

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0075.7
(777 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
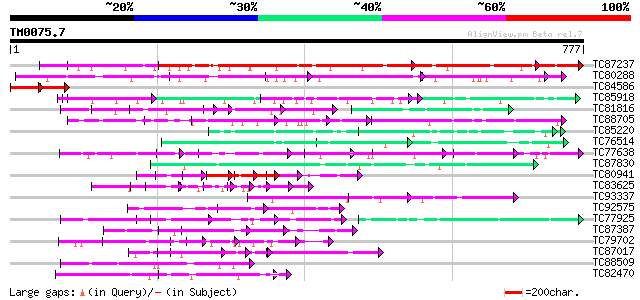
Score E
Sequences producing significant alignments: (bits) Value
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 584 e-167
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 128 7e-30
TC84586 similar to PIR|T04991|T04991 hypothetical protein T16L1.... 82 9e-29
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 103 2e-22
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 90 4e-18
TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unkn... 72 1e-12
TC85220 weakly similar to PIR|S33520|S33520 Lea protein - soybea... 71 1e-12
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 68 2e-11
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 64 3e-10
TC87830 similar to GP|15451168|gb|AAK96855.1 putative protein {A... 63 4e-10
TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein ... 62 6e-10
TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 62 1e-09
TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical... 61 1e-09
TC92575 weakly similar to GP|21428434|gb|AAM49877.1 LD13350p {Dr... 61 1e-09
TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1 ... 60 2e-09
TC87387 homologue to PIR|T47775|T47775 hypothetical protein F24I... 60 4e-09
TC79702 MtN14 59 7e-09
TC87017 similar to PIR|A96564|A96564 unknown protein 23094-2177... 59 7e-09
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 58 1e-08
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 58 1e-08
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 584 bits (1506), Expect = e-167
Identities = 361/615 (58%), Positives = 431/615 (69%), Gaps = 39/615 (6%)
Frame = +1
Query: 202 EENKH--EAENKEAAEVHEEKSQQENEETKDK-ENEEIKNEVEDTEAGEVKVEKSQQENE 258
EE H E+KE E EEKS ENEE+K+ E+ E K+ +E+ E + EKS+QENE
Sbjct: 1 EEKSHLENEESKETGESSEEKSHLENEESKETGESSEEKSHLENEENKDE--EKSKQENE 174
Query: 259 ETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKE-----NEEIKNEVEDKEAGEVK 313
E KD E + +NE E+K+ EEKSQQENEE KD+E NE KNE +KE GE+
Sbjct: 175 EIKDGEKIQQENE-ENKD-----EEKSQQENEENKDEEKSQQENELKKNEGGEKETGEIT 336
Query: 314 EEKSRQENEET-----KDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEA-EKEE 367
EEKS+QENEET KDKENEE NQNG+D K+ V ENHEQDSK+ EE+NGTE EKEE
Sbjct: 337 EEKSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNGTEGGEKEE 516
Query: 368 QGKIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAV-DHKSQDIFNESYNKTE-QVDK 425
KI+E+ SS+ QVQDGE NNE REENY+GDNASSAV D+KSQ ES NKTE Q DK
Sbjct: 517 HDKIKEDTSSDNQVQDGEKNNEA-REENYSGDNASSAVVDNKSQ----ESSNKTEEQFDK 681
Query: 426 NEKNESEMESQKSGTETNEVTDSNATTNIQENGGE-NQAQTDNGSEKRSTSESGGQ-QHQ 483
EKNE E+ESQK+ ET E TDS T N Q N E +QAQT+N + K S SES Q Q Q
Sbjct: 682 KEKNEFELESQKNSNETTESTDSTITQNSQGNESEKDQAQTENDTPKGSASESDEQKQEQ 861
Query: 484 EQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKTEDSNAD 543
EQNN T D V+T D+SSQNGN+TT +KQNETSEDANSK EDSS TTP EDS +
Sbjct: 862 EQNNTTKDDVQTTDTSSQNGNDTT----EKQNETSEDANSKKEDSSALNTTPNNEDSKSG 1029
Query: 544 AAGDDVDSTYKSSSGNS-DSNANNGAYKETTNLVQNENPNGNSVQEGKQD---------- 592
AGD DST +SS + D N N+G YK+TT NENP NS QEG Q+
Sbjct: 1030VAGDQADSTTTTSSSETQDGNTNHGEYKDTT----NENPEKNSGQEGTQESGSSSNTFDN 1197
Query: 593 ---ASHEAQL-TASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEAS 648
AS++ QL T SDTSSEQKKDESS+AE+ ++S QNDN NS SNT S+ SANDNK++S
Sbjct: 1198KDAASNKVQLTTTSDTSSEQKKDESSSAESKSESSQNDNANSGQSNTTSDESANDNKDSS 1377
Query: 649 QVNSSDVSSEESKEGSSNSENSSDANQNGSNSNE--NDNGNASHDA----NENENSVQRK 702
QV + SSE S EG+SN+EN+SD NQN S +NE ND+GN S+DA N+NEN+ Q K
Sbjct: 1378QVTT---SSENSAEGNSNTENNSDENQNDSKNNENTNDSGNTSNDANVNENQNENAAQTK 1548
Query: 703 TSENDGEAAQNESVESEKAKDESAHSDEGSNSNTNDQGSSSDPSITEEVKESHIDLGTLP 762
TSEN+G+ AQNESVES+K +ESAH D +NSN+NDQG SSD S+T++ KES +DLGTLP
Sbjct: 1549TSENEGD-AQNESVESKKENNESAHKDVDNNSNSNDQG-SSDTSVTQDDKESRVDLGTLP 1722
Query: 763 ESNAESNHRDVSSAE 777
ESN ES+H DVSSAE
Sbjct: 1723ESNGESHHNDVSSAE 1767
Score = 179 bits (455), Expect = 3e-45
Identities = 165/663 (24%), Positives = 288/663 (42%), Gaps = 22/663 (3%)
Frame = +1
Query: 79 HKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDENEVENKPEEH 138
H E +E E +EE+ +L+ S + E E+ H+ ENE ENK EE
Sbjct: 13 HLENEESKETGESSEEKSHLENEESKETGESSE--EKSHL---------ENE-ENKDEEK 156
Query: 139 IKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDNTESKENDSLV 198
K E E + + E+E++ ++ + +N E+K+ +
Sbjct: 157 SK-------------------QENEEIKDGEKIQQENEENKDEEKSQQENEENKDEEKSQ 279
Query: 199 KENEENKHEAENKEAAEVHEEKSQQENEET-----KDKENEEIKNEVEDT--EAGEVKVE 251
+ENE K+E KE E+ EEKS+QENEET KDKENEE D + GE +
Sbjct: 280 QENELKKNEGGEKETGEITEEKSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQ 459
Query: 252 KSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENE-EIKNEVEDKEAG 310
S+Q EET E E +E ++KE+ S + +K NE +N D +
Sbjct: 460 DSKQGTEETNGTEGG------EKEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASS 621
Query: 311 EVKEEKSRQENEETKDK-ENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQG 369
V + KS++ + +T+++ + +E N+ + + + E E + S G E+EK +Q
Sbjct: 622 AVVDNKSQESSNKTEEQFDKKEKNEFELESQKNSNETTESTDSTITQNSQGNESEK-DQA 798
Query: 370 KIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKN 429
+ + + + E EQ + D ++ ++ + E N+T + D N K
Sbjct: 799 QTENDTPKGSASESDEQKQEQEQNNTTKDDVQTTDTSSQNGNDTTEKQNETSE-DANSKK 975
Query: 430 ESEM--------ESQKSGTETNEVTDSNATTNIQENGGENQAQTDNGSEKRSTSE----S 477
E E KSG + DS TT+ E N T++G K +T+E +
Sbjct: 976 EDSSALNTTPNNEDSKSGV-AGDQADSTTTTSSSETQDGN---TNHGEYKDTTNENPEKN 1143
Query: 478 GGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKT 537
GQ+ +++ +S+ + D++S TT T+D +E +D E SS ++ + +
Sbjct: 1144SGQEGTQESGSSSNTFDNKDAASNKVQLTT--TSDTSSEQKKD-----ESSSAESKSESS 1302
Query: 538 EDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGNSVQEGKQDASHEA 597
++ NA++ S+ SD +AN+ + GNS E D
Sbjct: 1303QNDNANSG---------QSNTTSDESANDNKDSSQVTTSSENSAEGNSNTENNSD----- 1440
Query: 598 QLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVNSSDVSS 657
+ ++ K +E++N NT + N N N + + ++ S N+ D +
Sbjct: 1441-----ENQNDSKNNENTNDSGNTSNDANVNENQNENAAQTKTSENE---------GDAQN 1578
Query: 658 EESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKT-SENDGEAAQNESV 716
E + N+E++ N SNSN+ + + S ++ E+ V T E++GE+ N+
Sbjct: 1579ESVESKKENNESAHKDVDNNSNSNDQGSSDTSVTQDDKESRVDLGTLPESNGESHHNDVS 1758
Query: 717 ESE 719
+E
Sbjct: 1759SAE 1767
Score = 134 bits (338), Expect = 1e-31
Identities = 132/597 (22%), Positives = 259/597 (43%), Gaps = 86/597 (14%)
Frame = +1
Query: 41 NEKKASYGESAKTGNDVVRLGRKDLNPLVEETTVVDARHKEQDEESEHESKT-------- 92
NE+ GES++ + + K+ EE + ++ + +E+S+ E++
Sbjct: 22 NEESKETGESSEEKSHLENEESKETGESSEEKSHLENEENKDEEKSKQENEEIKDGEKIQ 201
Query: 93 -EEQINLDEVNSVDENEP----EHKPEEEHIKLDDANGVDENEV--ENKPEEHIKLDDAN 145
E + N DE S ENE E +E +K ++ + E+ E +E+ + + N
Sbjct: 202 QENEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKETGEITEEKSKQENEETSETN 381
Query: 146 GVDENSEHSQ----DLVEHEGEGSSENARTDIESEQHVEKVGEESDNTESKEN---DSLV 198
D+ +E S D E GE ++++ E E GE+ ++ + KE+ D+ V
Sbjct: 382 SKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNGTEG-GEKEEHDKIKEDTSSDNQV 558
Query: 199 KENEENKHEAENKEAAE-----VHEEKSQQENEETKDKENEEIKNEVE------------ 241
++ E+N E + + V + KSQ+ + +T+++ +++ KNE E
Sbjct: 559 QDGEKNNEAREENYSGDNASSAVVDNKSQESSNKTEEQFDKKEKNEFELESQKNSNETTE 738
Query: 242 --------DTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKE-----EKSQQE 288
+++ E + +++Q EN+ K +E + + E ++ K+ + S Q
Sbjct: 739 STDSTITQNSQGNESEKDQAQTENDTPKGSASESDEQKQEQEQNNTTKDDVQTTDTSSQN 918
Query: 289 NEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEE-------------TKDKENEEINQN 335
+T +K+NE ++ KE E+ + T E ++ N N
Sbjct: 919 GNDTTEKQNETSEDANSKKEDSSALNTTPNNEDSKSGVAGDQADSTTTTSSSETQDGNTN 1098
Query: 336 GTDVKDHVEENHEQDSKEEKEESNG-------------------TEAEKEEQGKIQEEAS 376
+ KD EN E++S +E + +G T ++ + K E +S
Sbjct: 1099HGEYKDTTNENPEKNSGQEGTQESGSSSNTFDNKDAASNKVQLTTTSDTSSEQKKDESSS 1278
Query: 377 SEKQVQDGEGNNEQHREENYAGDNASSAVDHK-SQDIFNESYNKTEQVDKNEKNESEMES 435
+E + + + +N + N D SA D+K S + S N E E N E ++
Sbjct: 1279AESKSESSQNDNANSGQSNTTSD--ESANDNKDSSQVTTSSENSAEGNSNTENNSDENQN 1452
Query: 436 -QKSGTETNEVTDSNATTNIQENGGENQAQTDNGSEKRSTSESGGQQHQEQNNPTSDGVE 494
K+ TN+ +++ N+ EN EN AQT SE +++ + +++NN ++ +
Sbjct: 1453DSKNNENTNDSGNTSNDANVNENQNENAAQTKT-SENEGDAQNESVESKKENNESAH--K 1623
Query: 495 TVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDS 551
VD++S + ++ +S T + T +D S+V+ T P +SN ++ +DV S
Sbjct: 1624DVDNNSNSNDQGSSDT----SVTQDDKESRVDLG----TLP---ESNGESHHNDVSS 1761
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 128 bits (322), Expect = 7e-30
Identities = 107/423 (25%), Positives = 190/423 (44%), Gaps = 20/423 (4%)
Frame = +3
Query: 8 RSQRSKGFKVKHALQICVLLGVCIWLVYQIKHTNEKKASYGES-----AKTGND-VVRLG 61
R+QRSKGFKVKH LQ +LLGVC WL+YQ+KH ++KK + ++ +T D +++LG
Sbjct: 171 RNQRSKGFKVKHVLQAILLLGVCFWLIYQVKHNHDKKKEFDKNDTKLPIRTETDQILKLG 350
Query: 62 RKDLNPLVEETTVVDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLD 121
RKDL+P E + H+E++E+ + + DE E +H+ EEE
Sbjct: 351 RKDLHPGKVEADKNEG-HEEEEEDEHIVYNMQNKREHDEQQQEGEEGNKHETEEE----- 512
Query: 122 DANGVDENEVENKPEEHIKLDDANG--VDENSEHSQDLVEHEGE----GSSENARTDIES 175
E+ V + EE + ++ +G V E +E + VE EG +++ +++ ++
Sbjct: 513 -----SEDNVHERREEQDEEENKHGAEVQEENESKSEEVEDEGGDVEIDENDHEKSEADN 677
Query: 176 EQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEE 235
++ E V EE D E ++++ ENE+ K + E E HE +E D +
Sbjct: 678 DREDEVVDEEKDKEEEGDDET---ENED-KEDEEKGGLVENHENHEAREEHYKADDASSA 845
Query: 236 IKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDK 295
+ ++ +T +E S + T+ ENE ++ E Q + K
Sbjct: 846 VAHDTHETSTETGNLEHSDLSLQNTRKPENETNHSD----------ESYGSQNVSDLKVT 995
Query: 296 ENEEIKNEVEDKEAG-EVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHE---QDS 351
E E + AG E + E +TK +++ N T V N D+
Sbjct: 996 EGELTDGVSSNATAGKETGNDSFSNETAKTKPDSQLDLSSNLTAVITEASSNSSGTGDDT 1175
Query: 352 KEEKEESNG---TEAEKEEQGKIQEEASSE-KQVQDGEGNNEQHREENYAGDNASSAVDH 407
E+ +E++ + + + + KQ + E + + E N G+++ +V
Sbjct: 1176SSSSEQIKAVILSESDHAQNATVNTTITGDMKQTEGLEQSGSKISEGNLPGNDSIVSVKP 1355
Query: 408 KSQ 410
+S+
Sbjct: 1356ESR 1364
Score = 99.8 bits (247), Expect = 4e-21
Identities = 89/372 (23%), Positives = 167/372 (43%), Gaps = 25/372 (6%)
Frame = +3
Query: 217 HEEKSQQENEETK----DKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKE---NEEIK 269
H++K + + +TK + ++ +K +D G+V+ +K++ EE +D+ N + K
Sbjct: 270 HDKKKEFDKNDTKLPIRTETDQILKLGRKDLHPGKVEADKNEGHEEEEEDEHIVYNMQNK 449
Query: 270 NEVEDKEAGEVKEEKSQQENEETKDK--ENEEIKNEVEDKEAGEVKEEKSRQENEETKDK 327
E D++ E +E + EE++D E E ++E E+K EV+EE +
Sbjct: 450 RE-HDEQQQEGEEGNKHETEEESEDNVHERREEQDEEENKHGAEVQEE---------NES 599
Query: 328 ENEEINQNGTDVKDHVEEN-HEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEG 386
++EE+ G DV+ ++EN HE+ + E + EK+++ + +E +E + + +G
Sbjct: 600 KSEEVEDEGGDVE--IDENDHEKSEADNDREDEVVDEEKDKEEEGDDETENEDKEDEEKG 773
Query: 387 ----NNEQH--REENYAGDNASSAVDHKSQDIFNESYN--------KTEQVDKNEKNESE 432
N+E H REE+Y D+ASSAV H + + E+ N + + +NE N S+
Sbjct: 774 GLVENHENHEAREEHYKADDASSAVAHDTHETSTETGNLEHSDLSLQNTRKPENETNHSD 953
Query: 433 MESQKSGTETNEVTDSNATTNIQENGGENQAQTDNGSEKRSTSESGGQQHQEQNNPTSDG 492
+VT+ T + N + ++ + Q +N T+
Sbjct: 954 ESYGSQNVSDLKVTEGELTDGVSSNATAGKETGNDSFSNETAKTKPDSQLDLSSNLTAVI 1133
Query: 493 VETVDSSSQNGNETTSHTTD-KQNETSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDS 551
E +SS G++T+S + K SE +++ +T T + + +G
Sbjct: 1134TEASSNSSGTGDDTSSSSEQIKAVILSESDHAQNATVNTTITGDMKQTEGLEQSGS---- 1301
Query: 552 TYKSSSGNSDSN 563
K S GN N
Sbjct: 1302--KISEGNLPGN 1331
Score = 62.0 bits (149), Expect = 8e-10
Identities = 77/414 (18%), Positives = 156/414 (37%), Gaps = 28/414 (6%)
Frame = +3
Query: 347 HEQDSKEEKEESNGTEAEKEEQGKI----QEEASSEKQVQDGEGNNEQHREENYAGDNAS 402
H D K+E ++++ + E +I +++ K D +E+ E+ + N
Sbjct: 264 HNHDKKKEFDKNDTKLPIRTETDQILKLGRKDLHPGKVEADKNEGHEEEEEDEHIVYNMQ 443
Query: 403 SAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGENQ 462
+ +H Q ++ ++ K+E+E ES+ + E E D +EN +
Sbjct: 444 NKREHDEQQ---------QEGEEGNKHETEEESEDNVHERREEQDE------EENKHGAE 578
Query: 463 AQTDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDAN 522
Q +N S+ + GG ++N+ +E + D+++E ++
Sbjct: 579 VQEENESKSEEVEDEGGDVEIDEND----------------HEKSEADNDREDEVVDEEK 710
Query: 523 SKVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPN 582
K E+ +T + ED + G V++ + A++ + + +
Sbjct: 711 DKEEEGDDET---ENEDKEDEEKGGLVENHENHEAREEHYKADDASSAVAHDTHETSTET 881
Query: 583 GNSVQEGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNS---EG 639
GN + + Q T + DES ++ +D + +DG ++N+ +
Sbjct: 882 GNL-----EHSDLSLQNTRKPENETNHSDESYGSQNVSDLKVTEGELTDGVSSNATAGKE 1046
Query: 640 SANDN--------KEASQVNSSDVSSEESKEGSSNSENSSDANQNGSN------------ 679
+ ND+ K SQ++ S + E SSNS + D + S
Sbjct: 1047TGNDSFSNETAKTKPDSQLDLSSNLTAVITEASSNSSGTGDDTSSSSEQIKAVILSESDH 1226
Query: 680 -SNENDNGNASHDANENENSVQRKTSENDGEAAQNESVESEKAKDESAHSDEGS 732
N N + D + E Q + ++G N+S+ S K + A +E S
Sbjct: 1227AQNATVNTTITGDMKQTEGLEQSGSKISEGNLPGNDSIVSVKPESRVAAPEESS 1388
Score = 56.6 bits (135), Expect = 4e-08
Identities = 47/216 (21%), Positives = 89/216 (40%), Gaps = 21/216 (9%)
Frame = +3
Query: 560 SDSNANNGAYKETTNLV---QNENPNGNSVQEGKQDASHEAQLTASDTSSEQKKDESSNA 616
+D N + +E ++V QN+ + QEG++ HE + + D E+++++
Sbjct: 381 ADKNEGHEEEEEDEHIVYNMQNKREHDEQQQEGEEGNKHETEEESEDNVHERREEQDEEE 560
Query: 617 ETNTDSVQNDN------VNSDGSN-----TNSEGSANDNKEASQVNSSDVSSEESKEGSS 665
+ VQ +N V +G + + E S DN +V + EE + +
Sbjct: 561 NKHGAEVQEENESKSEEVEDEGGDVEIDENDHEKSEADNDREDEVVDEEKDKEEEGDDET 740
Query: 666 NSENSSDANQNGSNSN-ENDNGNASH-DANENENSVQRKTSENDGEAAQNESVE-----S 718
+E+ D + G N EN H A++ ++V T E E E + +
Sbjct: 741 ENEDKEDEEKGGLVENHENHEAREEHYKADDASSAVAHDTHETSTETGNLEHSDLSLQNT 920
Query: 719 EKAKDESAHSDEGSNSNTNDQGSSSDPSITEEVKES 754
K ++E+ HSDE S ++ +T+ V +
Sbjct: 921 RKPENETNHSDESYGSQNVSDLKVTEGELTDGVSSN 1028
>TC84586 similar to PIR|T04991|T04991 hypothetical protein T16L1.230 -
Arabidopsis thaliana, partial (18%)
Length = 499
Score = 81.6 bits (200), Expect(2) = 9e-29
Identities = 39/46 (84%), Positives = 43/46 (92%), Gaps = 1/46 (2%)
Frame = +3
Query: 1 MFRLSPRRSQRSK-GFKVKHALQICVLLGVCIWLVYQIKHTNEKKA 45
MFRLSPRRSQRSK GFKVKHALQICVL+GVC+WLVYQI H+ E+KA
Sbjct: 252 MFRLSPRRSQRSKNGFKVKHALQICVLVGVCVWLVYQIGHSREQKA 389
Score = 64.3 bits (155), Expect(2) = 9e-29
Identities = 29/39 (74%), Positives = 34/39 (86%)
Frame = +1
Query: 43 KKASYGESAKTGNDVVRLGRKDLNPLVEETTVVDARHKE 81
KK SYG S KTGN+VV+LGRKDL P VEET+V+DARH+E
Sbjct: 382 KKLSYGASTKTGNEVVKLGRKDLKPRVEETSVIDARHRE 498
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 103 bits (258), Expect = 2e-22
Identities = 168/752 (22%), Positives = 297/752 (39%), Gaps = 57/752 (7%)
Frame = -1
Query: 79 HKEQDEESEHESKTEEQINLDEVN-SVDENEPEHKPEEEHIKLDDANGVDENEVENKP-- 135
HK Q + E E K E + ++ + SV P+EE + ++ + E N
Sbjct: 3040 HKIQV*KKEDEVKAEPAVAAEKTDDSVPIETVSDNPKEEKDLITESETISVTEPVNDTAI 2861
Query: 136 EEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDNTESKEND 195
H D+ + +++ + +E E ++E + IE E + SD TE+ + +
Sbjct: 2860 SHH---DETPSTESDAKETSQQIEKELVETNEENQPKIEDIPEAETTEKSSDATETNQFE 2690
Query: 196 SLVKENEENKHEAENKEAAEVHEEKSQQENEETKD--KENEEIKNEVEDTEAGEVKVEKS 253
VK+ E ++ AE +E E+ E + +E + TK+ KE++E E E +A + VE +
Sbjct: 2689 EKVKQAEISETSAEKEEKPEL-EPLATEEPKVTKEPEKESQEKSEEAEQPKAIAI-VEST 2516
Query: 254 QQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVK 313
+ NEE E +E + VKEE E + + E + EVE E +K
Sbjct: 2515 VEANEEKTAAAISEETKIIEVEPTETVKEEPMVTEVATAEIVKEEPVVTEVEATET--LK 2342
Query: 314 EEKSRQENEETKDKENE----EINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQG 369
EE E E T+ + E E+ T VKD ++ KEE TEAE E
Sbjct: 2341 EELVATEVEATETVKEEPAATEVKVTET-VKDEPVVTEVDPTETVKEEPLATEAEATETV 2165
Query: 370 K---IQEEASSEKQVQDGEGNNEQHREENYAGDNASSAVDH----KSQDIFNESYNKTEQ 422
K + E + V++ E E + A++ V+ K + + E+ TE
Sbjct: 2164 KEEPVATEVEPTETVKEEPVATEVEATETVKDEPAATEVEATETVKEEPVATET-EATET 1988
Query: 423 VDKNEKNESEMESQKSGTETNEVTDSNATTNIQE-------NGGENQAQTDNGSEKRSTS 475
V K E +E+E+ ++ E + T+ AT ++E E +E +T
Sbjct: 1987 V-KEEPVATEVEATETVKEESVATEVEATETVKEEPVTTVVEPTEMVKDEPAATEFEATE 1811
Query: 476 ESGGQQHQEQNNPTS-----------DGVETVDSSSQNGNETTSHTTDKQNETSEDANSK 524
+ + +PT + +TV + T K T D
Sbjct: 1810 TVKDETIVTEVDPTETVKEEAVATEVEITKTVKEPLVTEVDLTESVKVKPVVTEVDPREM 1631
Query: 525 VEDSSTDTTTPKTEDSNADAAGDDVD--STYKSSSGNSDSNANNGAYKE-TTNLVQNENP 581
V++ T TE + +V+ T K ++ N ++ +T + E P
Sbjct: 1630 VKEEPVATEVQATETVKEEPVATEVEPTETVKGEPVVTEVEENQKEPEQHSTERREEEQP 1451
Query: 582 NGNSVQE---------GKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDG 632
N +S+ E ++ S E + A+ E K DE+ AET + + V ++
Sbjct: 1450 NASSIPEQSTETNDVIAVEEKSRELEFEAA-ILKETKNDEAGPAETE----KVEPVVTEV 1286
Query: 633 SNTNSEGSANDNKEASQVNSSDVSSEESKEG---------SSNSENSSDANQNGSNS--N 681
SE K+ +V + + + + G S+ E + N +GS +
Sbjct: 1285 DENQSEPGIQSFKQEEEVKPKEETENQEQTGDRVEVHPPKDSDIEAVKETNNSGSEAILV 1106
Query: 682 ENDNGNASHDANENENSVQRKTSENDGEAAQNESVESEKAKDESAHSDEGSNSNTNDQGS 741
+ +N + + E + S Q + E GE + VE E+A +++ EGSN ++
Sbjct: 1105 KEENIDPLSNGVEEKLSEQLQVGEEVGEII--KEVEPEQATEKT----EGSNGTIKEE-- 950
Query: 742 SSDPSITEEVKESHIDLGTLPESNAESNHRDV 773
++P+ T + L E + E + V
Sbjct: 949 ETEPNTTVNAAQVQTTPANLVEPSPEVEEKIV 854
Score = 79.3 bits (194), Expect = 5e-15
Identities = 118/523 (22%), Positives = 212/523 (39%), Gaps = 47/523 (8%)
Frame = -1
Query: 72 TTVVDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDENEV 131
TTVV+ +DE + E + E + + + V E +P +EE + E E+
Sbjct: 1879 TTVVEPTEMVKDEPAATEFEATETVKDETI--VTEVDPTETVKEEAV-------ATEVEI 1727
Query: 132 ENKPEEHI--KLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHV--EKVGEESD 187
+E + ++D V ++ V+ E T++++ + V E V E +
Sbjct: 1726 TKTVKEPLVTEVDLTESVKVKPVVTE--VDPREMVKEEPVATEVQATETVKEEPVATEVE 1553
Query: 188 NTESKENDSLVKENEENK-----HEAENKEAAEVHEEKSQQENEETKD------KENE-- 234
TE+ + + +V E EEN+ H E +E + + +++ ET D K E
Sbjct: 1552 PTETVKGEPVVTEVEENQKEPEQHSTERREEEQPNASSIPEQSTETNDVIAVEEKSRELE 1373
Query: 235 ---EIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQEN-- 289
I E ++ EAG + EK + E + ++E + +E + KEE QE
Sbjct: 1372 FEAAILKETKNDEAGPAETEKVEPVVTEVDENQSEPGIQSFKQEEEVKPKEETENQEQTG 1193
Query: 290 -----EETKDKENEEIK-NEVEDKEAGEVKEEKSRQENEETKDKENEEINQNGTDVKDHV 343
KD + E +K EA VKEE + ++K +E++ Q G +V + +
Sbjct: 1192 DRVEVHPPKDSDIEAVKETNNSGSEAILVKEENIDPLSNGVEEKLSEQL-QVGEEVGEII 1016
Query: 344 EENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGN----NEQHREENYAGD 399
+E + + E+ E SNGT E+E + A+ QVQ N + + E+ D
Sbjct: 1015 KEVEPEQATEKTEGSNGTIKEEETEPNTTVNAA---QVQTTPANLVEPSPEVEEKIVVED 845
Query: 400 NASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGG 459
V + + QV E+NE++ + ++G +V + + N E
Sbjct: 844 GKKEVVVAAEDEKKEPEASDAVQVSSREQNEAKTATIEAGETETKVEEISKAVN--EPVR 671
Query: 460 ENQAQTDNGSEKRSTSESGGQQHQEQ-NNPTSDGVETVDSSSQNGNETTSHTTDKQNET- 517
E A E+ + +E+ P VE S N+TT+ + D ET
Sbjct: 670 ETLASKFEKDEEETIQNGVDNLEKERIEEPVKTEVE-----STKENDTTTISKDLPKETP 506
Query: 518 -------SEDANSKVEDS------STDTTTPKTEDSNADAAGD 547
S + +KV+ S + +P +++ ++D GD
Sbjct: 505 AKPAQKQSNNIIAKVKQSLVKAKKAITGKSPSSKNLSSDPKGD 377
Score = 45.4 bits (106), Expect = 8e-05
Identities = 52/230 (22%), Positives = 97/230 (41%), Gaps = 11/230 (4%)
Frame = -3
Query: 341 DHVEENHEQDSKEEKEE----------SNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQ 390
DH HEQ K+EK+E + + ++ Q +A+ +++D E E
Sbjct: 3731 DHASTTHEQFVKQEKDEAIAEKMNSLANEDGDKHNDKPDNAQGQATPSSKIED-EVKAEL 3555
Query: 391 HREENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNA 450
EE D+ SSA+ +DI K E + + + + E K + + T S A
Sbjct: 3554 AAEEIEKSDD-SSALKVSVEDIV--KLEKDEAIAEKMNSLANEEGDKHDDKPDN-TQSQA 3387
Query: 451 TTNIQENGGENQAQTDNGSEKRSTSESGGQQHQEQ-NNPTSDGVETVDSSSQNGNETTSH 509
T + G E + + G +RS S + E NP D ++ ++ N +
Sbjct: 3386 TPSSNTEGDEVKTELAAGEIERSDDSSALKDFVEDIVNPEKD-----EAIAEKINSLANE 3222
Query: 510 TTDKQNETSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSGN 559
DK + +DA + +T ++T + ++ A+ A +++ + SS+ N
Sbjct: 3221 DGDKHDGKPDDA----QGQATPSSTTEGDEIKAELAAAEIEKSDDSSALN 3084
Score = 42.4 bits (98), Expect = 7e-04
Identities = 40/146 (27%), Positives = 66/146 (44%), Gaps = 12/146 (8%)
Frame = -1
Query: 66 NPLVEETTVVDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEH---KPEEEHIKLDD 122
+P VEE VV+ KE +E E K E + +V+S ++NE + + E K+++
Sbjct: 880 SPEVEEKIVVEDGKKEVVVAAEDEKKEPEASDAVQVSSREQNEAKTATIEAGETETKVEE 701
Query: 123 ANGVDENEVENKPEEHIKLDD----ANGVDE-NSEHSQDLVEHEGEGSSENART----DI 173
+ V + D+ NGVD E ++ V+ E E + EN T D+
Sbjct: 700 ISKAVNEPVRETLASKFEKDEEETIQNGVDNLEKERIEEPVKTEVESTKENDTTTISKDL 521
Query: 174 ESEQHVEKVGEESDNTESKENDSLVK 199
E + ++S+N +K SLVK
Sbjct: 520 PKETPAKPAQKQSNNIIAKVKQSLVK 443
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 89.7 bits (221), Expect = 4e-18
Identities = 60/216 (27%), Positives = 117/216 (53%), Gaps = 4/216 (1%)
Frame = +3
Query: 163 EGSSENARTDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENK---EAAEVHEE 219
E SS TDI +E+ N K +D + +E+K E E ++ E +E
Sbjct: 18 EASSLFTTTDITVSSKMEE-----SNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDE 182
Query: 220 KSQQENEETKDKEN-EEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAG 278
K +++++E KDK + +E K++ + + + K E++ + EE D++ ++ K + E KE G
Sbjct: 183 KKEKKDKEKKDKTDVDEGKDKKDKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKG 362
Query: 279 EVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKENEEINQNGTD 338
+ ++K +E + KDKE ++ KN ED + GE +K + ++++ K KE +E +
Sbjct: 363 KEDKDKDGEEKKSKKDKEKKKDKN--EDDDEGEDGSKKKKNKDKKEKKKEEDEKEEGKVS 536
Query: 339 VKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEE 374
V+D E ++ KE+K++ E +K++ GK +++
Sbjct: 537 VRDIDIEETAKEGKEKKKKKEDKEEKKKKSGKDKDQ 644
Score = 77.0 bits (188), Expect = 3e-14
Identities = 55/195 (28%), Positives = 95/195 (48%), Gaps = 14/195 (7%)
Frame = +3
Query: 264 ENEEIKNEVEDKEAGEVKEEKSQQENE-ETKDKENEEIKNEVEDKEAGEVKEEKSRQENE 322
E I +++DK AG+VKE+K + E + E K E ++ K E +DKE K++ E +
Sbjct: 69 EESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKEK---KDKTDVDEGK 239
Query: 323 ETKDKENEEINQNGTDVKDHVEENHEQDSKE----EKEESNGTEAEK---EEQGKIQEEA 375
+ KDKE ++ + +VK E+ E+ KE EK+E + +K E++ K +E
Sbjct: 240 DKKDKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEK 419
Query: 376 SSEKQVQDGEGNNEQHREENYAGDNASSAVDHKSQ------DIFNESYNKTEQVDKNEKN 429
+K D EG + +++N D K + DI E K + K +K
Sbjct: 420 KKDKNEDDDEGEDGSKKKKNKDKKEKKKEEDEKEEGKVSVRDIDIEETAKEGKEKKKKKE 599
Query: 430 ESEMESQKSGTETNE 444
+ E + +KSG + ++
Sbjct: 600 DKEEKKKKSGKDKDQ 644
Score = 72.8 bits (177), Expect = 5e-13
Identities = 57/200 (28%), Positives = 106/200 (52%), Gaps = 8/200 (4%)
Frame = +3
Query: 112 KPEEEHI--KLDD--ANGVDENEVENKPEEHIKL---DDANGVDENSEHSQDLVEHEGEG 164
K EE +I K+DD A V E++VE + + IK DD ++ E EG+
Sbjct: 63 KMEESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKEKKDKTDVDEGKD 242
Query: 165 SSENARTDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQE 224
+ + E ++ K GEE D E K+ + KE +E E ++K+ E+KS+++
Sbjct: 243 KKDKEKKKKEKKEENVK-GEEEDGDEKKDKEKKKKEKKEKGKEDKDKDG---EEKKSKKD 410
Query: 225 NEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKE-AGEVKEE 283
E+ KDK ++ +E ED + +K +++ EE + +E + +++ +E A E KE+
Sbjct: 411 KEKKKDKNEDD--DEGEDGSKKKKNKDKKEKKKEEDEKEEGKVSVRDIDIEETAKEGKEK 584
Query: 284 KSQQENEETKDKENEEIKNE 303
K ++E++E K K++ + K++
Sbjct: 585 KKKKEDKEEKKKKSGKDKDQ 644
Score = 62.8 bits (151), Expect = 5e-10
Identities = 47/216 (21%), Positives = 110/216 (50%), Gaps = 1/216 (0%)
Frame = +3
Query: 69 VEETTV-VDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVD 127
+EE+ + + K + E + + E+ + + V DE + E K +E+ K D G D
Sbjct: 66 MEESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKK-EKKDKEKKDKTDVDEGKD 242
Query: 128 ENEVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESD 187
+ + E K +E K ++ G +E+ + +D + + E + ++ +K GEE
Sbjct: 243 KKDKEKKKKEK-KEENVKGEEEDGDEKKDKEKKKKEKKEKG-------KEDKDKDGEEKK 398
Query: 188 NTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGE 247
+ + KE K+ ++N+ + E ++ ++ +++N++ K+K+ EE + E +
Sbjct: 399 SKKDKE-----KKKDKNEDDDEGEDGSK------KKKNKDKKEKKKEEDEKEEGKVSVRD 545
Query: 248 VKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEE 283
+ +E++ +E +E K K+ ++ E + K++G+ K++
Sbjct: 546 IDIEETAKEGKEKKKKKEDK---EEKKKKSGKDKDQ 644
Score = 43.5 bits (101), Expect = 3e-04
Identities = 43/221 (19%), Positives = 79/221 (35%), Gaps = 1/221 (0%)
Frame = +3
Query: 464 QTDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANS 523
QT+ S +T + + +E N P ++ ++ E K E ++
Sbjct: 12 QTEASSLFTTTDITVSSKMEESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKE 191
Query: 524 KVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNG 583
K + D T + D D D K ++ G ++ E
Sbjct: 192 KKDKEKKDKT-------DVDEGKDKKDKEKKKKEKKEENV--KGEEEDGDEKKDKEKKKK 344
Query: 584 NSVQEGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSAND 643
++GK+D + + S E+KKD++ + + D + N D E D
Sbjct: 345 EKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDEGEDGSKKKK-NKDKKEKKKE---ED 512
Query: 644 NKEASQVNSSDVSSEE-SKEGSSNSENSSDANQNGSNSNEN 683
KE +V+ D+ EE +KEG + D + S ++
Sbjct: 513 EKEEGKVSVRDIDIEETAKEGKEKKKKKEDKEEKKKKSGKD 635
Score = 34.3 bits (77), Expect = 0.19
Identities = 31/168 (18%), Positives = 73/168 (43%), Gaps = 1/168 (0%)
Frame = +3
Query: 34 VYQIKHTNEKKASYGESAKTGNDVVR-LGRKDLNPLVEETTVVDARHKEQDEESEHESKT 92
+ ++ +EKK + K DV +KD +E + + +E+D + + + +
Sbjct: 156 IKSVEKDDEKKEKKDKEKKDKTDVDEGKDKKDKEKKKKEKKEENVKGEEEDGDEKKDKEK 335
Query: 93 EEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDENEVENKPEEHIKLDDANGVDENSE 152
+++ E + + + EE+ K D D+NE +++ E+ K D+ +
Sbjct: 336 KKK----EKKEKGKEDKDKDGEEKKSKKDKEKKKDKNEDDDEGEDGSK--KKKNKDKKEK 497
Query: 153 HSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDNTESKENDSLVKE 200
++ + EG+ S + + +++ EK ++ D E K+ K+
Sbjct: 498 KKEEDEKEEGKVSVRDIDIEETAKEGKEKKKKKEDKEEKKKKSGKDKD 641
>TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1685
Score = 71.6 bits (174), Expect = 1e-12
Identities = 67/294 (22%), Positives = 130/294 (43%), Gaps = 6/294 (2%)
Frame = +1
Query: 79 HKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDENEVENKPEEH 138
HKEQ +++E + + ++ E + E E K E+ +K ++ ++ +EN+ E+H
Sbjct: 31 HKEQIDKAEEKERLQK-----EKEEKQKKEAEEKANEKQVKTNE----EDTGIENEAEKH 183
Query: 139 IKLDD--ANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDNTESKENDS 196
++D A + E E +D + E + A T +E + +D +ES+ S
Sbjct: 184 SDIEDNFAASIHEKIEVKEDSPVDQDEAGEKLADT-------LENFDKATDTSESE--GS 336
Query: 197 LVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQE 256
L + EEN EAE + E E + T K TE+ + ++ ++
Sbjct: 337 LFDKVEENAKEAEREPTVE-------SETDLTTGK-----------TESSDEAIDTGKEA 462
Query: 257 NEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEI----KNEVEDKEAGEV 312
+E T EE+ V + GE +KS + N +++ E+I NE + A E
Sbjct: 463 SENTDGLSKEELGRLVASRWTGEDVGKKSVEANTALDNEDQEDILHGTNNEENEGYASET 642
Query: 313 KEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKE 366
++ S+ +++ K +++ + D+ D E ++D E+ S ++ E E
Sbjct: 643 DDDTSKYDDD--TGKYDDDTGKYDEDIND---EEFQEDEHEDLSSSYKSDVESE 789
Score = 65.5 bits (158), Expect = 8e-11
Identities = 71/272 (26%), Positives = 114/272 (41%), Gaps = 19/272 (6%)
Frame = +1
Query: 184 EESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENE-EIKNEVED 242
E+ D E KE L KE EE + KEA E EK + NEE ENE E +++ED
Sbjct: 37 EQIDKAEEKER--LQKEKEEK----QKKEAEEKANEKQVKTNEEDTGIENEAEKHSDIED 198
Query: 243 TEAG------EVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKE 296
A EVK E S + +E +K + ++N + + E + + E K+ E
Sbjct: 199 NFAASIHEKIEVK-EDSPVDQDEAGEKLADTLENFDKATDTSESEGSLFDKVEENAKEAE 375
Query: 297 NEEIKNEVEDKEAGEVKE-----EKSRQENEETKDKENEEINQ------NGTDV-KDHVE 344
E D G+ + + ++ +E T EE+ + G DV K VE
Sbjct: 376 REPTVESETDLTTGKTESSDEAIDTGKEASENTDGLSKEELGRLVASRWTGEDVGKKSVE 555
Query: 345 ENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQHREENYAGDNASSA 404
N D++++++ +GT E E +G E + D G Y D
Sbjct: 556 ANTALDNEDQEDILHGTNNE-ENEGYASETDDDTSKYDDDTG--------KYDDDTGK-- 702
Query: 405 VDHKSQDIFNESYNKTEQVDKNEKNESEMESQ 436
+DI +E + + E D + +S++ES+
Sbjct: 703 ---YDEDINDEEFQEDEHEDLSSSYKSDVESE 789
Score = 57.4 bits (137), Expect = 2e-08
Identities = 59/270 (21%), Positives = 124/270 (45%), Gaps = 26/270 (9%)
Frame = +1
Query: 247 EVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENE-EIKNEVE 305
++K K Q + E K++ +E K E + KEA E EK + NEE ENE E +++E
Sbjct: 19 QLKDHKEQIDKAEEKERLQKE-KEEKQKKEAEEKANEKQVKTNEEDTGIENEAEKHSDIE 195
Query: 306 DKEAGEVKEE-KSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAE 364
D A + E+ + ++++ +D+ E++ + + + + S +K E N EAE
Sbjct: 196 DNFAASIHEKIEVKEDSPVDQDEAGEKLADTLENFDKATDTSESEGSLFDKVEENAKEAE 375
Query: 365 KEEQGKIQEE------ASSEKQVQDGEGNNEQ----HREE-------NYAGDN------- 400
+E + + + SS++ + G+ +E +EE + G++
Sbjct: 376 REPTVESETDLTTGKTESSDEAIDTGKEASENTDGLSKEELGRLVASRWTGEDVGKKSVE 555
Query: 401 ASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGE 460
A++A+D++ Q+ N E ++ +E++ ++ K +T + D T E+ +
Sbjct: 556 ANTALDNEDQEDILHGTNNEE--NEGYASETDDDTSKYDDDTGKYDDD--TGKYDEDIND 723
Query: 461 NQAQTDNGSEKRSTSESGGQQHQEQNNPTS 490
+ Q D + S+ +S + + ++ S
Sbjct: 724 EEFQEDEHEDLSSSYKSDVESEPDLSDDPS 813
Score = 50.1 bits (118), Expect = 3e-06
Identities = 55/279 (19%), Positives = 116/279 (40%), Gaps = 15/279 (5%)
Frame = +1
Query: 491 DGVETVDSSSQNGNETTSHTTDKQNETSEDANSK-VEDSSTDTTTPKTEDSNADAAGDDV 549
D E +D + + ++ E E AN K V+ + DT + ++D +
Sbjct: 28 DHKEQIDKAEEKERLQKEKEEKQKKEAEEKANEKQVKTNEEDTGIENEAEKHSDIEDNFA 207
Query: 550 DSTYKSSSGNSDSNANNGA-----------YKETTNLVQNENPNGNSVQEGKQDASHEAQ 598
S ++ DS + + + T+ ++E + V+E ++A E
Sbjct: 208 ASIHEKIEVKEDSPVDQDEAGEKLADTLENFDKATDTSESEGSLFDKVEENAKEAEREPT 387
Query: 599 LTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVNSSDVSSE 658
+ S+T K ESS+ +T ++N +DG + G AS+ DV +
Sbjct: 388 VE-SETDLTTGKTESSDEAIDTGKEASEN--TDGLSKEELGRL----VASRWTGEDVGKK 546
Query: 659 ESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSENDGEAAQNESVES 718
+ ++ + +G+N+ EN+ G AS + +++ + T + D + + + +
Sbjct: 547 SVEANTALDNEDQEDILHGTNNEENE-GYAS-ETDDDTSKYDDDTGKYDDDTGKYDEDIN 720
Query: 719 EKAKDESAHSDEGSNSNTNDQGS---SSDPSITEEVKES 754
++ E H D S+ ++ + S DPS E++++S
Sbjct: 721 DEEFQEDEHEDLSSSYKSDVESEPDLSDDPSWLEKIQKS 837
>TC85220 weakly similar to PIR|S33520|S33520 Lea protein - soybean, partial
(69%)
Length = 1880
Score = 71.2 bits (173), Expect = 1e-12
Identities = 103/493 (20%), Positives = 191/493 (37%), Gaps = 9/493 (1%)
Frame = +1
Query: 270 NEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKEN 329
NE E ++ +VK + ET +K +E K E KEA E E ++++ E ++
Sbjct: 160 NEDEGRDFQDVKAKTM-----ETANKASETGK---EGKEAAETWTEWAKEKITEGLGFKD 315
Query: 330 EEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNE 389
++ + TD + +K +E+ KE K + A + G +
Sbjct: 316 KDKDSTTTDKASEYASDAAHKTKHYADET--AHKSKETAQKASDHAHDAAEKTKGYAWDA 489
Query: 390 QHREENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSN 449
+ +YA D AS H + D ESY + ++ ++ + K+G E T S
Sbjct: 490 SKKIRDYATD-ASEKTKHYAAD---ESYADDDDDAEDNRDFAGEAIHKAG----EYTGSA 645
Query: 450 ATTNIQENGGENQAQTDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSH 509
A + E + +++ + G H+ + E ++Q E
Sbjct: 646 A-----QKAKEYATDAAQKAAQKAKEYATGAAHKSK--------EYAGDAAQKTKEYAGD 786
Query: 510 TTDKQNETSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSS--SGNSDSNANNG 567
T+K E + DA K +D + T E + D+ YKS +G++ A +
Sbjct: 787 ATEKTKEYAGDAAQKTKDYAGGATQKSKEYAT--------DAAYKSKEYAGDAAQMAKDA 942
Query: 568 AYKETTNLVQNENPNGNSVQEGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDN 627
AYK ++ G++ ++ K S AQ A D + + K AE D+
Sbjct: 943 AYK-------TKDYAGDAAEKTKDYGSDAAQ-KAKDAAYKTKDYAGDAAEKTKDAAYKTK 1098
Query: 628 VNSDGSNTNSEGSANDNKEASQVNSSDVSSEESKEGSSNSENSSDANQNGSNSNENDNGN 687
+ + + AN A+ S D + + +++ + + D + ++ G+
Sbjct: 1099DYAGDATEKTREYAN----AAAYKSKDYAGDAAEKTKDAAYKTKDYAGEAAQKAKDYAGD 1266
Query: 688 ASHDANENENSVQRKTSENDGEAAQNESVESEKAKDES-------AHSDEGSNSNTNDQG 740
A+H+ E N K+ + G+AAQ SE A D + A + + + + D
Sbjct: 1267AAHNTREYANDAAYKSKDYAGDAAQKTKEASEYANDTTQKTAKYGADTAQKTKNKIQDVA 1446
Query: 741 SSSDPSITEEVKE 753
S + E+ KE
Sbjct: 1447SGTGEHSAEKAKE 1485
Score = 47.0 bits (110), Expect = 3e-05
Identities = 56/276 (20%), Positives = 108/276 (38%), Gaps = 6/276 (2%)
Frame = +1
Query: 474 TSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTT 533
++E G+ Q+ T + + + G E T+ E + + TT
Sbjct: 157 SNEDEGRDFQDVKAKTMETANKASETGKEGKEAAETWTEWAKEKITEGLGFKDKDKDSTT 336
Query: 534 TPKTEDSNADAA------GDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGNSVQ 587
T K + +DAA D+ K ++ + +A++ A K T + + ++
Sbjct: 337 TDKASEYASDAAHKTKHYADETAHKSKETAQKASDHAHDAAEK-TKGYAWDAS---KKIR 504
Query: 588 EGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEA 647
+ DAS + + A+D E D+ +AE N D + ++ G T GSA +
Sbjct: 505 DYATDASEKTKHYAAD---ESYADDDDDAEDNRD-FAGEAIHKAGEYT---GSAAQKAKE 663
Query: 648 SQVNSSDVSSEESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSEND 707
+++ +++++KE ++ + + S + G+A+ E KT E
Sbjct: 664 YATDAAQKAAQKAKEYATGAAHKS----------KEYAGDAAQKTKEYAGDATEKTKEYA 813
Query: 708 GEAAQNESVESEKAKDESAHSDEGSNSNTNDQGSSS 743
G+AAQ K KD + + + S D S
Sbjct: 814 GDAAQ-------KTKDYAGGATQKSKEYATDAAYKS 900
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 67.8 bits (164), Expect = 2e-11
Identities = 69/346 (19%), Positives = 135/346 (38%), Gaps = 6/346 (1%)
Frame = +3
Query: 207 EAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENE 266
++ K A +V ++ K + + EV+ A + KVE+ + + K + E
Sbjct: 108 KSSKKSATKVDAAPVVVSPVKSGKKGKRQAEEEVKAVSAKKQKVEEVAAKQKALKVVKKE 287
Query: 267 EIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKD 326
E +E + E K+ +++T K+ K + E + S E E K
Sbjct: 288 ESSSEESSESEDEQPVVKAPAPSKKTPAKKGNVKKAQPETTSEESDSDSSSSDEEEVKKP 467
Query: 327 KENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEG 386
++NG+ V+ + E+DS+E +E A K K A ++K D E
Sbjct: 468 VSKAVPSKNGSAPAKKVDTSEEEDSEESSDEDK-KPAAKAVPSK-NGSAPAKKAASDEED 641
Query: 387 NNEQHREENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVT 446
+E E+ A+ AV K+ + + + TE D++ ++ E E +K + ++
Sbjct: 642 TDESSDEDEEDEKPAAKAVPSKNGSVPAKKAD-TESSDEDSESSDE-EDKKPAAKASKNV 815
Query: 447 DSNATTNIQENGGENQAQTDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQNGNET 506
+ + E+ ++D + + S+ + + + + E DSSS +
Sbjct: 816 SAPTKKAASSSDEESDEESDEDEDAKPVSKPAAVAKKSKKDSSDSDDEDDDSSSDEDKKP 995
Query: 507 TS------HTTDKQNETSEDANSKVEDSSTDTTTPKTEDSNADAAG 546
+ DK S+ + + ED + T K +D AG
Sbjct: 996 VAAKKEDKMNVDKDGSDSDQSEEESEDEPSKTPQKKIKDVEMVDAG 1133
Score = 59.7 bits (143), Expect = 4e-09
Identities = 71/363 (19%), Positives = 137/363 (37%), Gaps = 20/363 (5%)
Frame = +3
Query: 416 SYNKTEQVDKNEKNESEMESQKSGTET--NEVTDSNATTNIQENGGENQAQTDNGSEKRS 473
S +VD S ++S K G EV +A E Q ++ S
Sbjct: 114 SKKSATKVDAAPVVVSPVKSGKKGKRQAEEEVKAVSAKKQKVEEVAAKQKALKVVKKEES 293
Query: 474 TSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTT 533
+SE + EQ + + + ++ GN + E+ D++S E+
Sbjct: 294 SSEESSESEDEQPVVKAPA-PSKKTPAKKGNVKKAQPETTSEESDSDSSSSDEEEVKKPV 470
Query: 534 TPKTEDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGNSVQEGKQDA 593
+ N A VD++ + S S A K P+ N K+ A
Sbjct: 471 SKAVPSKNGSAPAKKVDTSEEEDSEESSDEDKKPAAKAV--------PSKNGSAPAKKAA 626
Query: 594 SHEAQLT-ASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVNS 652
S E +SD E +K + + SV +++ S+ +SE S ++K+ + S
Sbjct: 627 SDEEDTDESSDEDEEDEKPAAKAVPSKNGSVPAKKADTESSDEDSESSDEEDKKPAAKAS 806
Query: 653 SDVSSEESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSENDGEAAQ 712
+VS+ K SS+ E S + + D +E+ V + + + ++
Sbjct: 807 KNVSAPTKKAASSSDEESDEES----------------DEDEDAKPVSKPAAV--AKKSK 932
Query: 713 NESVESEKAKDESAHSD-----------------EGSNSNTNDQGSSSDPSITEEVKESH 755
+S +S+ D+S+ + +GS+S+ +++ S +PS T + K
Sbjct: 933 KDSSDSDDEDDDSSSDEDKKPVAAKKEDKMNVDKDGSDSDQSEEESEDEPSKTPQKKIKD 1112
Query: 756 IDL 758
+++
Sbjct: 1113VEM 1121
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 63.5 bits (153), Expect = 3e-10
Identities = 47/184 (25%), Positives = 80/184 (42%)
Frame = +1
Query: 200 ENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEE 259
E+ + ++E KE EV E+ S+ +N + E+ + ED G+ V ++ E
Sbjct: 442 EDVPQESKSEVKEQTEVREQVSETDNSNARQFEDNP-GDLPEDATKGDSNVSSEEKSEEN 618
Query: 260 TKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQ 319
+ +K +E+ K E E K K++ E T E+ + GE K +
Sbjct: 619 STEKSSEDTKTEDEGK--------KTEDEGSNT------------ENNKDGEEASTKESE 738
Query: 320 ENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEK 379
+E K E+EE N++ +D E + E +SN E ++ Q K +E +SEK
Sbjct: 739 SDESEKKDESEENNKSDSD-----ESEKKSSDSNETTDSNVEEKVEQSQNKESDENASEK 903
Query: 380 QVQD 383
D
Sbjct: 904 NTDD 915
Score = 61.2 bits (147), Expect = 1e-09
Identities = 49/215 (22%), Positives = 90/215 (41%), Gaps = 1/215 (0%)
Frame = +1
Query: 256 ENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKD-KENEEIKNEVEDKEAGEVKE 314
+NE+ + E+K + EV+E+ S+ +N + ++N E K V
Sbjct: 436 QNEDVPQESKSEVKEQT------EVREQVSETDNSNARQFEDNPGDLPEDATKGDSNVSS 597
Query: 315 EKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEE 374
E+ +EN K E+ + G +D E ++ ++ EE++ E+E +E
Sbjct: 598 EEKSEENSTEKSSEDTKTEDEGKKTED---EGSNTENNKDGEEASTKESESDE------- 747
Query: 375 ASSEKQVQDGEGNNEQHREENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKNESEME 434
SEK+ + E N E +++ D ++ +S NK + +EKN +
Sbjct: 748 --SEKKDESEENNKSDSDESEKKSSDSNETTDSNVEEKVEQSQNKESDENASEKNTDDNA 921
Query: 435 SQKSGTETNEVTDSNATTNIQENGGENQAQTDNGS 469
+S +NEV S A + + N+ T GS
Sbjct: 922 KDQS---SNEVFPSGAQSELL-----NETTTQTGS 1002
Score = 57.0 bits (136), Expect = 3e-08
Identities = 54/185 (29%), Positives = 79/185 (42%), Gaps = 10/185 (5%)
Frame = +1
Query: 603 DTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVNSSDVSSEESKE 662
D E K + E + DN N+ E + D E + S+VSSEE E
Sbjct: 445 DVPQESKSEVKEQTEVREQVSETDNSNA----RQFEDNPGDLPEDATKGDSNVSSEEKSE 612
Query: 663 GSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSENDGEAAQ---NESVESE 719
+S ++S D + D G + D N + DGE A +ES ESE
Sbjct: 613 ENSTEKSSED-------TKTEDEGKKTEDEGSNTEN------NKDGEEASTKESESDESE 753
Query: 720 KAKDES-----AHSDEGSNSNTNDQGSSSDPSITEEVKESHIDLG--TLPESNAESNHRD 772
K KDES + SDE S ++D ++D ++ E+V++S E N + N +D
Sbjct: 754 K-KDESEENNKSDSDE-SEKKSSDSNETTDSNVEEKVEQSQNKESDENASEKNTDDNAKD 927
Query: 773 VSSAE 777
SS E
Sbjct: 928 QSSNE 942
Score = 55.5 bits (132), Expect = 8e-08
Identities = 47/186 (25%), Positives = 89/186 (47%), Gaps = 17/186 (9%)
Frame = +1
Query: 68 LVEETTVVDARHKEQDEESEHESKTEEQINLDEVNSVD-------ENEPEHKPEEEHIKL 120
++ ++VV ++++ +ES+ E K + ++ ++V+ D E+ P PE+ K
Sbjct: 406 MMTSSSVVPVQNEDVPQESKSEVKEQTEVR-EQVSETDNSNARQFEDNPGDLPEDA-TKG 579
Query: 121 DDANGVDENEVENKPE----------EHIKLDDANGVDENSEHSQDLVEHEGEGSSENAR 170
D +E EN E E K +D EN++ ++ E E S E+ +
Sbjct: 580 DSNVSSEEKSEENSTEKSSEDTKTEDEGKKTEDEGSNTENNKDGEEASTKESE-SDESEK 756
Query: 171 TDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKD 230
D ESE++ + +ES+ S N++ EE +++NKE+ E EK+ +N KD
Sbjct: 757 KD-ESEENNKSDSDESEKKSSDSNETTDSNVEEKVEQSQNKESDENASEKNTDDN--AKD 927
Query: 231 KENEEI 236
+ + E+
Sbjct: 928 QSSNEV 945
Score = 54.7 bits (130), Expect = 1e-07
Identities = 42/205 (20%), Positives = 91/205 (43%), Gaps = 7/205 (3%)
Frame = +1
Query: 492 GVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKT-EDSNADAAGD--- 547
GV + SSS + + ++E E + + S TD + + ED+ D D
Sbjct: 397 GVWMMTSSSVVPVQNEDVPQESKSEVKEQTEVREQVSETDNSNARQFEDNPGDLPEDATK 576
Query: 548 ---DVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGNSVQEGKQDASHEAQLTASDT 604
+V S KS +++ ++ + ++ ++E N + ++G++ ++ E++ S+
Sbjct: 577 GDSNVSSEEKSEENSTEKSSEDTKTEDEGKKTEDEGSNTENNKDGEEASTKESESDESEK 756
Query: 605 SSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVNSSDVSSEESKEGS 664
E +++ S+++ + + N +D SN + + NKE+ + S + + +K+ S
Sbjct: 757 KDESEENNKSDSDESEKKSSDSNETTD-SNVEEKVEQSQNKESDENASEKNTDDNAKDQS 933
Query: 665 SNSENSSDANQNGSNSNENDNGNAS 689
SN S A N G+ S
Sbjct: 934 SNEVFPSGAQSELLNETTTQTGSFS 1008
Score = 53.1 bits (126), Expect = 4e-07
Identities = 48/214 (22%), Positives = 89/214 (41%), Gaps = 21/214 (9%)
Frame = +1
Query: 401 ASSAVDHKSQDIFNESYNKT-------EQVDKNEKNESEMESQKSGTETNEVTDSNATTN 453
+SS V +++D+ ES ++ EQV + + + + G + T ++ +
Sbjct: 415 SSSVVPVQNEDVPQESKSEVKEQTEVREQVSETDNSNARQFEDNPGDLPEDATKGDSNVS 594
Query: 454 IQENGGENQAQTDNGSEKRSTSESGGQQHQE----QNNPTS----------DGVETVDSS 499
+E EN T+ SE T + G + E +NN D E D S
Sbjct: 595 SEEKSEENS--TEKSSEDTKTEDEGKKTEDEGSNTENNKDGEEASTKESESDESEKKDES 768
Query: 500 SQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSGN 559
+N + + K ++++E +S VE+ + + K D NA D ++ +SS+
Sbjct: 769 EENNKSDSDESEKKSSDSNETTDSNVEE-KVEQSQNKESDENASEKNTDDNAKDQSSNEV 945
Query: 560 SDSNANNGAYKETTNLVQNENPNGNSVQEGKQDA 593
S A + ETT + + + V+E K D+
Sbjct: 946 FPSGAQSELLNETTTQTGSFSTQCSRVKE*KGDS 1047
>TC87830 similar to GP|15451168|gb|AAK96855.1 putative protein {Arabidopsis
thaliana}, partial (35%)
Length = 2189
Score = 63.2 bits (152), Expect = 4e-10
Identities = 111/547 (20%), Positives = 203/547 (36%), Gaps = 22/547 (4%)
Frame = +1
Query: 192 KENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEE-----IKNEVEDTEA- 245
+ N+ + + +EN E+ +K AE H ++ E + +E+ E E+ EA
Sbjct: 55 EHNEPMAVQPQENAEESYSKTNAETHGVEAHNEPMAVQPQEDAEESFSKTNAEIHAVEAH 234
Query: 246 GEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVE 305
E + Q++ EE+ K N E E V + Q++ EE+ K N E
Sbjct: 235 NEPMAVQPQEDAEESYSKTNAETHGVEAHNEPMTV---QPQEDAEESYSKTNAETHAVEA 405
Query: 306 DKEAGEVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEK 365
E V+ ++ +E+ + E + + + +E+ E+ + E +G EA
Sbjct: 406 HNEPMAVQPQEDAEESYSKTNAETHGVEAHNEPIAVQPQEDAEESFSKTNAEIHGVEAHN 585
Query: 366 EEQGKIQEEASSEKQVQDGEGNNEQHREE-NYAGDNASSAVDHKS--QDIFNESYNKTEQ 422
E IQ + +E Q + +E H+ E ++ N A H + D+ ++T
Sbjct: 586 EPTA-IQPQEDAEAQPSEIPVPSECHQSEVDFGSHNNIEAHGHTNIISDVRELGCSQTA- 759
Query: 423 VDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGENQAQTDNGSEKR-----STSES 477
E N + + + S E + T ++ E EN+ N + T +
Sbjct: 760 ----EMNNAGINFEISSAENYSFVPGHETLSLTE-VFENELCRPNFFDASLPLMDKTDDL 924
Query: 478 GGQQHQEQ-NNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPK 536
G H + + PTS + +DSS NE D+ N + + + T T
Sbjct: 925 VGSIHTDMLSIPTS---QKMDSSPMLENEFAEDQHDRNNAGATEIAENAMEIRTQVETDS 1095
Query: 537 TEDSNADAAGDDVDSTYKS-SSGNSDSNANNGAYKETTNLVQNENP---NGNSVQEG-KQ 591
E D Y S ++G+ ++N Y + + P NGN++ G +
Sbjct: 1096LE----------ADHLYASMATGSKEAN----EYTDNQVFYNGDLPVEENGNNMLGGLNE 1233
Query: 592 DASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEASQVN 651
D L D ++ S N E D + + + ++ S + E
Sbjct: 1234DQIISPGLGCDDKDAKSGGLFSENVE--VDCLHSAALINESSLNDEENPVCQEAALQNTM 1407
Query: 652 SSDVSSEESKEGSSNSENSSDANQNGSNSNENDN--GNASHDANENENSVQRKTSENDGE 709
DVS+ S EN+ G + +D + DA + + EN G
Sbjct: 1408YPDVSAIRSPFADQTDENNMGGIDTGFLNVGDDEIIEDDDDDAGGFASGAEGTHLENSGW 1587
Query: 710 AAQNESV 716
+++ +V
Sbjct: 1588SSRTRAV 1608
>TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (15%)
Length = 557
Score = 62.4 bits (150), Expect = 6e-10
Identities = 38/104 (36%), Positives = 58/104 (55%), Gaps = 1/104 (0%)
Frame = +2
Query: 235 EIKNEVEDTEAGE-VKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETK 293
E+K ED E E ++ EK + + E K++ N+E K EVE+KE G+ EKS++E EE
Sbjct: 83 EVKETTEDKEEKEKIEAEKEAEGSIEEKEEGNDEEKTEVEEKEEGD-DNEKSEEETEE-- 253
Query: 294 DKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKENEEINQNGT 337
K E ++KE E + E+ + +E+T DK EE G+
Sbjct: 254 -------KEEGDEKEKTEAETEEEEEADEDTIDKSKEEDKAEGS 364
Score = 62.0 bits (149), Expect = 8e-10
Identities = 36/101 (35%), Positives = 62/101 (60%), Gaps = 1/101 (0%)
Frame = +2
Query: 267 EIKNEVEDKEAGE-VKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETK 325
E+K EDKE E ++ EK + + E K++ N+E K EVE+KE G+ EKS +E EE +
Sbjct: 83 EVKETTEDKEEKEKIEAEKEAEGSIEEKEEGNDEEKTEVEEKEEGD-DNEKSEEETEEKE 259
Query: 326 DKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKE 366
+ + +E + T+ ++ +E+ SKEE +++ G++ KE
Sbjct: 260 EGDEKEKTEAETEEEEEADEDTIDKSKEE-DKAEGSKGGKE 379
Score = 61.6 bits (148), Expect = 1e-09
Identities = 41/106 (38%), Positives = 59/106 (54%)
Frame = +2
Query: 198 VKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQEN 257
VKE E+K E E EA EK + + E K++ N+E K EVE+ E G+ EKS++E
Sbjct: 86 VKETTEDKEEKEKIEA-----EKEAEGSIEEKEEGNDEEKTEVEEKEEGDDN-EKSEEET 247
Query: 258 EETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNE 303
EE K E ++KE E + E+ ++ +E+T DK EE K E
Sbjct: 248 EE---------KEEGDEKEKTEAETEEEEEADEDTIDKSKEEDKAE 358
Score = 50.8 bits (120), Expect = 2e-06
Identities = 32/93 (34%), Positives = 49/93 (52%)
Frame = +2
Query: 305 EDKEAGEVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAE 364
E KE E KEEK + E E+ + EE + + K VEE E D E+ EE TE +
Sbjct: 83 EVKETTEDKEEKEKIEAEKEAEGSIEEKEEGNDEEKTEVEEKEEGDDNEKSEEE--TEEK 256
Query: 365 KEEQGKIQEEASSEKQVQDGEGNNEQHREENYA 397
+E K + EA +E++ + E ++ +EE+ A
Sbjct: 257 EEGDEKEKTEAETEEEEEADEDTIDKSKEEDKA 355
Score = 44.7 bits (104), Expect = 1e-04
Identities = 36/130 (27%), Positives = 56/130 (42%)
Frame = +2
Query: 349 QDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAVDHK 408
+++ E+KEE EAEKE +G I+E+ EGN+E+ E V+ K
Sbjct: 89 KETTEDKEEKEKIEAEKEAEGSIEEKE---------EGNDEEKTE-----------VEEK 208
Query: 409 SQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGENQAQTDNG 468
E+ D NEK+E E E ++ G E + + A T +E E+
Sbjct: 209 ------------EEGDDNEKSEEETEEKEEGDEKEK---TEAETEEEEEADEDTIDKSKE 343
Query: 469 SEKRSTSESG 478
+K S+ G
Sbjct: 344 EDKAEGSKGG 373
Score = 44.3 bits (103), Expect = 2e-04
Identities = 29/98 (29%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Frame = +2
Query: 172 DIESEQHVEKVGEESDNTESKE--NDSLVKENEENKHEAENKEAAEVHEEKSQQENEETK 229
D E ++ +E E + E KE ND E EE + +N+++ E EEK + + +E
Sbjct: 104 DKEEKEKIEAEKEAEGSIEEKEEGNDEEKTEVEEKEEGDDNEKSEEETEEKEEGDEKEKT 283
Query: 230 DKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEE 267
+ E E E+ EA E ++KS++E++ K +E
Sbjct: 284 EAETE------EEEEADEDTIDKSKEEDKAEGSKGGKE 379
Score = 33.9 bits (76), Expect = 0.24
Identities = 31/127 (24%), Positives = 51/127 (39%), Gaps = 1/127 (0%)
Frame = +3
Query: 204 NKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDK 263
N + + + + K +Q + + N+ + VKVEK+ E EE KD
Sbjct: 231 NLKKRQRRRKRAMKRRKLKQRRRRRRRPTRILLINQKKKIRPRVVKVEKNDNEVEEVKDD 410
Query: 264 -ENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENE 322
E + +K VEDK N++ +E EE +DK E + EK + E
Sbjct: 411 MEVDSVKEPVEDK-------------NDDDDIQEMEE-----DDKNVAESENEKMDVDAE 536
Query: 323 ETKDKEN 329
+ E+
Sbjct: 537 VKETTED 557
>TC83625 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (9%)
Length = 908
Score = 61.6 bits (148), Expect = 1e-09
Identities = 41/142 (28%), Positives = 76/142 (52%)
Frame = +2
Query: 165 SSENARTDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQE 224
SS+ ++I + + G+ ++ E D +K E++ + E K+ E+K + +
Sbjct: 449 SSKMEESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDK----EKKDKTD 616
Query: 225 NEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEK 284
+E KDK+++E K + E E V+ +++ +E KDKE K + E KE G+ ++K
Sbjct: 617 VDEGKDKKDKEKKKK----EKKEENVKGEEEDGDEKKDKE----KKKKEKKEKGKEDKDK 772
Query: 285 SQQENEETKDKENEEIKNEVED 306
+E + KDKE ++ KNE +D
Sbjct: 773 DGEEKKSKKDKEKKKDKNEDDD 838
Score = 57.4 bits (137), Expect = 2e-08
Identities = 37/134 (27%), Positives = 71/134 (52%), Gaps = 13/134 (9%)
Frame = +2
Query: 264 ENEEIKNEVEDKEAGEVKEEKSQ-------------QENEETKDKENEEIKNEVEDKEAG 310
E I +++DK AG+VKE+K + E +E KDKE ++ K +V++ +
Sbjct: 461 EESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKEKKD-KTDVDEGKDK 637
Query: 311 EVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGK 370
+ KE+K +++ EE E E+ G + KD +E +++ KE+ +E + E+++ K
Sbjct: 638 KDKEKKKKEKKEENVKGEEED----GDEKKD--KEKKKKEKKEKGKEDKDKDGEEKKSKK 799
Query: 371 IQEEASSEKQVQDG 384
+E+ + + DG
Sbjct: 800 DKEKKKDKNEDDDG 841
Score = 57.0 bits (136), Expect = 3e-08
Identities = 45/147 (30%), Positives = 80/147 (53%)
Frame = +2
Query: 185 ESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTE 244
E N K +D + +E+K E E E KS ++++E K+K+++E K++ D +
Sbjct: 461 EESNIPIKIDDKSAGDVKEDKVEIEKDL-----EIKSVEKDDEKKEKKDKEKKDKT-DVD 622
Query: 245 AGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEV 304
G+ K +K +++ K+K+ E +K E ED G+ K++K E K KE +E E
Sbjct: 623 EGKDKKDKEKKK----KEKKEENVKGEEED---GDEKKDK------EKKKKEKKEKGKED 763
Query: 305 EDKEAGEVKEEKSRQENEETKDKENEE 331
+DK+ EEK ++++E K +NE+
Sbjct: 764 KDKDG----EEKKSKKDKEKKKDKNED 832
Score = 55.8 bits (133), Expect = 6e-08
Identities = 40/140 (28%), Positives = 72/140 (50%), Gaps = 16/140 (11%)
Frame = +2
Query: 232 ENEEIKNEVEDTEAGEVKVEKSQ-------------QENEETKDKENEEIKNEVEDKEAG 278
E I +++D AG+VK +K + E +E KDKE K + D + G
Sbjct: 461 EESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKE----KKDKTDVDEG 628
Query: 279 EVKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENE---ETKDKENEEINQN 335
+ K++K +++ K+K+ E +K E ED + + KE+K +++ E E KDK+ EE ++
Sbjct: 629 KDKKDKEKKK----KEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGEE-KKS 793
Query: 336 GTDVKDHVEENHEQDSKEEK 355
D + ++N + D E+
Sbjct: 794 KKDKEKKKDKNEDDDGG*ER 853
Score = 52.0 bits (123), Expect = 9e-07
Identities = 31/126 (24%), Positives = 66/126 (51%), Gaps = 10/126 (7%)
Frame = +2
Query: 296 ENEEIKNEVEDKEAGEVKEE----------KSRQENEETKDKENEEINQNGTDVKDHVEE 345
E I +++DK AG+VKE+ KS ++++E K+K+++E ++ TDV + ++
Sbjct: 461 EESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKE-KKDKTDVDEGKDK 637
Query: 346 NHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAV 405
++ K+EK+E N E++ K +E +++ + G+ + ++ EE + +
Sbjct: 638 KDKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEKKK 817
Query: 406 DHKSQD 411
D D
Sbjct: 818 DKNEDD 835
Score = 49.3 bits (116), Expect = 6e-06
Identities = 36/130 (27%), Positives = 64/130 (48%), Gaps = 6/130 (4%)
Frame = +2
Query: 112 KPEEEHI--KLDDANGVDENEVENKPEEHIKLDDANGVDENSEHSQDLVEHE---GEGSS 166
K EE +I K+DD + D E + + E+ +++ DE E + + EG
Sbjct: 455 KMEESNIPIKIDDKSAGDVKEDKVEIEKDLEIKSVEKDDEKKEKKDKEKKDKTDVDEGKD 634
Query: 167 ENARTDIESEQHVEKV-GEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQEN 225
+ + + E+ E V GEE D E K+ + KE +E E ++K+ E+KS+++
Sbjct: 635 KKDKEKKKKEKKEENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDG---EEKKSKKDK 805
Query: 226 EETKDKENEE 235
E+ KDK ++
Sbjct: 806 EKKKDKNEDD 835
>TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical protein
{Dictyostelium discoideum}, partial (7%)
Length = 743
Score = 61.2 bits (147), Expect = 1e-09
Identities = 67/244 (27%), Positives = 100/244 (40%), Gaps = 20/244 (8%)
Frame = +1
Query: 323 ETKDKENEEINQNGTDVKDHVEENHEQDSKEEK-----------EESNGTEAEKEEQGKI 371
ETK KE+EE K+ E E D +E EE EKEE+G
Sbjct: 4 ETKVKEDEESRTRDGGEKEDGERGGEDDEMDENDPEKPEVVDRSEEFVDEGKEKEEEGDE 183
Query: 372 QEEASSEKQVQDG--EGNN-EQHREENYAGDNASSAVDHKSQDIFNESYN-KTEQVDKN- 426
+E +S+ + G E NN + REE Y GD+ASSAV H ++ E+ E D N
Sbjct: 184 KENENSKDEGNQGLVESNNIHEAREEQYKGDDASSAVTHDTRTTSTETETVSMENSDVNA 363
Query: 427 EKNESEMESQKSGTETNEVTDSNATTNIQE----NGGENQAQTDNGSEKRSTSESGGQQH 482
E ++ E++ + TE + + +I E G + + + N + S S H
Sbjct: 364 EMGITKPENKPNYTEDGIRSQPGSNFSITEVKLVVGAYSNSSSSNETGNNSLSNPVAGSH 543
Query: 483 QEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSKVEDSSTDTTTPKTEDSNA 542
Q N T+ S + N T K N + DA+S+ + K + N+
Sbjct: 544 Q---NNTALIYSVRHSEETSSNLTIVVPAAKNNMSGTDASSEHHKMVMVSGLDKNDTGNS 714
Query: 543 DAAG 546
AG
Sbjct: 715 TVAG 726
Score = 57.0 bits (136), Expect = 3e-08
Identities = 56/241 (23%), Positives = 97/241 (40%), Gaps = 11/241 (4%)
Frame = +1
Query: 460 ENQAQTDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQ---NGNETTSHTTDKQNE 516
+ +++T +G EK G ++N+P E VD S + G E +K+NE
Sbjct: 22 DEESRTRDGGEKEDGERGGEDDEMDENDPEKP--EVVDRSEEFVDEGKEKEEEGDEKENE 195
Query: 517 TSEDANSKVEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSG--------NSDSNANNGA 568
S+D ++ S + + E D A V +++S NSD NA G
Sbjct: 196 NSKDEGNQGLVESNNIHEAREEQYKGDDASSAVTHDTRTTSTETETVSMENSDVNAEMGI 375
Query: 569 YKETTNLVQNENPNGNSVQEGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNV 628
K N +G Q G + E +L S+ SS+ ET +S+ N
Sbjct: 376 TKPENK--PNYTEDGIRSQPGSNFSITEVKLVVGAYSNS-----SSSNETGNNSLSNPVA 534
Query: 629 NSDGSNTNSEGSANDNKEASQVNSSDVSSEESKEGSSNSENSSDANQNGSNSNENDNGNA 688
S +NT S ++E S + V + ++ +++ + S ++ND GN+
Sbjct: 535 GSHQNNTALIYSVRHSEETSSNLTIVVPAAKNNMSGTDASSEHHKMVMVSGLDKNDTGNS 714
Query: 689 S 689
+
Sbjct: 715 T 717
Score = 40.0 bits (92), Expect = 0.003
Identities = 49/262 (18%), Positives = 95/262 (35%), Gaps = 12/262 (4%)
Frame = +1
Query: 114 EEEHIKLDDANGVDENEVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDI 173
E+E + D ++ E + +E +DEN ++V+ E E
Sbjct: 19 EDEESRTRDGGEKEDGERGGEDDE---------MDENDPEKPEVVDRSEEFVDEG----- 156
Query: 174 ESEQHVEKVGEESDNTESKE--NDSLVKEN-----EENKHEAENKEAAEVHEEKSQQENE 226
+ E+ G+E +N SK+ N LV+ N E +++ ++ +A H+ ++
Sbjct: 157 ---KEKEEEGDEKENENSKDEGNQGLVESNNIHEAREEQYKGDDASSAVTHDTRTTSTET 327
Query: 227 ETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQ 286
ET EN ++ E+ + K + + T+D + + E V S
Sbjct: 328 ETVSMENSDVN--------AEMGITKPENKPNYTEDGIRSQPGSNFSITEVKLVVGAYS- 480
Query: 287 QENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKENEEI-----NQNGTDVKD 341
+ + + N + N V S + +EET + N +GTD
Sbjct: 481 -NSSSSNETGNNSLSNPVAGSHQNNTALIYSVRHSEETSSNLTIVVPAAKNNMSGTDASS 657
Query: 342 HVEENHEQDSKEEKEESNGTEA 363
+ ++ + N T A
Sbjct: 658 EHHKMVMVSGLDKNDTGNSTVA 723
Score = 31.6 bits (70), Expect = 1.2
Identities = 22/112 (19%), Positives = 46/112 (40%), Gaps = 12/112 (10%)
Frame = +1
Query: 658 EESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSENDGEAAQNESVE 717
EES+ + + +END E + K E +G+ +NE+ +
Sbjct: 25 EESRTRDGGEKEDGERGGEDDEMDENDPEKPEVVDRSEEFVDEGKEKEEEGDEKENENSK 204
Query: 718 SE------------KAKDESAHSDEGSNSNTNDQGSSSDPSITEEVKESHID 757
E +A++E D+ S++ T+D ++S + T ++ S ++
Sbjct: 205 DEGNQGLVESNNIHEAREEQYKGDDASSAVTHDTRTTSTETETVSMENSDVN 360
>TC92575 weakly similar to GP|21428434|gb|AAM49877.1 LD13350p {Drosophila
melanogaster}, partial (3%)
Length = 557
Score = 61.2 bits (147), Expect = 1e-09
Identities = 54/191 (28%), Positives = 88/191 (45%), Gaps = 1/191 (0%)
Frame = -1
Query: 161 EGEGSSENARTDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEK 220
EGE E+ + D E + E+ + E+D L +E E +K E E+ EA + +
Sbjct: 539 EGEELKEDKKVDAGDE-----IEEDKKDDNGFEDDKLSEETELSKEERESVEAKKPELDA 375
Query: 221 SQQENE-ETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGE 279
+E+ E KD+ +E+ EKSQ++ E+ KDK + + NE K
Sbjct: 374 MDEESILEGKDEGSEK---------------EKSQKDREDDKDKVDNK-SNEENIKVEKR 243
Query: 280 VKEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKENEEINQNGTDV 339
+K+ + N E K+ +E+K + EA + EK R +EE DKE G +V
Sbjct: 242 LKKRGKGKVNGENVKKKMKELKKIEDIPEAKDESIEKERSRDEEVGDKE----KVGGENV 75
Query: 340 KDHVEENHEQD 350
K ++E E +
Sbjct: 74 KKKMKELKEAE 42
Score = 43.5 bits (101), Expect = 3e-04
Identities = 40/182 (21%), Positives = 78/182 (41%), Gaps = 10/182 (5%)
Frame = -1
Query: 282 EEKSQQENEETKDKENEEIKNEVE-DKEAGEVKEEKSRQENEETKDKENEEINQNGTDVK 340
EE+ + E EE K+ + + +E+E DK+ E+ E E +E E + ++
Sbjct: 557 EEEKKAEGEELKEDKKVDAGDEIEEDKKDDNGFEDDKLSEETELSKEERESVEAKKPEL- 381
Query: 341 DHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQ---------DGEGNNEQH 391
D ++E + K+E E ++ ++E+ + S+E+ ++ G+ N E
Sbjct: 380 DAMDEESILEGKDEGSEKEKSQKDREDDKDKVDNKSNEENIKVEKRLKKRGKGKVNGENV 201
Query: 392 REENYAGDNASSAVDHKSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNAT 451
+++ + K + I E ++ E+V EK E +K E E + T
Sbjct: 200 KKKMKELKKIEDIPEAKDESIEKER-SRDEEVGDKEKVGGE-NVKKKMKELKEAEPTTPT 27
Query: 452 TN 453
TN
Sbjct: 26 TN 21
>TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1
{Arabidopsis thaliana}, partial (47%)
Length = 1538
Score = 60.5 bits (145), Expect = 2e-09
Identities = 60/268 (22%), Positives = 127/268 (47%), Gaps = 3/268 (1%)
Frame = +3
Query: 192 KENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVE 251
+ N S + N+ ++E+E+ E E EE + + ++++ ++ V
Sbjct: 282 RNNGSYLDRNDSRRNESESDE------ELKGLSYEEYRRLKRQKMRKSLKHC-IWNVTPS 440
Query: 252 KSQQENEETKDK-ENEEIKNEVEDKEAGEVKEE-KSQQENEETKDKENEEIKNEVEDKEA 309
++EN++ +D + EEI ++ K+ E+K++ KS+ E+EE+K+ ++E K
Sbjct: 441 PPRRENDDLEDYGKPEEISEIIDVKKDKEIKQKAKSESESEESKESDSELRK-------- 596
Query: 310 GEVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQG 369
K KS +++ E+ D E+E + VE+ + S++ E + T +E+E++
Sbjct: 597 ---KRRKSYKKSRES-DSESES--------ESEVEDRKRRKSRKYSESDSDTNSEEEDRK 740
Query: 370 KIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAVDHKS-QDIFNESYNKTEQVDKNEK 428
+ + SS ++ + R++ D+ S D +S D ++++ K++K
Sbjct: 741 RRRRRKSSRRR-------RKSSRKKRRCSDSDESETDDESGYDDSGRKRKRSKRSRKSKK 899
Query: 429 NESEMESQKSGTETNEVTDSNATTNIQE 456
SE S+ G+E E+ +A I E
Sbjct: 900 KSSEPVSE--GSEEIELGSDSAVAKINE 977
Score = 53.1 bits (126), Expect = 4e-07
Identities = 47/199 (23%), Positives = 80/199 (39%), Gaps = 5/199 (2%)
Frame = +3
Query: 172 DIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDK 231
D+E E++ E D + KE K E++ E+ KS +++ E+ D
Sbjct: 462 DLEDYGKPEEISEIIDVKKDKEIKQKAKSESESEESKESDSELRKKRRKSYKKSRES-DS 638
Query: 232 ENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEE 291
E+E ++EVED K KS++ +E D +EE + + + KS ++
Sbjct: 639 ESES-ESEVEDR-----KRRKSRKYSESDSDTNSEEEDRKRRRRRKSSRRRRKSSRKKRR 800
Query: 292 TKDKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKE-----NEEINQNGTDVKDHVEEN 346
D + E +E ++G ++ R + K E +EEI + E
Sbjct: 801 CSDSDESETDDESGYDDSGRKRKRSKRSRKSKKKSSEPVSEGSEEIELGSDSAVAKINEE 980
Query: 347 HEQDSKEEKEESNGTEAEK 365
+ D EEK EA K
Sbjct: 981 ID-DVDEEKMAEMNAEALK 1034
Score = 46.2 bits (108), Expect = 5e-05
Identities = 46/208 (22%), Positives = 87/208 (41%), Gaps = 1/208 (0%)
Frame = +3
Query: 70 EETTVVDARH-KEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDE 128
E + ++D + KE ++++ ES++EE D E + + + + D+ E
Sbjct: 489 EISEIIDVKKDKEIKQKAKSESESEESKESDS-----ELRKKRRKSYKKSRESDSESESE 653
Query: 129 NEVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDN 188
+EVE++ + + D NSE +D SS R ++ E +
Sbjct: 654 SEVEDRKRRKSRKYSESDSDTNSE-EEDRKRRRRRKSSRRRRKSSRKKRRCSDSDESETD 830
Query: 189 TESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEV 248
ES +DS K + K+++E E S +E E D +I E++D + E
Sbjct: 831 DESGYDDSGRKRKRSKRSRKSKKKSSEPVSEGS-EEIELGSDSAVAKINEEIDDVD--EE 1001
Query: 249 KVEKSQQENEETKDKENEEIKNEVEDKE 276
K+ + E + K+ + K ++D E
Sbjct: 1002KMAEMNAEALKLKELFESQKKPALDDDE 1085
Score = 45.8 bits (107), Expect = 6e-05
Identities = 69/310 (22%), Positives = 124/310 (39%), Gaps = 5/310 (1%)
Frame = +3
Query: 473 STSESGGQQHQEQNNPTSDGVETVDSSSQNGNET--TSHTTDKQNETSEDANSKVEDSST 530
ST E+ Q+H+ +D T DS N S + D ++ D + + S
Sbjct: 48 STIETPYQRHRH-----NDRRRTPDSDVSNDRPRYRRSPSYDSYDDRHTDRHRR-RSISP 209
Query: 531 DTTTPKTEDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGNSVQEGK 590
+ P+ D N ++ G+ + + + NNG+Y + + +NE+ S +E K
Sbjct: 210 ENRNPRHHD-NGNSNGNHLPKKF--------GHRNNGSYLDRNDSRRNES---ESDEELK 353
Query: 591 QDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEG--SANDNKEAS 648
+ E + + K N + +ND++ G +KE
Sbjct: 354 GLSYEEYRRLKRQKMRKSLKHCIWNVTPSPPRRENDDLEDYGKPEEISEIIDVKKDKEIK 533
Query: 649 QVNSSDVSSEESKEGSSNSENSSDANQNGSNSNENDNGNASH-DANENENSVQRKTSEND 707
Q S+ SEESKE S+SE ++ S E+D+ + S + + + RK SE+D
Sbjct: 534 QKAKSESESEESKE--SDSELRKKRRKSYKKSRESDSESESESEVEDRKRRKSRKYSESD 707
Query: 708 GEAAQNESVESEKAKDESAHSDEGSNSNTNDQGSSSDPSITEEVKESHIDLGTLPESNAE 767
+ E + + +S+ S S + S SD S T++ + + D G + +
Sbjct: 708 SDTNSEEEDRKRRRRRKSSRRRRKS-SRKKRRCSDSDESETDD-ESGYDDSGRKRKRSKR 881
Query: 768 SNHRDVSSAE 777
S S+E
Sbjct: 882 SRKSKKKSSE 911
>TC87387 homologue to PIR|T47775|T47775 hypothetical protein F24I3.230 -
Arabidopsis thaliana, partial (14%)
Length = 1074
Score = 59.7 bits (143), Expect = 4e-09
Identities = 45/185 (24%), Positives = 92/185 (49%), Gaps = 8/185 (4%)
Frame = +3
Query: 202 EENKHEAENKEAAEVHEEKSQ---QENEETKDKENEEIKNEVEDTEAGE-VKVEKSQQEN 257
E+ K E ++E +V ++ + ++ E+ D K++V + + E VK +K
Sbjct: 186 EDVKKEVADEEGEKVKKDDGEGRKRKKHESADSPAPAKKSKVAEVDGEEKVKTKKVDDAA 365
Query: 258 EETKDKENEEIKNEVEDKEAGEVK--EEKSQQENEETKDKENEEIKNEVEDKEAGEVKEE 315
E +D + E+ K + +DKE G EEK ++E ++ ++ E+ EVE K +
Sbjct: 366 VEVEDDKKEKKKKKKKDKENGAAASDEEKVEKEKKKKHKEKGEDGSPEVE-------KSD 524
Query: 316 KSRQENEETKDKENEEI--NQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQE 373
K +++++ET + + E+ ++ KD +++ D +ESN +EK+ + K +
Sbjct: 525 KKKKKHKETSEVGSPEVDKSEKKKKKKDKEAKDNAADISNGNDESNADRSEKKHKKKKNK 704
Query: 374 EASSE 378
+A E
Sbjct: 705 DAQEE 719
Score = 53.9 bits (128), Expect = 2e-07
Identities = 48/200 (24%), Positives = 91/200 (45%)
Frame = +3
Query: 128 ENEVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESD 187
+ EV ++ E +K DD G S D S+ A D E + +KV + +
Sbjct: 195 KKEVADEEGEKVKKDDGEGRKRKKHESAD--SPAPAKKSKVAEVDGEEKVKTKKVDDAAV 368
Query: 188 NTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGE 247
E + KE ++ K + + AA EEK ++E + K E E G
Sbjct: 369 EVEDDK-----KEKKKKKKKDKENGAAASDEEKVEKEKK----------KKHKEKGEDGS 503
Query: 248 VKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDK 307
+VEKS + ++ K KE E+ + DK E+K +++++E KD +I N ++
Sbjct: 504 PEVEKS--DKKKKKHKETSEVGSPEVDK-----SEKKKKKKDKEAKDNA-ADISNGNDES 659
Query: 308 EAGEVKEEKSRQENEETKDK 327
A +++ +++N++ +++
Sbjct: 660 NADRSEKKHKKKKNKDAQEE 719
Score = 50.1 bits (118), Expect = 3e-06
Identities = 54/238 (22%), Positives = 106/238 (43%)
Frame = +3
Query: 234 EEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETK 293
E++K EV D E GE KV+K E + K K+E D A K + ++ + EE
Sbjct: 186 EDVKKEVADEE-GE-KVKKDDGEGRKRK-------KHESADSPAPAKKSKVAEVDGEE-- 332
Query: 294 DKENEEIKNEVEDKEAGEVKEEKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKE 353
++K + D A EV+++K ++ ++ KDKEN G D EE E++ K+
Sbjct: 333 -----KVKTKKVDDAAVEVEDDKKEKKKKKKKDKEN------GAAASD--EEKVEKEKKK 473
Query: 354 EKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQHREENYAGDNASSAVDHKSQDIF 413
+ +E K E G + E S +K+ ++H+E + G
Sbjct: 474 KHKE-------KGEDGSPEVEKSDKKK--------KKHKETSEVG--------------- 563
Query: 414 NESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGENQAQTDNGSEK 471
+ +VDK+EK + + + + ++++ N +N + +++ + + +++
Sbjct: 564 ------SPEVDKSEKKKKKKDKEAKDNAA-DISNGNDESNADRSEKKHKKKKNKDAQE 716
Score = 37.7 bits (86), Expect = 0.017
Identities = 34/170 (20%), Positives = 72/170 (42%)
Frame = +3
Query: 588 EGKQDASHEAQLTASDTSSEQKKDESSNAETNTDSVQNDNVNSDGSNTNSEGSANDNKEA 647
EG++ HE ++D+ + KK + + + + V+ V+ + K+
Sbjct: 246 EGRKRKKHE----SADSPAPAKKSKVAEVD-GEEKVKTKKVDDAAVEVEDDKKEKKKKKK 410
Query: 648 SQVNSSDVSSEESKEGSSNSENSSDANQNGSNSNENDNGNASHDANENENSVQRKTSEND 707
+ +S+E K + + ++GS E ++ + ++TSE
Sbjct: 411 KDKENGAAASDEEKVEKEKKKKHKEKGEDGSPEVEK---------SDKKKKKHKETSE-V 560
Query: 708 GEAAQNESVESEKAKDESAHSDEGSNSNTNDQGSSSDPSITEEVKESHID 757
G ++S + +K KD+ A + SN ND+ S++D S + K+ + D
Sbjct: 561 GSPEVDKSEKKKKKKDKEAKDNAADISNGNDE-SNADRSEKKHKKKKNKD 707
>TC79702 MtN14
Length = 973
Score = 58.9 bits (141), Expect = 7e-09
Identities = 54/216 (25%), Positives = 98/216 (45%), Gaps = 14/216 (6%)
Frame = +2
Query: 67 PLVEETTVVDARHKEQDEESE----HESKTEEQI----------NLDEVNSVDENEPEHK 112
P++ VV A+ D+ ++ + KT E+I E +S +E +P K
Sbjct: 50 PIMATVEVVSAQTALSDDTAQLSLKSDEKTTEEIVESPITLAASETKEESSAEEAKPLVK 229
Query: 113 PEEEHIKLDDANGVDENEVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTD 172
E + + + V + E E K EE + + V+E E+ L E + EN +
Sbjct: 230 AETKEVVEEVVVPVKDTEEETKKEEQTETKEP--VEETKENGNSLNVEE---TKENGDSV 394
Query: 173 IESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKE 232
+E+ Q EK EES+ +++++V E E + + + E +EEK ++E+ + KE
Sbjct: 395 VEAVQ--EKPAEESETVNVVKDENVVAEPETKDNVKTEETSEEKNEEKVEKEDAMDEKKE 568
Query: 233 NEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEI 268
E I N + K+EK+++E K + E +
Sbjct: 569 EEVITN-------NDAKIEKNEEEAPIEKSE*GENV 655
Score = 57.8 bits (138), Expect = 2e-08
Identities = 58/214 (27%), Positives = 95/214 (44%), Gaps = 15/214 (7%)
Frame = +2
Query: 240 VEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEE 299
VE A + + Q + ++ +K EEI A E KEE S +E + E +E
Sbjct: 65 VEVVSAQTALSDDTAQLSLKSDEKTTEEIVESPITLAASETKEESSAEEAKPLVKAETKE 244
Query: 300 IKNE----VEDKEAGEVKEE--KSRQENEETKDKEN----EEINQNGTDVKDHVEENHEQ 349
+ E V+D E KEE ++++ EETK+ N EE +NG V + V+E +
Sbjct: 245 VVEEVVVPVKDTEEETKKEEQTETKEPVEETKENGNSLNVEETKENGDSVVEAVQEKPAE 424
Query: 350 DSKE---EKEESNGTEAEKEEQGKIQE--EASSEKQVQDGEGNNEQHREENYAGDNASSA 404
+S+ K+E+ E E ++ K +E E +E++V+ + +E+ EE ++A
Sbjct: 425 ESETVNVVKDENVVAEPETKDNVKTEETSEEKNEEKVEKEDAMDEKKEEEVITNNDA--- 595
Query: 405 VDHKSQDIFNESYNKTEQVDKNEKNESEMESQKS 438
K EKNE E +KS
Sbjct: 596 --------------------KIEKNEEEAPIEKS 637
Score = 57.4 bits (137), Expect = 2e-08
Identities = 49/184 (26%), Positives = 85/184 (45%), Gaps = 8/184 (4%)
Frame = +2
Query: 219 EKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEV----ED 274
+ + Q + ++ +K EEI A E K E S +E + E +E+ EV +D
Sbjct: 98 DDTAQLSLKSDEKTTEEIVESPITLAASETKEESSAEEAKPLVKAETKEVVEEVVVPVKD 277
Query: 275 KEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEAG----EVKEEKSRQENEETKDKENE 330
E KEE+++ + + KEN N E KE G E +EK +E+E ++E
Sbjct: 278 TEEETKKEEQTETKEPVEETKENGNSLNVEETKENGDSVVEAVQEKPAEESETVNVVKDE 457
Query: 331 EINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQ 390
+ + KD+V+ +++ EEK E EK E+ +E E+ + + + E+
Sbjct: 458 NVVAE-PETKDNVKT---EETSEEKNE------EKVEKEDAMDEKKEEEVITNNDAKIEK 607
Query: 391 HREE 394
+ EE
Sbjct: 608 NEEE 619
Score = 57.4 bits (137), Expect = 2e-08
Identities = 54/186 (29%), Positives = 86/186 (46%), Gaps = 1/186 (0%)
Frame = +2
Query: 127 DENEVENKPEEHIKLDDANGVDENS-EHSQDLVEHEGEGSSENARTDIESEQHVEKVGEE 185
DE E E I L + +E+S E ++ LV+ E + E ++ + K E+
Sbjct: 128 DEKTTEEIVESPITLAASETKEESSAEEAKPLVKAETKEVVEEVVVPVKDTEEETKKEEQ 307
Query: 186 SDNTESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEA 245
TE+KE KEN + + E KE + E Q+ K E E N V+D
Sbjct: 308 ---TETKEPVEETKENGNSLNVEETKENGDSVVEAVQE-----KPAEESETVNVVKDENV 463
Query: 246 GEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVE 305
K + EET +++NEE +VE ++A + K+E+ N + K ++NEE + +E
Sbjct: 464 VAEPETKDNVKTEETSEEKNEE---KVEKEDAMDEKKEEEVITNNDAKIEKNEE-EAPIE 631
Query: 306 DKEAGE 311
E GE
Sbjct: 632 KSE*GE 649
>TC87017 similar to PIR|A96564|A96564 unknown protein 23094-21772
[imported] - Arabidopsis thaliana, partial (30%)
Length = 1701
Score = 58.9 bits (141), Expect = 7e-09
Identities = 41/142 (28%), Positives = 72/142 (49%), Gaps = 4/142 (2%)
Frame = +3
Query: 219 EKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAG 278
E + + E T + + E+ K+E E + +++ + E++ T + + + N +D
Sbjct: 342 ESTAKSGEATSETQPEKAKDE----ETKQPEIKAPEVEDKSTSNNDVADRNNAGKDLAEK 509
Query: 279 EVKEEKSQQENEETKD--KENEEIKNEVEDKE-AGEV-KEEKSRQENEETKDKENEEINQ 334
E + S+ +NE++KD K E K EV DKE AGE+ KE + ++N E DK N
Sbjct: 510 ETDSDNSKVDNEQSKDGSKIENEDKKEVADKESAGEINKEPPTEEKNTENSDKNG---NS 680
Query: 335 NGTDVKDHVEENHEQDSKEEKE 356
D +D V E+ +++ K E
Sbjct: 681 ESKDKEDKVSESKDKEDKVSDE 746
Score = 55.8 bits (133), Expect = 6e-08
Identities = 37/130 (28%), Positives = 68/130 (51%), Gaps = 4/130 (3%)
Frame = +3
Query: 201 NEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIKNEVEDTEAGEVKVEKSQQENEET 260
+E +A+++E + + + E++ T + + + N +D E + S+ +NE++
Sbjct: 372 SETQPEKAKDEETKQPEIKAPEVEDKSTSNNDVADRNNAGKDLAEKETDSDNSKVDNEQS 551
Query: 261 KD--KENEEIKNEVEDKE-AGEV-KEEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEK 316
KD K E K EV DKE AGE+ KE ++++N E DK + EDK E K+++
Sbjct: 552 KDGSKIENEDKKEVADKESAGEINKEPPTEEKNTENSDKNGNSESKDKEDK-VSESKDKE 728
Query: 317 SRQENEETKD 326
+ +E T +
Sbjct: 729 DKVSDEPTAE 758
Score = 50.1 bits (118), Expect = 3e-06
Identities = 41/140 (29%), Positives = 66/140 (46%), Gaps = 7/140 (5%)
Frame = +3
Query: 162 GEGSSENA---RTDIESEQHVEKVGEESDNTESKENDSLVKENEENKHEAENKEAAEVHE 218
GE +SE D E++Q K E D + S N+ + N K AE E
Sbjct: 360 GEATSETQPEKAKDEETKQPEIKAPEVEDKSTS--NNDVADRNNAGKDLAEK----ETDS 521
Query: 219 EKSQQENEETKDK---ENEEIKNEVEDTEAGEVKVE-KSQQENEETKDKENEEIKNEVED 274
+ S+ +NE++KD ENE+ K + AGE+ E ++++N E DK + ED
Sbjct: 522 DNSKVDNEQSKDGSKIENEDKKEVADKESAGEINKEPPTEEKNTENSDKNGNSESKDKED 701
Query: 275 KEAGEVKEEKSQQENEETKD 294
K E K+++ + +E T +
Sbjct: 702 K-VSESKDKEDKVSDEPTAE 758
Score = 43.1 bits (100), Expect = 4e-04
Identities = 32/162 (19%), Positives = 68/162 (41%), Gaps = 3/162 (1%)
Frame = +3
Query: 349 QDSKEEKEESNGTEAEKEEQGKIQEEASSEKQVQDGEGNNEQHREENYAG-DNASSAVDH 407
+ + + E ++ T+ EK + + ++ +V+D +N + N AG D A D
Sbjct: 342 ESTAKSGEATSETQPEKAKDEETKQPEIKAPEVEDKSTSNNDVADRNNAGKDLAEKETDS 521
Query: 408 KSQDIFNESYNKTEQVDKNEKNESEMESQKSGTETNEVTDSNATTNIQENGGENQAQTDN 467
+ + NE +++ +K E + T+ T N +NG +++
Sbjct: 522 DNSKVDNEQSKDGSKIENEDKKEVADKESAGEINKEPPTEEKNTENSDKNGN-----SES 686
Query: 468 GSEKRSTSESGGQQHQEQNNPTSDG--VETVDSSSQNGNETT 507
++ SES ++ + + PT++G ++ S N N T
Sbjct: 687 KDKEDKVSESKDKEDKVSDEPTAEGGPFKSFQQLSSNQNAFT 812
Score = 36.2 bits (82), Expect = 0.049
Identities = 43/189 (22%), Positives = 73/189 (37%), Gaps = 12/189 (6%)
Frame = +3
Query: 465 TDNGSEKRSTSESGGQQHQEQNNPTSDGVETVDSSSQNGNETTSHTTDKQNETSEDANSK 524
T E S ++ + +E P E D S+ N + D+ N + A +
Sbjct: 348 TAKSGEATSETQPEKAKDEETKQPEIKAPEVEDKSTSNND-----VADRNNAGKDLAEKE 512
Query: 525 VEDSSTDTTTPKTEDSNADAAGDDVDSTYKSSSGNSDSNANNGAYKETTNLVQNENPNGN 584
+ ++ +++D + D + K S+G N E N +N + NGN
Sbjct: 513 TDSDNSKVDNEQSKDGSKIENEDKKEVADKESAGE----INKEPPTEEKN-TENSDKNGN 677
Query: 585 SVQEGKQDASHEAQLTASDTSSE-----------QKKDESSNAETN-TDSVQNDNVNSDG 632
S + K+D E++ S E Q+ + NA T S + ++ S G
Sbjct: 678 SESKDKEDKVSESKDKEDKVSDEPTAEGGPFKSFQQLSSNQNAFTGLAGSGFSSSLFSFG 857
Query: 633 SNTNSEGSA 641
S TN +GSA
Sbjct: 858 STTN-DGSA 881
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 58.2 bits (139), Expect = 1e-08
Identities = 62/287 (21%), Positives = 121/287 (41%), Gaps = 24/287 (8%)
Frame = +1
Query: 70 EETTVVDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDDANGVDEN 129
+ +D +E++ K + ++ + VD E + E+H ++ A V
Sbjct: 205 DREVALDKAFPRHEEDTAIVRKDDRRLRRLIESRVDNRE---EVREDHRRIRQAEIVSTI 375
Query: 130 EVENKPEEHIKLDDANGVDENSEHSQDLVEHEGEGSSENARTDIESEQHVEKVGEESDNT 189
E E K +E + L++ + D +E + + E + E A E E+ E+ EE ++
Sbjct: 376 EEEAKRQEGLDLEEQDE-DAMAERRRRIKEKLRQRDQEEALPQEEEEEEEEEEEEEEESD 552
Query: 190 ESKEND------SLVKE-----------NEENKHEAENKEAAEVHEEKSQQENEETKDKE 232
++D ++VK E + EAE + E + + ++ ET+
Sbjct: 553 YETDSDEEYTGVAMVKPVFVPKSERDTIAERERLEAEEEALEEARKRRMEERRNETRQIV 732
Query: 233 NEEIKNEVEDTEAGEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKE---EKSQQEN 289
EEI+ + E ++K+ + D + ++ NE E+ EA +V+E K +E+
Sbjct: 733 VEEIRKDQE--------IQKNIELEANIDDVDTDDELNEAEEYEAWKVREIGRIKRDRED 888
Query: 290 EETKDKENEEIKN----EVEDKEAGEVKEEKSRQENEETKDKENEEI 332
E KE EE++ E++ E K K Q +++ + E I
Sbjct: 889 REALLKEKEEVERVRNMTEEERREWERKNPKPSQSSKQEMEIYAENI 1029
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 58.2 bits (139), Expect = 1e-08
Identities = 64/315 (20%), Positives = 143/315 (45%), Gaps = 13/315 (4%)
Frame = +2
Query: 63 KDLNPLVEETTVVDARHKEQDEESEHESKTEEQINLDEVNSVDENEPEHKPEEEHIKLDD 122
K + L E+ +V+ ++ +++ E + + I D ++ E + K + ++ +
Sbjct: 104 KSFSTLKEKLVLVEKSFEDIRSKTQVEERRLQSIKRDIEECCEDLENKKKEIRDVGRIIE 283
Query: 123 ANGVDENEVENKPEEHIKLDDANGVDEN--SEHSQDLVEHEGEGSSENARTDIESEQHVE 180
A + +++ ++ + + G+ E+ EH ++L E E +I ++ +E
Sbjct: 284 ARKKMQGKIDECVKDFVAKEGQLGLMEDLIGEHKKELKTKELE--LRQVMDNISKQKELE 457
Query: 181 KVGEESDN---TESKENDSLVKENEENKHEAENKEAAEVHEEKSQQENEETKDKENEEIK 237
+E N ++ K +S +KE E + + E + EEK + + + K
Sbjct: 458 SQVKELVNDLVSKQKHFESHIKELESKERQLEGRLKEHELEEKEFEGRMNELESKERHFK 637
Query: 238 NEVEDTEA------GEVKVEKSQQENEETKDKENEEIKNEVEDKEAGEVKEEKSQQENEE 291
+EVE+ A G++K S+++ + KE E KN+ E++ +KE +S+++ E
Sbjct: 638 SEVEEINAKLMPLKGQIKELASKEKQLNGQVKELESKKNQFENR----IKELESKEKQHE 805
Query: 292 TKDKENEEIKNEVEDK-EAGEVKEEKSRQENEETKDKENEEINQ-NGTDVKDHVEENHEQ 349
+ KE+ + E E + + K++ + + + KEN+ ++Q G +K +
Sbjct: 806 GRVKEHASKEREFESQVMEQQFKKKLFEIQVKALESKENQLVDQMKGV*IK--------R 961
Query: 350 DSKEEKEESNGTEAE 364
D + E NG E++
Sbjct: 962 DGI*KPNEGNGIESK 1006
Score = 45.1 bits (105), Expect = 1e-04
Identities = 50/189 (26%), Positives = 87/189 (45%), Gaps = 9/189 (4%)
Frame = +2
Query: 203 ENKHEAENKEAAEVHEEKSQQENEETKDKENE-EIKNEVEDTEAGEVKVEKSQQENEETK 261
E+K E + KE E Q + +K KE E ++K V D S+Q++ E+
Sbjct: 377 EHKKELKTKEL-----ELRQVMDNISKQKELESQVKELVNDLV--------SKQKHFESH 517
Query: 262 DKENEEIKNEVEDKEAGEVKEEKSQQENEETKDKENEEIKNEVEDKEA------GEVKE- 314
KE E + ++E + EEK + + + K+EVE+ A G++KE
Sbjct: 518 IKELESKERQLEGRLKEHELEEKEFEGRMNELESKERHFKSEVEEINAKLMPLKGQIKEL 697
Query: 315 -EKSRQENEETKDKENEEINQNGTDVKDHVEENHEQDSKEEKEESNGTEAEKEEQGKIQE 373
K +Q N + K+ E+++ NQ +K E +SKE K+ +G+++E
Sbjct: 698 ASKEKQLNGQVKELESKK-NQFENRIK-------ELESKE-----------KQHEGRVKE 820
Query: 374 EASSEKQVQ 382
AS E++ +
Sbjct: 821 HASKEREFE 847
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.288 0.112 0.277
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,210,390
Number of Sequences: 36976
Number of extensions: 167926
Number of successful extensions: 2422
Number of sequences better than 10.0: 458
Number of HSP's better than 10.0 without gapping: 1418
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2001
length of query: 777
length of database: 9,014,727
effective HSP length: 104
effective length of query: 673
effective length of database: 5,169,223
effective search space: 3478887079
effective search space used: 3478887079
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 45 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0075.7


