

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
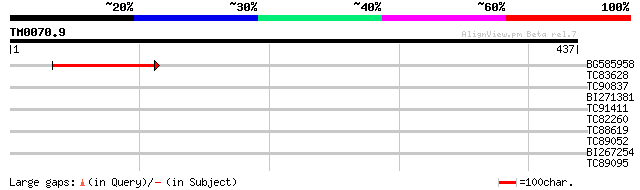
Query= TM0070.9
(437 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 99 3e-21
TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_18... 36 0.026
TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical... 34 0.097
BI271381 similar to GP|21397269|gb Unknown protein {Oryza sativa... 29 4.1
TC91411 similar to GP|15192738|gb|AAK91588.1 mRNA cap binding pr... 29 4.1
TC82260 similar to GP|19310478|gb|AAL84973.1 AT3g05350/T12H1_32 ... 28 5.3
TC88619 homologue to SP|Q43077|AMO_PEA Amine oxidase [copper-con... 28 5.3
TC89052 weakly similar to GP|166949|gb|AAA32913.1|| cytochrome P... 28 5.3
BI267254 similar to GP|19310478|gb AT3g05350/T12H1_32 {Arabidops... 28 5.3
TC89095 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_18... 28 7.0
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 99.0 bits (245), Expect = 3e-21
Identities = 46/82 (56%), Positives = 63/82 (76%)
Frame = +2
Query: 34 MEFYQQQQHANTVRPKKTRRVIKRDREAGNERLMKDYFSKNPVYTEELFRRRFRMRKHVF 93
++ Y Q++ A++ RP+ RR I+R+RE G +RL DYFS+ PVY +E FRRR+RM KHVF
Sbjct: 425 LKVYPQEEFASSSRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVF 604
Query: 94 LRIVEALGSYNPYFLMSVDAVG 115
LRIVEALG ++ YF ++VDA G
Sbjct: 605 LRIVEALGQHDEYFQLTVDATG 670
>TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_180
{Arabidopsis thaliana}, partial (23%)
Length = 773
Score = 36.2 bits (82), Expect = 0.026
Identities = 28/102 (27%), Positives = 43/102 (41%)
Frame = +2
Query: 77 YTEELFRRRFRMRKHVFLRIVEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAYGS 136
+ ++ FRR FRM K F I L S + + R + Q+ I LA G
Sbjct: 443 FPDDEFRRCFRMSKQTFNMICNELDS----SVTKKNTTLRDAIPVRQRVAVCIYRLATGE 610
Query: 137 PADSVDEYVRIGESTAIECLKNFVEGVCEVFGGQYLRRPNEE 178
P V + +G ST + + + V ++LR P+EE
Sbjct: 611 PLRLVSKKFGLGISTCHKLVLEVCAAIKSVLMQKFLRWPDEE 736
>TC90837 weakly similar to GP|22202752|dbj|BAC07409. hypothetical
protein~similar to Oryza sativa chromosome10
OSJNBa0079H13.17, partial (7%)
Length = 596
Score = 34.3 bits (77), Expect = 0.097
Identities = 24/91 (26%), Positives = 40/91 (43%), Gaps = 11/91 (12%)
Frame = +1
Query: 286 YYLADGIYPEWGTFVKTIPM-----------PQGEKKQKFAKRQEAARKDVERAFGVLQS 334
YYL D YP+ ++ P P ++ F + + R +ER+FGVL+
Sbjct: 10 YYLVDKGYPDKEGYMVPYPRIRYHQSQFEHEPPTNAQEAFNRAHSSLRSCIERSFGVLKK 189
Query: 335 RFAIVRGPSRWWHPNDMKSIIYACIILHNMI 365
R+ I+ ++ + +I A LHN I
Sbjct: 190 RWKILNKMPQFSVKTQI-DVIIAAFALHNYI 279
>BI271381 similar to GP|21397269|gb Unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 407
Score = 28.9 bits (63), Expect = 4.1
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = +2
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVR 340
++ F R + R +ER FGVL++RF I++
Sbjct: 137 EELFNYRHSSLRMTIERCFGVLKNRFPILK 226
>TC91411 similar to GP|15192738|gb|AAK91588.1 mRNA cap binding protein
{Arabidopsis thaliana}, partial (21%)
Length = 740
Score = 28.9 bits (63), Expect = 4.1
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = +1
Query: 209 CPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWH 241
CP + Q + D G P+ M A+ASQD ++W+
Sbjct: 217 CPSS*ARQLSN*DFGVPSYMC*AIASQDSFLWN 315
>TC82260 similar to GP|19310478|gb|AAL84973.1 AT3g05350/T12H1_32
{Arabidopsis thaliana}, partial (20%)
Length = 700
Score = 28.5 bits (62), Expect = 5.3
Identities = 13/33 (39%), Positives = 17/33 (51%), Gaps = 4/33 (12%)
Frame = -3
Query: 186 WGESR----GFPGMLGSIDCMHWEWKNCPVAWK 214
WG SR GF G+ G + + W WK C W+
Sbjct: 236 WGRSRNLRNGFYGV-GKMVSLRWNWKWCNCGWR 141
>TC88619 homologue to SP|Q43077|AMO_PEA Amine oxidase [copper-containing]
precursor (EC 1.4.3.6). [Garden pea] {Pisum sativum},
partial (67%)
Length = 1519
Score = 28.5 bits (62), Expect = 5.3
Identities = 24/102 (23%), Positives = 43/102 (41%), Gaps = 5/102 (4%)
Frame = +1
Query: 281 MYHIGYYLADGIYPEWG-TFVKTIPMPQGEKKQKFAKRQEA----ARKDVERAFGVLQSR 335
MY++ + + DG+ + T +KT+ + G K+K E D + G+
Sbjct: 766 MYYLDFDI-DGVQNSFEKTSLKTVKITDGSSKRKSYWTTETQIAKTESDAKITLGLAPGE 942
Query: 336 FAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDERNTYKGNF 377
AIV + ND+ + I H ++ ED+ +G F
Sbjct: 943 LAIVNPNKKTIVGNDVGYRLIPAIPAHPLLTEDDYPQIRGAF 1068
>TC89052 weakly similar to GP|166949|gb|AAA32913.1|| cytochrome P-450LXXIA1
(cyp71A1) {Persea americana}, partial (38%)
Length = 856
Score = 28.5 bits (62), Expect = 5.3
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +2
Query: 324 DVERAFGVLQSRFAIVRGPSRWWHPNDMK 352
D+E V+ + +AI R PS W HP + K
Sbjct: 464 DIEAGTEVIINAWAIARDPSSWEHPLEFK 550
>BI267254 similar to GP|19310478|gb AT3g05350/T12H1_32 {Arabidopsis
thaliana}, partial (21%)
Length = 620
Score = 28.5 bits (62), Expect = 5.3
Identities = 13/33 (39%), Positives = 17/33 (51%), Gaps = 4/33 (12%)
Frame = -2
Query: 186 WGESR----GFPGMLGSIDCMHWEWKNCPVAWK 214
WG SR GF G+ G + + W WK C W+
Sbjct: 145 WGRSRNLRNGFYGV-GKMVSLRWNWKWCNCGWR 50
>TC89095 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_180
{Arabidopsis thaliana}, partial (15%)
Length = 696
Score = 28.1 bits (61), Expect = 7.0
Identities = 17/37 (45%), Positives = 22/37 (58%), Gaps = 2/37 (5%)
Frame = +1
Query: 68 KDYFSK--NPVYTEELFRRRFRMRKHVFLRIVEALGS 102
KD++ K +P + EE FRR FRM K F I + L S
Sbjct: 481 KDWWQKCSSPDFPEEEFRRCFRMSKATFEFICQELES 591
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,070,218
Number of Sequences: 36976
Number of extensions: 174421
Number of successful extensions: 869
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 867
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 869
length of query: 437
length of database: 9,014,727
effective HSP length: 99
effective length of query: 338
effective length of database: 5,354,103
effective search space: 1809686814
effective search space used: 1809686814
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0070.9


