HP+ G
Sbjct: 539 SQVDYVLGNNPLEMSYMVGYGQKYPQKIHHRGSTLPSLDVHPKHXG 676
Score = 62.4 bits (150), Expect(2) = 8e-76
Identities = 30/51 (58%), Positives = 34/51 (65%)
Frame = +3
Query: 424 GCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYIN 474
GC G +F SS PN N+L+GAIVGGP D F D R + S SEPTTYIN
Sbjct: 675 GCRDG-DKYFQSSTPNINVLIGAIVGGPANDDSFQDSRYNVSQSEPTTYIN 824
>TC88778 similar to GP|1039431|gb|AAC78504.1| cellulase {Phaseolus
vulgaris}, partial (49%)
Length = 781
Score = 244 bits (623), Expect = 5e-65
Identities = 122/254 (48%), Positives = 170/254 (66%), Gaps = 3/254 (1%)
Frame = +1
Query: 68 LANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDY 125
L V+L GGYYDAGDNVKF +PM+FT ++LSW+A+EY + Q+ R AIRWGT++
Sbjct: 37 LIKVNLVGGYYDAGDNVKFGWPMSFTVSLLSWAAVEYESEISSVNQIGYLRRAIRWGTNF 216
Query: 126 LLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAA 185
+L+ TS P L+ VG+ N DH CWERPEDMDT RT+Y + N+PG++ AAE AAAL+A
Sbjct: 217 ILQSHTS-PITLFTQVGEGNADHNCWERPEDMDTPRTLYKIDANSPGTEAAAEAAAALSA 393
Query: 186 ASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGA 245
AS+VFKK D YS L+R ++ ++ FA +Y+G+Y+ S CPFYCSYSG++DELLW A
Sbjct: 394 ASIVFKKKDTKYSSKLLRHSKSLFDFADKYRGTYTGS-----CPFYCSYSGYQDELLWAA 558
Query: 246 AWLFRATNAVKYYNLI-KTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQE 304
AWL++A+ KY I G + FSWD+K+ G LL++ G K+ + +
Sbjct: 559 AWLYKASGESKYLKYITDNQGWNQAASEFSWDNKFVGVQTLLTQE-FYGGKKDLAKIHSD 735
Query: 305 AENFMCKILPNSPS 318
E+F+C ++ S S
Sbjct: 736 GESFICALMQGSYS 777
>TC88779 similar to GP|1039431|gb|AAC78504.1| cellulase {Phaseolus
vulgaris}, partial (47%)
Length = 803
Score = 210 bits (535), Expect = 9e-55
Identities = 105/262 (40%), Positives = 152/262 (57%), Gaps = 32/262 (12%)
Frame = +3
Query: 228 CPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIK-TLGADDQPDIFSWDDKYAGAHVLL 286
CPFYCSYSG++DELLW AAWL++A+ KY I G + FSWD+K+ G LL
Sbjct: 15 CPFYCSYSGYQDELLWAAAWLYKASGESKYLKYITDNQGWNQAASEFSWDNKFVGVQTLL 194
Query: 287 SRRALLN------------------------------GDKNFDQYRQEAENFMCKILPNS 316
++ + G K+ + + E+F+C ++ S
Sbjct: 195 TQVFIFTHTYNKLMIFY*TLVVPKKELTQ*FY*EFYGGKKDLAKIHSDGESFICALMQGS 374
Query: 317 PSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYT-FNCGNVLVTPNTLRS 375
S + T GGL++ +NLQY T+ T +L +K ++ +CG+ T + +++
Sbjct: 375 YSLQIKKTPGGLLYTRDSNNLQYTTTSTMVLFIVSKILNKNNIDGIHCGSTNFTSSEIKA 554
Query: 376 VAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHS 435
AK QVDYILG NP++MSYMVGYG +PK++HHRGSS+PS+ H +GC+ G+ +F+S
Sbjct: 555 FAKSQVDYILGNNPMKMSYMVGYGSKYPKQLHHRGSSIPSIKVHQTKVGCNDGYTDYFYS 734
Query: 436 SNPNPNILVGAIVGGPNQSDGF 457
SNPNPNI VGAIVGGP+ +D F
Sbjct: 735 SNPNPNIHVGAIVGGPDFNDQF 800
>TC88503 similar to GP|1657380|emb|CAA65600.1 endo-beta-1 4-glucanase
{Prunus persica}, partial (37%)
Length = 1215
Score = 202 bits (515), Expect = 2e-52
Identities = 94/161 (58%), Positives = 119/161 (73%)
Frame = +2
Query: 326 GGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYIL 385
GG+++K +SNLQYVTS +FLL YAKY++ +CG +T L +AK+QVDYIL
Sbjct: 410 GGILYKGSESNLQYVTSTSFLLLIYAKYLNTNGGAVSCGTSKITEQNLIKLAKKQVDYIL 589
Query: 386 GENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVG 445
G+NP +MSYMVG+G +PK IHHRGSSLPS+ PQ I C+ GFQ + HS +PNPN+LVG
Sbjct: 590 GDNPTKMSYMVGFGEKYPKHIHHRGSSLPSIRVQPQQISCNNGFQ-YLHSGSPNPNVLVG 766
Query: 446 AIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAG 486
AIVGGP+ SD F DDR++Y SEP TYIN VG LA+F+G
Sbjct: 767 AIVGGPDSSDNFSDDRNNYQQSEPATYINAPFVGALAFFSG 889
Score = 33.1 bits (74), Expect = 0.25
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +2
Query: 301 YRQEAENFMCKILPNSPSSSTQYTQG 326
Y+ A+N++C ++P SP QYT G
Sbjct: 14 YKAHADNYICSLVPGSPGFQAQYTPG 91
>AJ388754 similar to GP|16648911|gb putative endo-1 4-beta-glucanase
{Arabidopsis thaliana}, partial (30%)
Length = 601
Score = 201 bits (512), Expect = 4e-52
Identities = 97/149 (65%), Positives = 113/149 (75%)
Frame = +3
Query: 336 NLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYM 395
NLQ+ TSI FL YAKY+S T NCGNV V+ TLR AKRQVDYILG+NPL +SYM
Sbjct: 3 NLQHATSIAFLELVYAKYLSRTSQTINCGNVYVSAQTLRERAKRQVDYILGDNPLGLSYM 182
Query: 396 VGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSD 455
VGYG N+P+RIHHRGSSLPS+ HPQ I C G +F+S+NPNPN+LVGA+VGGP + D
Sbjct: 183 VGYGNNYPQRIHHRGSSLPSIKDHPQQIACKEG-SIYFNSTNPNPNVLVGAVVGGPGEDD 359
Query: 456 GFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
+ DDR+DY SEPTTYIN VG LAYF
Sbjct: 360 VYVDDRADYRKSEPTTYINAPFVGGLAYF 446
>TC89276 weakly similar to GP|21740771|emb|CAD41248. OSJNBa0067K08.12 {Oryza
sativa}, partial (15%)
Length = 1389
Score = 121 bits (304), Expect(2) = 1e-50
Identities = 65/161 (40%), Positives = 97/161 (59%), Gaps = 3/161 (1%)
Frame = +1
Query: 324 TQGGLMFKLPDSN--LQYVTSITFLLTTYAKYMSAKKYT-FNCGNVLVTPNTLRSVAKRQ 380
T GGL+ PD+ LQY + +FL Y+ Y+ K + +C + + LR + Q
Sbjct: 682 TPGGLVIPKPDNGPLLQYAVTASFLSKLYSDYIDHLKISGASCETDTFSVSMLRDFSSSQ 861
Query: 381 VDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNP 440
V+YILG+NP++MSY+VGYG FP ++HHR +S+P + CD G + + +S NPNP
Sbjct: 862 VNYILGQNPMKMSYLVGYGDKFPVQVHHRSASIP---WDKRLYNCDDG-KTWLNSKNPNP 1029
Query: 441 NILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPL 481
+L+GA+VGGP+ +D F D RS+ +EPT N +V L
Sbjct: 1030QVLLGAMVGGPDTNDHFTDQRSNKRFTEPTISSNAGLVSAL 1152
Score = 97.1 bits (240), Expect(2) = 1e-50
Identities = 73/191 (38%), Positives = 99/191 (51%), Gaps = 23/191 (12%)
Frame = +2
Query: 120 RWGTDYLLKCATSTPGR---LYVGVG-------DPNVDHKCWERPEDMDTARTVYWVSPN 169
RWGTDYL K G LY VG +PN D CW+RPEDM R V +
Sbjct: 8 RWGTDYLHKVFIPPNGSNLTLYSQVGSTISTNHEPN-DISCWQRPEDMSYGRPVSVC--D 178
Query: 170 NPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQY----QGSYS--DSL 223
+D+A E AAL+AAS+VFK+ D YS LV+ A+ +Y+ + QG+Y+ D+
Sbjct: 179 GSATDLAGEIVAALSAASMVFKE-DKEYSGKLVQAAESLYEVVTKEDPKKQGTYTAVDAC 355
Query: 224 GSAVCPFYCSYSGFKDELLWGAAWLFRAT-------NAVKYYNLIKTLGADDQPDIFSWD 276
G Y S S +KDEL WGA WLF AT NA +++ K + +F W+
Sbjct: 356 GKQARMLYNSTS-YKDELAWGATWLFLATKNTDYLANATQFFLSAKKDETNLDKGVFYWN 532
Query: 277 DKYAGAHVLLS 287
+K + VLL+
Sbjct: 533 NKLSAVAVLLT 565
>AW256351 similar to GP|12957206|dbj beta-1 4-glucanase {Atriplex
lentiformis}, partial (17%)
Length = 262
Score = 123 bits (309), Expect = 1e-28
Identities = 61/87 (70%), Positives = 70/87 (80%)
Frame = +1
Query: 156 DMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQY 215
DMDT R VY VS NP SDVAAETAAALAA+SLVFK DP+YS L++ A +V+ FA +Y
Sbjct: 1 DMDTPRNVYKVSAQNPSSDVAAETAAALAASSLVFKDTDPSYSSKLLQAAIQVFNFADRY 180
Query: 216 QGSYSDSLGSAVCPFYCSYSGFKDELL 242
+GSYSDSL S VCPFYCSYSG+ DELL
Sbjct: 181 RGSYSDSLNSVVCPFYCSYSGYHDELL 261
>TC91867 weakly similar to GP|11094813|gb|AAG29742.1 endo-beta-1 4-glucanase
putative {Arabidopsis thaliana}, partial (33%)
Length = 963
Score = 121 bits (303), Expect = 7e-28
Identities = 53/115 (46%), Positives = 77/115 (66%)
Frame = +3
Query: 368 VTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDG 427
+ P L + K Q DYILG+NP MSY++GYGP +P +HHRGSS+ S+ + +GC
Sbjct: 15 IQPQELLNFVKSQADYILGKNPEDMSYLIGYGPKYPIHVHHRGSSIASVFSLHSEVGCAQ 194
Query: 428 GFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
GF +++ PNPN++ G +VGGP+++D + DDRS+Y SEPT + +VG LA
Sbjct: 195 GFDAWYNRIEPNPNVIHGGLVGGPDRNDDYTDDRSNYEQSEPTLSGSAPLVGILA 359
>BI311609 similar to PIR|T01108|T011 cellulase (EC 3.2.1.4) T21L14.7 -
Arabidopsis thaliana, partial (16%)
Length = 550
Score = 96.3 bits (238), Expect = 2e-20
Identities = 42/94 (44%), Positives = 61/94 (64%)
Frame = +3
Query: 394 YMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQ 453
Y+VGYGP++PKR+HHRG+S+ S + IGC G+ ++ +PNPN+LVGA+VGGP+
Sbjct: 3 YLVGYGPSYPKRVHHRGASIMSYKENKGFIGCTQGYDNWYSKEDPNPNVLVGALVGGPDW 182
Query: 454 SDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAGI 487
D F D R+++ +E TY +V A F I
Sbjct: 183 QDNFEDKRNNFMQTEACTYNTAPLVALFAKFLHI 284
>BG454971 similar to GP|1039431|gb|A cellulase {Phaseolus vulgaris}, partial
(14%)
Length = 474
Score = 95.1 bits (235), Expect = 5e-20
Identities = 43/74 (58%), Positives = 53/74 (71%)
Frame = +2
Query: 413 LPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTY 472
+PS+ H +GC+ G +F SSNPNPNI VGAIVGGPN +D + D RSDYSH+EPTTY
Sbjct: 2 IPSIKVHQTKVGCNDGQSNYFSSSNPNPNIHVGAIVGGPNSNDQYNDARSDYSHAEPTTY 181
Query: 473 INGAVVGPLAYFAG 486
+N A VG +A G
Sbjct: 182 MNAAFVGSVAALLG 223
>TC92360 similar to PIR|A96668|A96668 probable endo-beta-1 4-glucanase
F15H21.9 [imported] - Arabidopsis thaliana, partial
(22%)
Length = 519
Score = 58.5 bits (140), Expect = 5e-09
Identities = 38/126 (30%), Positives = 63/126 (49%), Gaps = 2/126 (1%)
Frame = +3
Query: 143 DPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAA--LAAASLVFKKVDPTYSKL 200
D + DH CW+RPEDM T T + + + K++ T +
Sbjct: 3 DGDTDHYCWQRPEDMTTF*TSF*DR*EESRFRSGWRNCCS*WQLLPLCLGKQIHITLNYF 182
Query: 201 LVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNL 260
+ +++F +Y+G Y S+G V +Y S SG+ DELL GA WL++AT+ +Y++
Sbjct: 183 CTMLST-LFEFGDKYRGKYDGSVG-VVKSYYASVSGYMDELLRGAMWLYKATDKQEYFDY 356
Query: 261 IKTLGA 266
I + G+
Sbjct: 357 IISNGS 374
>CA918779 similar to GP|6009979|dbj| endo-1 4-beta-glucanase {Pisum sativum},
partial (6%)
Length = 330
Score = 44.3 bits (103), Expect = 1e-04
Identities = 18/29 (62%), Positives = 22/29 (75%)
Frame = -1
Query: 457 FPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
FPD RSD+ SEP TY+N +VG LA+FA
Sbjct: 330 FPDKRSDFEQSEPATYVNAPLVGTLAFFA 244
>TC85996 weakly similar to GP|16648871|gb|AAL24287.1 Unknown protein
{Arabidopsis thaliana}, partial (53%)
Length = 1446
Score = 30.4 bits (67), Expect = 1.6
Identities = 33/126 (26%), Positives = 56/126 (44%), Gaps = 6/126 (4%)
Frame = +2
Query: 238 KDELLWGAAWLFRATNAV-KYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDK 296
KDE+L AW+ A ++ + +N I+TL D + + WDDK+ + +S + L
Sbjct: 305 KDEIL-SLAWMTLAMKSLCETHNDIRTLITDLELPVSVWDDKWVDVYFDISTKLL----- 466
Query: 297 NFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITF-----LLTTYA 351
+C I S ++ QG L K NL +S +F LL +
Sbjct: 467 -----------DICNIF---SSELSRLNQGNLPLKCALHNLGPASSKSFVRACSLLDDWR 604
Query: 352 KYMSAK 357
++++AK
Sbjct: 605 RHINAK 622
>TC79971 similar to GP|11994319|dbj|BAB02278. casein kinase {Arabidopsis
thaliana}, partial (21%)
Length = 869
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/53 (30%), Positives = 23/53 (43%)
Frame = +2
Query: 389 PLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPN 441
PL S++ NFP++IH S + HP + P FH + N N
Sbjct: 47 PLFFSFLFPESRNFPRKIHTLSSKTDPKSTHPYPLSTLFTILPQFHFIDSNLN 205
>TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regulatory
particle chain RPT6-like protein {Arabidopsis thaliana},
partial (77%)
Length = 1564
Score = 30.0 bits (66), Expect = 2.1
Identities = 14/35 (40%), Positives = 22/35 (62%)
Frame = -3
Query: 165 WVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSK 199
W SP++PG+DV AA+ VF++V+ +SK
Sbjct: 1241 WESPDSPGNDVLDVILLYSFAATDVFREVERAFSK 1137
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,320,046
Number of Sequences: 36976
Number of extensions: 247045
Number of successful extensions: 1104
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1063
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1073
length of query: 488
length of database: 9,014,727
effective HSP length: 100
effective length of query: 388
effective length of database: 5,317,127
effective search space: 2063045276
effective search space used: 2063045276
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0083.8


