

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
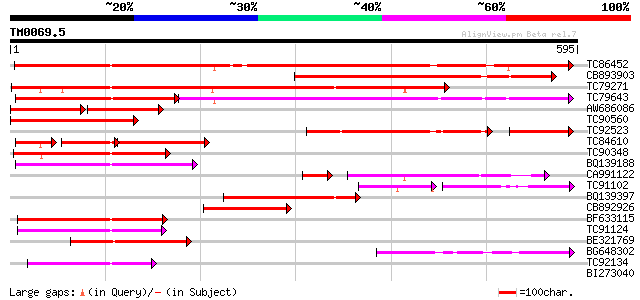
Query= TM0069.5
(595 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86452 similar to PIR|T09892|T09892 hypothetical protein T22A6.... 485 e-137
CB893903 weakly similar to GP|20466444|gb unknown protein {Arabi... 436 e-123
TC79271 weakly similar to GP|20466444|gb|AAM20539.1 unknown prot... 436 e-122
TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported]... 294 e-122
AW686086 similar to GP|20466444|gb| unknown protein {Arabidopsis... 142 3e-70
TC90560 weakly similar to GP|20466444|gb|AAM20539.1 unknown prot... 241 4e-64
TC92523 similar to PIR|T09892|T09892 hypothetical protein T22A6.... 156 7e-61
TC84610 similar to GP|20466444|gb|AAM20539.1 unknown protein {Ar... 99 4e-49
TC90348 similar to GP|15810435|gb|AAL07105.1 unknown protein {Ar... 175 5e-44
BQ139188 weakly similar to PIR|C86410|C864 protein F3M18.18 [imp... 149 4e-36
CA991122 similar to PIR|C86420|C86 unknown protein 124288-12173... 118 6e-35
TC91102 homologue to PIR|C86420|C86420 unknown protein 124288-1... 97 1e-34
BQ139397 weakly similar to PIR|T09892|T098 hypothetical protein ... 141 6e-34
CB892926 136 2e-32
BF633115 weakly similar to GP|15809820|gb At1g29690/F15D2_24 {Ar... 127 2e-29
TC91124 weakly similar to GP|15809820|gb|AAL06838.1 At1g29690/F1... 100 2e-21
BE321769 weakly similar to PIR|C86410|C864 protein F3M18.18 [imp... 99 4e-21
BG648302 similar to GP|22202725|dbj contains ESTs AU032617(S1163... 89 4e-18
TC92134 similar to GP|14209545|dbj|BAB56041. contains ESTs C7286... 86 3e-17
BI273040 similar to GP|6721157|gb| putative GDSL-motif lipase/ac... 30 2.6
>TC86452 similar to PIR|T09892|T09892 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (80%)
Length = 2233
Score = 485 bits (1249), Expect = e-137
Identities = 270/601 (44%), Positives = 373/601 (61%), Gaps = 15/601 (2%)
Frame = +3
Query: 6 VEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKG 65
VE + S+G G+DLT+D RLKFCK + +L+ ++ R + +PG + +V I CDKG
Sbjct: 198 VEDVIKSVGLGYDLTNDLRLKFCKYDSKLIAIDHDNLRTVELPGRVSIPNVPKSINCDKG 377
Query: 66 DLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGY 125
D R SD+LSF QMSE FNQ+ S+ GKIP+G+FN+ F F +G W DAANTK L DG
Sbjct: 378 DRMRLCSDVLSFQQMSEQFNQEVSLSGKIPTGHFNSAFQF-SGVWQRDAANTKSLAFDGV 554
Query: 126 VIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQ 185
I L+N+ +D+ +VLS V AVPSSWDP ALARFIEK+GTH +VG+ IGG D++ KQ
Sbjct: 555 SITLYNIALDKTHVVLSDHVKRAVPSSWDPAALARFIEKYGTHAVVGVKIGGTDIIYAKQ 734
Query: 186 DVSSNLEPSELKKNLDELGDQLFTGTCN--------FLPKTKDQKHKVPPAFDVFGPQIV 237
SS L+PS+++K L ++ D+LF G F K K + D+ Q
Sbjct: 735 QYSSPLQPSDVQKKLKDMADELFRGQAGQNNANDGTFNSKEKFMRDNGLGFLDI---QAQ 905
Query: 238 AFNNSTCVCAKDGITVICAKRGGD-TQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAP 296
++ + I +C ++GG+ Q +H+EW TV ++PD + SFIP TSLL
Sbjct: 906 SYRETEV----QDIKFMCKRKGGNGKQNLSHNEWCQTVLSQPDVISMSFIPITSLLGGIN 1073
Query: 297 GRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNL 356
G G+L+HAINLYLRYKP + +L FL++Q R WAP+ +L LGP ++ S SL +
Sbjct: 1074GSGYLTHAINLYLRYKPAIEELHQFLEFQLPRQWAPVFGELALGPDRKKSQSSSSLQFSF 1253
Query: 357 MGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSE 416
MGPKLYVNT+ V VG +PVTG+RL+LEG K N LAIHLQ+L + P K E S+
Sbjct: 1254MGPKLYVNTSPVVVGMKPVTGLRLYLEGKKSNCLAIHLQHLSSLPKTFQLKDETNRNVSD 1433
Query: 417 EIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRL 476
+ ++ E + K FSH+CTAPV+ D +VTG V + + VL LRL
Sbjct: 1434ASSERKYYEKVQWKSFSHICTAPVE-------SYDDNAVVTGAHFEVGETGLKKVLFLRL 1592
Query: 477 LFSKVSNAIVVKS-NW--TQGSTQRSGSGSGSGSSVFSAISTSIQGKDQ---KKPALVVV 530
F KV++A V++ W + G TQ+SG + + IST G + +P+ V V
Sbjct: 1593HFCKVADATRVRAPEWDGSPGLTQKSG-------MISTFISTRFSGPQKLPPPQPSDVNV 1751
Query: 531 DSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSL 590
+S+++P GPPVP QA KLLKFVDT+++ +GPQD PG+W+V+GARL ++KGKI L K+SL
Sbjct: 1752NSALYPGGPPVPAQAPKLLKFVDTTEMTRGPQDLPGYWVVSGARLYVEKGKISLKVKYSL 1931
Query: 591 L 591
L
Sbjct: 1932L 1934
>CB893903 weakly similar to GP|20466444|gb unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 815
Score = 436 bits (1122), Expect = e-123
Identities = 215/277 (77%), Positives = 244/277 (87%), Gaps = 3/277 (1%)
Frame = +3
Query: 300 FLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGP 359
FLSHAINLYL YKPP+SDL YFLDYQ +++WAP+HNDLPLGP +N +T SP L+LNLMGP
Sbjct: 3 FLSHAINLYLPYKPPMSDLSYFLDYQGYKIWAPVHNDLPLGPTTNISTISPFLTLNLMGP 182
Query: 360 KLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEII 419
KLYVNT KVTVGKRP+TGMRLFLEGMKCNRLAIH+++LLNTPTML+NKIEDTT WSEEI
Sbjct: 183 KLYVNTDKVTVGKRPITGMRLFLEGMKCNRLAIHVEHLLNTPTMLSNKIEDTTIWSEEIN 362
Query: 420 DDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSV-LHLRLLF 478
D+RF EAISGKKFSHVCTAPVKYNP WS++K+VAFIVTG QLHVKKH+++ V LHLRLLF
Sbjct: 363 DERFFEAISGKKFSHVCTAPVKYNPKWSTEKNVAFIVTGAQLHVKKHDTKRVLLHLRLLF 542
Query: 479 SKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQG--KDQKKPALVVVDSSVFP 536
SKVSN+ VVKSNWT+GS SG S +FSAISTSI G KDQKK + V++DSSVFP
Sbjct: 543 SKVSNSFVVKSNWTKGS-----SGLSQKSGIFSAISTSISGSSKDQKK-STVLLDSSVFP 704
Query: 537 TGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGA 573
TGPPVPVQ QK+LKFVDTS+LCKGPQ +PGHWLVTGA
Sbjct: 705 TGPPVPVQTQKMLKFVDTSELCKGPQHTPGHWLVTGA 815
>TC79271 weakly similar to GP|20466444|gb|AAM20539.1 unknown protein
{Arabidopsis thaliana}, partial (26%)
Length = 1719
Score = 436 bits (1121), Expect = e-122
Identities = 225/484 (46%), Positives = 320/484 (65%), Gaps = 25/484 (5%)
Frame = +2
Query: 3 EGIVEKALNSLGKGFDLTSDFRLKFCKG----------EERLVILNETERRELTVPGFGP 52
EGI AL SLGKGFDL SDFRL+F KG +RLV+L+E +R++ +PG G
Sbjct: 272 EGIERVALESLGKGFDLASDFRLRFAKGIRGGGNSNSGSKRLVVLDEQNKRDILIPGVGG 451
Query: 53 VT---DVSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGS 109
T VS +I+CDKGD R++SD+L F QMSE+ NQKS++ GKIPSGYFN +F +G
Sbjct: 452 ATVIKGVSENIRCDKGDRIRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDM-SGD 628
Query: 110 WASDAANTKFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHI 169
W DAA+ K+L DGY I L+ +H+ PLVL ++V ++VP+ WDP +L+RFI+ +GTHI
Sbjct: 629 WLRDAADIKYLAFDGYFISLYCLHLTASPLVLQEEVKKSVPAQWDPASLSRFIQTYGTHI 808
Query: 170 LVGLSIGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGT---CNFLPKTKDQKHKVP 226
+VG+++GG+D++ VKQ SS + P +++++L++LGD LF+ + KT D KHKVP
Sbjct: 809 VVGMAVGGQDVICVKQKHSSKIPPGDVRRHLEDLGDFLFSDVRSPSSLQRKTADSKHKVP 988
Query: 227 PAFD-VFGPQIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSF 285
F+ V F + + +KDG+T+IC+KRGGD +HS WL TVP+ P+A+ F F
Sbjct: 989 EVFNRVMQSNTTQFTSISETSSKDGLTIICSKRGGDVFKHSHSNWLQTVPSNPEAIIFKF 1168
Query: 286 IPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNR 345
+P +SLL PG G+LSHAINLYLRYKP DL YFL++Q R WAP+ +LPL R
Sbjct: 1169VPISSLLTGIPGSGYLSHAINLYLRYKPSPEDLQYFLEFQIPRQWAPMFCELPLRH-QRR 1345
Query: 346 TTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLN 405
T S SL +GPKL++++ +V ++PV G+RL+LEG K +RLA+H+ +L + P +
Sbjct: 1346KTSSLSLQFCCLGPKLHISSTEVVSEQKPVVGLRLYLEGKKSDRLALHINHLSSLPNKMI 1525
Query: 406 NKIEDTT-----TW---SEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVT 457
+ +T W E ++FLE + K+FS+VCTA VK++P+W +D +IVT
Sbjct: 1526LSSDASTPSIQSMWRGSDENESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNDCGGVYIVT 1705
Query: 458 GVQL 461
G QL
Sbjct: 1706GAQL 1717
>TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (74%)
Length = 2150
Score = 294 bits (752), Expect(2) = e-122
Identities = 174/425 (40%), Positives = 245/425 (56%), Gaps = 11/425 (2%)
Frame = +1
Query: 178 KDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCN---------FLPKTKDQKHKVPPA 228
KD+V +KQ SS++ P+EL+K L +L D+ F+ N K KD K+
Sbjct: 670 KDVVHIKQSKSSDIPPTELQKLLKQLADERFSAVSNQSSNVNPAAISGKLKDDHTKLRGL 849
Query: 229 FDVFGPQIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPF 288
P +V D I I +RGG +++WL T+ P+ + S +P
Sbjct: 850 HRNKPPSLVGRPIVKSHSKNDDIVSISVRRGGIDVFQPYNQWLSTISQSPNVISMSLVPI 1029
Query: 289 TSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPL-GPVSNRTT 347
TSLL + PG GFLSHA+NLYLRYKP + +L FL++Q R WAP++ DLPL +
Sbjct: 1030 TSLLNSVPGNGFLSHAVNLYLRYKPAIEELHQFLEFQLPRQWAPMYGDLPLVFDHKYKRN 1209
Query: 348 HSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNK 407
SPSL LMGPKLYVNT KV G RPVTG+RL+LEG K N L+IHLQ+L P +L
Sbjct: 1210 ASPSLQFTLMGPKLYVNTVKVDSGNRPVTGIRLYLEGKKNNHLSIHLQHLSEVPGVLEIS 1389
Query: 408 IEDTTTWSEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHE 467
+ + +E D + E + FSHV TAPV+YN S + IVT VK
Sbjct: 1390 EDHSYDPIDEPDDRGYYEPVKWSMFSHVYTAPVQYNSSRMDES--TSIVTKAWFEVKLMG 1563
Query: 468 SRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSA-ISTSIQGKDQKKPA 526
+ VL LRL +S V++A + +S W ST S SG S++ SA +S + + K +
Sbjct: 1564 MKKVLFLRLGYSTVASAKIRRSEWDGPST--SSRKSGFFSALMSAKLSQGLHSPE--KQS 1731
Query: 527 LVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWA 586
V ++S+++ GPPVP +A K+L FVDT ++ +GP+D PG+W+VTGA+L ++ G+I + A
Sbjct: 1732 KVDINSAIYHGGPPVPTRAPKMLNFVDTKEMVRGPEDPPGYWVVTGAKLCVEGGRISIKA 1911
Query: 587 KFSLL 591
K+SLL
Sbjct: 1912 KYSLL 1926
Score = 162 bits (409), Expect(2) = e-122
Identities = 80/172 (46%), Positives = 118/172 (68%)
Frame = +3
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKGD 66
EKA++ +G+G++L +D R CK + + + R+L P V +V + IK DKGD
Sbjct: 159 EKAVSVIGQGYNLCNDIRFSACKSRLIHIDNSSSNTRDLVFPSGVVVPNVPLSIKSDKGD 338
Query: 67 LTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYV 126
TR++SD+L+F QMSE FN++ S+ GKIPSG FN++F + W+ DAA+TK L DG+
Sbjct: 339 CTRFRSDVLTFIQMSEHFNRQLSLSGKIPSGQFNSMFDMKK-CWSRDAASTKSLAFDGWF 515
Query: 127 IKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGK 178
I L++V +DR + LS+ V + VP SW+P ALA FIEK+GTH++VG+ +GGK
Sbjct: 516 ITLYSVELDRTNITLSETVKKDVPCSWNPAALAEFIEKYGTHVVVGVKMGGK 671
>AW686086 similar to GP|20466444|gb| unknown protein {Arabidopsis
thaliana}, partial (13%)
Length = 666
Score = 142 bits (359), Expect(2) = 3e-70
Identities = 68/79 (86%), Positives = 74/79 (93%)
Frame = +2
Query: 1 MGEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDI 60
M EGIVEKALNSLGKGFDLTSDFRLKFCKGEERL++LNE E+REL+VPGFG + DVSVDI
Sbjct: 2 MDEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLILLNEIEKRELSVPGFGSIKDVSVDI 181
Query: 61 KCDKGDLTRYQSDILSFTQ 79
KCDKGD TRYQSDIL+FTQ
Sbjct: 182KCDKGDRTRYQSDILTFTQ 238
Score = 141 bits (355), Expect(2) = 3e-70
Identities = 64/80 (80%), Positives = 76/80 (95%)
Frame = +1
Query: 82 EMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYVIKLFNVHIDRYPLVL 141
++FN+KSSIPGKIPSGYFNTVFGF+ GSWA++AANTK LG+DGY+IKLFN+HID YPL+L
Sbjct: 238 KLFNRKSSIPGKIPSGYFNTVFGFDEGSWAAEAANTKCLGVDGYLIKLFNLHIDPYPLLL 417
Query: 142 SKQVIEAVPSSWDPTALARF 161
SKQVI+AVPSSWDP ALAR+
Sbjct: 418 SKQVIQAVPSSWDPPALARY 477
>TC90560 weakly similar to GP|20466444|gb|AAM20539.1 unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 439
Score = 241 bits (616), Expect = 4e-64
Identities = 114/135 (84%), Positives = 128/135 (94%)
Frame = +3
Query: 1 MGEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDI 60
M EGIVEKALNSLGKGFDLTSDFRLKFCKGEERL++LNE E+REL+VPGFG + DVSVDI
Sbjct: 30 MDEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLILLNEIEKRELSVPGFGSIKDVSVDI 209
Query: 61 KCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFL 120
KCDKGD TRYQSDIL+FTQMSE+FN+KSSIPGKIPSGYFNTVFGF+ GSWA++AANTK L
Sbjct: 210 KCDKGDRTRYQSDILTFTQMSELFNRKSSIPGKIPSGYFNTVFGFDEGSWAAEAANTKCL 389
Query: 121 GLDGYVIKLFNVHID 135
G+DGY+IKLFN+HID
Sbjct: 390 GVDGYLIKLFNLHID 434
>TC92523 similar to PIR|T09892|T09892 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (45%)
Length = 1327
Score = 156 bits (395), Expect(2) = 7e-61
Identities = 89/195 (45%), Positives = 122/195 (61%)
Frame = +1
Query: 312 KPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVG 371
KPP+ +L FL++Q R WAP+ +DLPLGP + + S SL + MGP+LYVNT V VG
Sbjct: 1 KPPIEELHQFLEFQLPRQWAPVFSDLPLGPQWKQRS-SASLQFSFMGPRLYVNTIPVDVG 177
Query: 372 KRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKK 431
KRPVTG+RL+LEG K NRLAIH+Q+L + P + + + + + D RF E + K
Sbjct: 178 KRPVTGLRLYLEGKKSNRLAIHMQHLSSLPKIFQLEDDSNENFRRKSYDKRFYEKVQWKN 357
Query: 432 FSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNW 491
FSHVCTAPV+ S+++++ +VTG QL V+ + +++L LRL FS V A VK
Sbjct: 358 FSHVCTAPVE------SEEELS-VVTGAQLQVENYGFKNILFLRLKFSTVLGAKEVKHPE 516
Query: 492 TQGSTQRSGSGSGSG 506
GS G G SG
Sbjct: 517 WDGS---PGLGPKSG 552
Score = 96.3 bits (238), Expect(2) = 7e-61
Identities = 43/67 (64%), Positives = 59/67 (87%)
Frame = +3
Query: 525 PALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICL 584
PA V ++S+V+P GPPVPVQA KLLKFVDT+++ +GPQ++PG+W+VTGARL+++KGKI L
Sbjct: 615 PADVNINSAVYPGGPPVPVQAPKLLKFVDTTEMTRGPQETPGYWVVTGARLLVEKGKISL 794
Query: 585 WAKFSLL 591
K+SLL
Sbjct: 795 RVKYSLL 815
>TC84610 similar to GP|20466444|gb|AAM20539.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 681
Score = 99.0 bits (245), Expect(3) = 4e-49
Identities = 42/96 (43%), Positives = 71/96 (73%)
Frame = +3
Query: 114 AANTKFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGL 173
A + K L DGY I L+N+H+ L+L +++ ++VP+ WDP +L+RFI +GTHI+VG+
Sbjct: 381 AQDIKSLAFDGYFISLYNLHLTASHLILQEELKKSVPAHWDPASLSRFIATYGTHIIVGM 560
Query: 174 SIGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFT 209
++GG+D++ VKQ SS + P +L+++L++L D LF+
Sbjct: 561 AVGGQDVICVKQKHSSKVPPGDLRRHLEDLXDFLFS 668
Score = 83.2 bits (204), Expect(3) = 4e-49
Identities = 39/61 (63%), Positives = 46/61 (74%)
Frame = +2
Query: 55 DVSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDA 114
DVS DI+CDKGD R++SD+L F QMSEM NQKS+I GKIPSGYFN VF +G W D
Sbjct: 206 DVSEDIRCDKGDRVRFKSDVLQFNQMSEMLNQKSAIQGKIPSGYFNAVFDL-SGDWFRDC 382
Query: 115 A 115
+
Sbjct: 383 S 385
Score = 52.4 bits (124), Expect(3) = 4e-49
Identities = 25/46 (54%), Positives = 35/46 (75%), Gaps = 3/46 (6%)
Frame = +1
Query: 7 EKALNSLGKGFDLTSDFRLKFCKG---EERLVILNETERRELTVPG 49
++AL+ LGKGFDLTSDFR+KF KG RL +++E +R++ VPG
Sbjct: 52 KRALDCLGKGFDLTSDFRMKFSKGLINGGRLXVIDEMNKRDIMVPG 189
>TC90348 similar to GP|15810435|gb|AAL07105.1 unknown protein {Arabidopsis
thaliana}, partial (48%)
Length = 663
Score = 175 bits (443), Expect = 5e-44
Identities = 88/168 (52%), Positives = 124/168 (73%), Gaps = 4/168 (2%)
Frame = +2
Query: 5 IVEKALNSLGKGFDLTSDFRLKFCKGEE---RLVILNE-TERRELTVPGFGPVTDVSVDI 60
+ E A+ S+G+G+D++SD RLKFCKG+ RL+ ++E + RE+ +PG + +VS I
Sbjct: 161 VAEIAIGSIGRGYDISSDIRLKFCKGDSIHSRLIEIDEDNDLREVVLPGGVSLPNVSKLI 340
Query: 61 KCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFL 120
KCDKG+ TR++SD+LSF QM+E FNQ+ S+ GKIPSG FN++F F +GSW DAA+TK L
Sbjct: 341 KCDKGERTRFRSDVLSFQQMTEQFNQELSLTGKIPSGLFNSMFEF-SGSWQKDAAHTKTL 517
Query: 121 GLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTH 168
DG +I L+ V +++ ++L V +AVPSSWDP ALARFI+ FGTH
Sbjct: 518 AFDGVLITLYTVALEKSQMLLCDHVKKAVPSSWDPPALARFIDTFGTH 661
>BQ139188 weakly similar to PIR|C86410|C864 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (10%)
Length = 667
Score = 149 bits (375), Expect = 4e-36
Identities = 82/194 (42%), Positives = 116/194 (59%), Gaps = 3/194 (1%)
Frame = +1
Query: 7 EKALNSLGKGFDLTSDFRLKFCK--GEERLVILNETERRELTVPGFGPVTDVSVDIKCDK 64
+KA+NS+G GFD+T D CK G + I N+ R L +PG + +VS +KC +
Sbjct: 79 QKAINSIGLGFDITLDINFDNCKSIGSPLIFINNQQHCRHLELPGGVTIPNVSNSVKCVR 258
Query: 65 GDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDG 124
G+ R SD+L+ QM + FN + + G SG+F FG +G D A+ K L DG
Sbjct: 259 GESIRIHSDVLTLHQMLQHFNHEMRLVGDTASGHFCASFGL-SGRCIKDLASIKSLAYDG 435
Query: 125 YVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVK 184
+ IK + V ++ Y L V EAVPSSWDP ALARFIE+FGTH++VG+S+G KD+ V
Sbjct: 436 WFIKRYAVELENYHGELHDHVKEAVPSSWDPEALARFIERFGTHVIVGVSMGXKDVFYVX 615
Query: 185 Q-DVSSNLEPSELK 197
Q D S +P+ ++
Sbjct: 616 QEDTSXXXDPTSIQ 657
>CA991122 similar to PIR|C86420|C86 unknown protein 124288-121737 [imported]
- Arabidopsis thaliana, partial (32%)
Length = 744
Score = 118 bits (296), Expect(2) = 6e-35
Identities = 75/218 (34%), Positives = 118/218 (53%), Gaps = 6/218 (2%)
Frame = +3
Query: 355 NLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTT- 413
++MG KLYV+ ++TVG+RPVTG+RL LEG K NRL++HLQ+L++ P +L +
Sbjct: 138 SIMGQKLYVSQEQITVGRRPVTGIRLCLEGNKQNRLSVHLQHLVSLPKILQPYWDSHVAI 317
Query: 414 ----WS-EEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHES 468
W E D R+ E + K FSHV TAP++ ++ D +IVTG QL V S
Sbjct: 318 GAPKWQGPEEQDSRWFEPVKWKNFSHVSTAPIENPETFIGDFSGVYIVTGAQLGVWDFGS 497
Query: 469 RSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALV 528
R+VL+++LL+S++ + +S W S S +G++ + ST++ ++
Sbjct: 498 RNVLYMKLLYSRLPGCTIRRSLWDH-IPNTSPKSSTAGNTSNTDNSTNLGSRENT*QT-- 668
Query: 529 VVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPG 566
L+K+VD S+L +GP+D PG
Sbjct: 669 ------------------SLVKYVDLSELRQGPEDPPG 728
Score = 47.8 bits (112), Expect(2) = 6e-35
Identities = 17/31 (54%), Positives = 25/31 (79%)
Frame = +1
Query: 308 YLRYKPPLSDLPYFLDYQAHRLWAPIHNDLP 338
YL YKPP+ +L YFL++Q R+WAP+H+ +P
Sbjct: 1 YLEYKPPIEELRYFLEFQIPRVWAPLHDRVP 93
>TC91102 homologue to PIR|C86420|C86420 unknown protein 124288-121737
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1061
Score = 97.4 bits (241), Expect(2) = 1e-34
Identities = 62/138 (44%), Positives = 78/138 (55%)
Frame = +1
Query: 455 IVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAIS 514
IVTG QL V +++VLHL+LLFSKV + +S W + G+S SA
Sbjct: 580 IVTGAQLGVWDFGAKNVLHLKLLFSKVPGCTIRRSVWDHNPSTPVAGHKSDGASSSSAKK 759
Query: 515 TSIQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGAR 574
TS D+KK DSSV KL K VD +++ KGPQD PGHWLVTGA+
Sbjct: 760 TS----DEKKE-----DSSV---------HIGKLAKIVDMTEMSKGPQDIPGHWLVTGAK 885
Query: 575 LMLDKGKICLWAKFSLLN 592
L ++KGKI L K+SLLN
Sbjct: 886 LGVEKGKIVLRIKYSLLN 939
Score = 67.8 bits (164), Expect(2) = 1e-34
Identities = 42/94 (44%), Positives = 53/94 (55%), Gaps = 12/94 (12%)
Frame = +3
Query: 367 KVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLN-----NKIEDTTTW-SEEIID 420
+VTVG++PVTG+RL LEG K NRLAIHLQ+L++ P L + W E D
Sbjct: 297 QVTVGRKPVTGLRLSLEGNKQNRLAIHLQHLVSLPKNLQPHWDAHMAIGAPKWQGPEEQD 476
Query: 421 DRFLEAISGKKFSHVCTAPVKY------NPSWSS 448
R+ E I K FSHV TAP++ P W S
Sbjct: 477 SRWFEPIKWKNFSHVSTAPIEIT*N*HRRPLWCS 578
>BQ139397 weakly similar to PIR|T09892|T098 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (13%)
Length = 618
Score = 141 bits (356), Expect = 6e-34
Identities = 68/145 (46%), Positives = 96/145 (65%), Gaps = 1/145 (0%)
Frame = +3
Query: 225 VPPAFD-VFGPQIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDF 283
VP F+ V + F + + +KDG+T+IC+KRGGD +HS WL T+ + P+A+ F
Sbjct: 6 VPEVFNRVLQSSTMQFTSISETSSKDGLTIICSKRGGDVFKHSHSSWLQTMASNPEAIHF 185
Query: 284 SFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVS 343
F+P +SLL PG G+LSHAINLYLRYKP DL YFL++Q R WAP+ ++LPL
Sbjct: 186 KFVPISSLLTGIPGSGYLSHAINLYLRYKPTPDDLQYFLEFQIPREWAPMFSELPLRH-Q 362
Query: 344 NRTTHSPSLSLNLMGPKLYVNTAKV 368
+ T+SP L + M PKL++N+ +V
Sbjct: 363 RKKTYSPPLQFSFMSPKLHINSTQV 437
>CB892926
Length = 282
Score = 136 bits (343), Expect = 2e-32
Identities = 63/92 (68%), Positives = 74/92 (79%)
Frame = +3
Query: 204 GDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNSTCVCAKDGITVICAKRGGDTQ 263
GDQLF+GTC FLP+ +DQKHKVP AFDVFGP VAF+ ST VCA DG+ V+C RGGDT
Sbjct: 3 GDQLFSGTCIFLPRNRDQKHKVPAAFDVFGPHFVAFSGSTSVCAVDGLFVLCVGRGGDTH 182
Query: 264 VSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAA 295
+S+HS+WLLTVP PDA+D S IP TSLL+ A
Sbjct: 183 LSSHSDWLLTVPITPDALDKSCIPITSLLRGA 278
>BF633115 weakly similar to GP|15809820|gb At1g29690/F15D2_24 {Arabidopsis
thaliana}, partial (48%)
Length = 630
Score = 127 bits (318), Expect = 2e-29
Identities = 71/159 (44%), Positives = 100/159 (62%), Gaps = 2/159 (1%)
Frame = +2
Query: 9 ALNSLGKGFDLTSDFRLKFCKGE--ERLVILNETERRELTVPGFGPVTDVSVDIKCDKGD 66
++ +LG+GFD+TSD RL +CKG RLV L+E R+L + V +VS+DI +G
Sbjct: 74 SIQALGRGFDVTSDIRLLYCKGAPGSRLVHLDEEHNRDLALSQELVVPNVSLDIDFSRGK 253
Query: 67 LTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYV 126
++ + SF +M+E FN++S I GKIP G FN++F F GS DAA TK L + GY
Sbjct: 254 SGIEKTPVCSFEKMAEYFNERSGIEGKIPLGSFNSMFNF-TGSSMVDAAATKSLAMVGYF 430
Query: 127 IKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKF 165
I LF V + + L L+ +V AVP SWDP +LAR ++ F
Sbjct: 431 IPLFEVKLTKQNLALNDEVRRAVPYSWDPASLARMLQSF 547
>TC91124 weakly similar to GP|15809820|gb|AAL06838.1 At1g29690/F15D2_24
{Arabidopsis thaliana}, partial (37%)
Length = 653
Score = 100 bits (248), Expect = 2e-21
Identities = 65/159 (40%), Positives = 90/159 (55%), Gaps = 3/159 (1%)
Frame = +3
Query: 9 ALNSLGKGFDLTSDFRLKFCKGE--ERLVILNETERRELTVPGFGPVTDVSVDIKCDKGD 66
++ +LG+GFD+TSD RL +CKG RLV L+E R+L + V +VS+DI +G
Sbjct: 159 SIQALGRGFDVTSDIRLLYCKGAPGSRLVHLDEEHNRDLALSQELVVPNVSLDIDFSRGK 338
Query: 67 LTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYV 126
++ + SF +M+E FN++S I GKIP G FN++F F GS DAA TK L +
Sbjct: 339 SGIEKTPVCSFEKMAEYFNERSGIEGKIPLGSFNSMFNF-TGSSMVDAAATKSLAMGWIF 515
Query: 127 IKLFNVHIDRYPLVL-SKQVIEAVPSSWDPTALARFIEK 164
I+ +V AVP SWDP +LA FI K
Sbjct: 516 HSSIRS*INETKFXP*MMKVRPAVPYSWDPXSLASFIGK 632
>BE321769 weakly similar to PIR|C86410|C864 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (9%)
Length = 378
Score = 99.4 bits (246), Expect = 4e-21
Identities = 53/126 (42%), Positives = 76/126 (60%)
Frame = +1
Query: 65 GDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDG 124
G+ R SD+LS QM + FN + + GK SG+F FG + + L DG
Sbjct: 1 GESIRINSDVLSLQQMLQHFNHEMRLDGKTASGHFCASFGLHFHG-T*ELDSIIHLAYDG 177
Query: 125 YVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVK 184
+ IK + V +++Y L V E VPS WD AL RFIE+FGTH++VG+S+GGKD++ V+
Sbjct: 178 WFIKRYAVELEKYHGQLHDHVKEVVPSLWDAGALTRFIERFGTHVIVGVSMGGKDVLYVR 357
Query: 185 QDVSSN 190
QD +S+
Sbjct: 358 QDDTSD 375
>BG648302 similar to GP|22202725|dbj contains ESTs AU032617(S11633)
D46758(S11633) C73071(E2861)~similar to Oryza sativa
chromosome 1, partial (5%)
Length = 816
Score = 89.4 bits (220), Expect = 4e-18
Identities = 70/208 (33%), Positives = 104/208 (49%), Gaps = 1/208 (0%)
Frame = +2
Query: 386 KCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKKFSHVCTAPVKYNPS 445
+ NRLAI+LQ+L++ P L S + ++ + + FS+VCTAP++ + S
Sbjct: 8 RSNRLAINLQHLVSLPKSLPLADNAPAYSSCDSYSCKYHKKLKWNCFSYVCTAPIESDDS 187
Query: 446 WSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKS-NWTQGSTQRSGSGSG 504
S IVTG QL V+K L LRL FSKV A + K W Q S
Sbjct: 188 LS-------IVTGAQLQVEK----KCLLLRLHFSKVIGATL*KPPEWDQPSNL------- 313
Query: 505 SGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDS 564
G S + I G++ P L + PV+ KL ++VD + +GP+++
Sbjct: 314 -GKSKEGYMDYPIPGEETVHPLL-------YSGALSRPVRTPKLQRYVDRMERIRGPKNT 469
Query: 565 PGHWLVTGARLMLDKGKICLWAKFSLLN 592
PG+W V+GA+L + GKI L K+SLL+
Sbjct: 470 PGYWAVSGAKLYVHNGKIYLLVKYSLLS 553
>TC92134 similar to GP|14209545|dbj|BAB56041. contains ESTs C72864(E2385)
AU082952(E2385)~similar to Arabidopsis thaliana
chromosome 1 F15D2.24~, partial (8%)
Length = 997
Score = 86.3 bits (212), Expect = 3e-17
Identities = 54/138 (39%), Positives = 76/138 (54%), Gaps = 2/138 (1%)
Frame = +3
Query: 19 LTSDFRLKFCKGEE--RLVILNETERRELTVPGFGPVTDVSVDIKCDKGDLTRYQSDILS 76
L D RL +CKG R+V ++E +R+L + V +VS DI+ + R S + S
Sbjct: 585 LIFDTRLLYCKGGSGSRVVEIDEQYQRDLFLYDDVVVPNVSRDIRSFPEPMGRLSSGVCS 764
Query: 77 FTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYVIKLFNVHIDR 136
F +M + FN K+SI G P G FN+ F F GS DAA TK L DG+ I L V + +
Sbjct: 765 FQEMVDYFNHKASISGSFPLGSFNSAFSF-TGSKHVDAAATKTLSSDGFYIPLAKVQLQK 941
Query: 137 YPLVLSKQVIEAVPSSWD 154
L+L + V A+P +WD
Sbjct: 942 IDLMLQENVKRAIPVNWD 995
>BI273040 similar to GP|6721157|gb| putative GDSL-motif lipase/acylhydrolase
{Arabidopsis thaliana}, partial (60%)
Length = 692
Score = 30.0 bits (66), Expect = 2.6
Identities = 30/106 (28%), Positives = 40/106 (37%), Gaps = 6/106 (5%)
Frame = +2
Query: 297 GRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHS----PSL 352
G + + N YL PY +DY HR N L + + + S P L
Sbjct: 62 GDSLVDNGNNNYLATTARADSYPYGIDYPTHRATGRFSNGLNMPDLISERIGSQPTLPYL 241
Query: 353 SLNLMGPKLYV--NTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQY 396
S L G L V N A +G TG++ F R+ LQY
Sbjct: 242 SPELNGEALLVGANFASAGIGILNDTGIQFF----NIIRITRQLQY 367
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,696,060
Number of Sequences: 36976
Number of extensions: 272802
Number of successful extensions: 1452
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 1376
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1409
length of query: 595
length of database: 9,014,727
effective HSP length: 102
effective length of query: 493
effective length of database: 5,243,175
effective search space: 2584885275
effective search space used: 2584885275
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0069.5


