

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
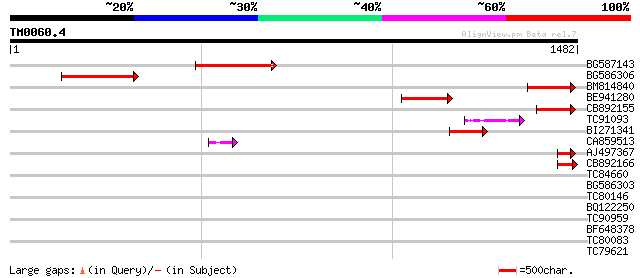
Query= TM0060.4
(1482 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F2... 219 6e-57
BG586306 weakly similar to GP|15128241|db helicase-like protein ... 165 1e-40
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 152 7e-37
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 152 9e-37
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 99 1e-20
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 82 2e-15
BI271341 80 5e-15
CA859513 58 2e-08
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 57 5e-08
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 56 9e-08
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 38 0.026
BG586303 weakly similar to PIR|A96586|A96 hypothetical protein F... 38 0.034
TC80146 36 0.097
BQ122250 32 2.4
TC90959 similar to GP|2117310|emb|CAB09116.1 hypothetical serine... 31 3.1
BF648378 similar to GP|9294677|dbj| gene_id:F14O13.28~pir||T2263... 31 4.1
TC80083 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 31 4.1
TC79621 MtN3 26 9.4
>BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (12%)
Length = 717
Score = 219 bits (558), Expect = 6e-57
Identities = 107/212 (50%), Positives = 143/212 (66%), Gaps = 1/212 (0%)
Frame = -3
Query: 486 SAYDRPDLACRVFRMKLDQLMKNLKKGKFFGQAISWMYTIEFQKRGLPHAHILLWLSTKD 545
S D+PD+ CRVF+MKLD+L+K+ KKG FF + ++ IEFQKRGL HAHILLW
Sbjct: 697 SPNDKPDIECRVFKMKLDELLKDFKKGTFFKPYTTALHRIEFQKRGLRHAHILLWFGNSS 518
Query: 546 KLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHGPCGVNRTNSPCMKNGRCSKFFPKN 605
+ S E +D +I AELP+ K P Y V+ +M+HGPCGV SPCM+N C+K +P+
Sbjct: 517 RTPSSEEVDEIISAELPNKKQDPEAYNLVTKHMIHGPCGVINPKSPCMENNVCTKKYPRP 338
Query: 606 FVEKTSFDSDGYPVYRRR-NTGISTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKS 664
+ TS D GY +YRRR N + G L+N F+VP+N KLL KY+AHIN+E+CN++
Sbjct: 337 YNGSTSIDKSGYVLYRRR*NETEHVVKNGAILNNTFIVPHNIKLLKKYEAHINMEWCNRT 158
Query: 665 NCIKYLFKYINKGVDRVTMSMSVTQSSNEETT 696
+ +KYLFKYI KGVDRV++ V + N TT
Sbjct: 157 SAVKYLFKYITKGVDRVSV---VIEKGNPATT 71
>BG586306 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (2%)
Length = 667
Score = 165 bits (417), Expect = 1e-40
Identities = 91/201 (45%), Positives = 126/201 (62%)
Frame = +2
Query: 135 FIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAEGDNPKFAQL 194
F D IR YNS+ AFTSIG K+D SV N G I GQ +HRI SL+P +G P++ Q+
Sbjct: 11 FRDTIRVYNSVLAFTSIGMKMDYSVVNAPGRYTIRIQGQTHHRIDSLIPRQGRPPEYLQI 190
Query: 195 NIYDTRNEFENRLNHLSDSTGKCSLNSDLVVELMAMVDEFNVLAKSFRRVRDHVQEANSN 254
I+DT NE NRLN + ++ + +L+ + L+ M+DE N LAK FRR RD+ + +
Sbjct: 191 YIFDTGNEVRNRLNAMGQTSTEGNLDETTLARLIEMIDENNCLAKLFRRARDY-YKGSGQ 367
Query: 255 QVALRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTSDCGRDIILRTSSGNLQRIYDTHSS 314
+ +RL + K Y+LP+ EVA LIV D ++ RDI+++ S LQ+I D HS
Sbjct: 368 EFNIRLLSDK-GKGKEYDLPSTSEVAGLIVGDMSSTIGVRDIVVQFQSDTLQQIRDYHSL 544
Query: 315 FLPLQYPLIFPYGEEGFSDEI 335
++ LQYPL+FPYGE GF EI
Sbjct: 545 YMSLQYPLLFPYGEYGFHPEI 607
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza sativa
(japonica cultivar-group)}, partial (6%)
Length = 733
Score = 152 bits (385), Expect = 7e-37
Identities = 75/127 (59%), Positives = 100/127 (78%)
Frame = +3
Query: 1353 NGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAM 1412
+GTR+++ L NV+ VI GTH G +I RM+L+P+ ++ I F+R QFPL+LSFAM
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAM 206
Query: 1413 TINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVF 1472
TINKSQG+TL+ VGLYLP PVF+HGQLYVA+SRVK+R+GLKILI +E+ S T N+V+
Sbjct: 207 TINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVY 386
Query: 1473 KEVFQRI 1479
+EVFQ+I
Sbjct: 387 QEVFQKI 407
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 152 bits (384), Expect = 9e-37
Identities = 77/134 (57%), Positives = 97/134 (71%)
Frame = -1
Query: 1023 VYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDS 1082
+Y YGGT KTF+W L+ LRSE +I+L ASSGI +LL+ GGRTAHS F IP DE S
Sbjct: 403 LYDYGGTEKTFIWRALSAALRSEGEIVLACASSGIDALLMPGGRTAHSRFGIPFIIDETS 224
Query: 1083 CCGIVQGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKV 1142
CG+ P A L+ + LIIWDEAPM+ ++ FEALDR+LRD+++ + + PFGGKV
Sbjct: 223 MCGVTPNIPLASLVIKAKLIIWDEAPMMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKV 44
Query: 1143 VVLGGDFRQILPVI 1156
VVLGGDFRQIL V+
Sbjct: 43 VVLGGDFRQILLVM 2
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1 [imported]
- Arabidopsis thaliana, partial (4%)
Length = 572
Score = 99.0 bits (245), Expect = 1e-20
Identities = 52/102 (50%), Positives = 67/102 (64%)
Frame = -2
Query: 1378 GRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHG 1437
GR V RM D S + + + FAMTINKSQG++L H+G+YLP+ VFSHG
Sbjct: 454 GRSVQYKRMKGADIDMSQGVSQMSTHESVEVYFAMTINKSQGQSLKHIGVYLPSSVFSHG 275
Query: 1438 QLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
QLYVA+SRV +R GLKILI N+D VT N+V++EVF +
Sbjct: 274 QLYVALSRVTSREGLKILISNDDGEDDCVTSNVVYREVFHNL 149
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 82.0 bits (201), Expect = 2e-15
Identities = 55/155 (35%), Positives = 80/155 (51%)
Frame = +3
Query: 1190 FSNSETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY 1249
F N + VE K FAE I G + + ND A++ P +P+ IV++ Y
Sbjct: 279 FPNIPRTKVE-FKTFAE----ILTGKMSEPNDSYAEVDTPP-------GDPIDAIVQSTY 422
Query: 1250 PEIMENVGCAKYYEDKAILAPTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIE 1309
P ++ ++ + +AIL T + VD +N YVL L PG ER S+ S DV
Sbjct: 423 PNLVSQYNNEQFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSANR--SEVNDVQ-A 593
Query: 1310 ADWITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRN 1344
D I EFL ++ S +P+HKL LK G P+ML+R+
Sbjct: 594 FDAIPPEFLQSLKTSDLPNHKLTLKVGTPIMLLRD 698
>BI271341
Length = 468
Score = 80.5 bits (197), Expect = 5e-15
Identities = 41/101 (40%), Positives = 67/101 (65%)
Frame = +1
Query: 1149 FRQILPVIPKGSRAEIVMATINSSRLWRFCKVLTLTENMRLFSNSETSDVEKIKVFAEWV 1208
FR ILPVIP+GSR++I+ ATINSS + C+V+ L +NM L N ++S+ + + F++ +
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPEFEQFSK*I 180
Query: 1209 LDIGDGVLGDYNDGDADIKVPKDTLVQQSSNPVADIVRAIY 1249
L +GDG + + ND ADI +P + L+ + + IV++ Y
Sbjct: 181 LKVGDGKIYEPNDSYADIDIPPELLISNYDDSLQTIVQSTY 303
>CA859513
Length = 363
Score = 58.2 bits (139), Expect = 2e-08
Identities = 31/75 (41%), Positives = 40/75 (53%)
Frame = +1
Query: 520 SWMYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMV 579
S +Y EF+K LPHAH+L H + D+ + AEL DP LYQ V + M+
Sbjct: 130 SHIYVAEFEKCDLPHAHMLF--------HRADNYDTFVSAELSDPVEQLRLYQTVVSVMI 285
Query: 580 HGPCGVNRTNSPCMK 594
HGP G N PCM+
Sbjct: 286 HGPYGPFNNNVPCMR 330
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 57.0 bits (136), Expect = 5e-08
Identities = 26/49 (53%), Positives = 38/49 (77%)
Frame = +1
Query: 1431 NPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
N FS+G+LYVA+SRV +R GLKIL+ +ED + + T N+V+KEVF+ +
Sbjct: 10 NQFFSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 56.2 bits (134), Expect = 9e-08
Identities = 30/50 (60%), Positives = 39/50 (78%)
Frame = -1
Query: 1433 VFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRIYSS 1482
VFSHGQLYVAISRV +R GLKIL+ +E+ D T N+V+K VFQ +++S
Sbjct: 274 VFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQNVWTS 128
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 38.1 bits (87), Expect = 0.026
Identities = 16/24 (66%), Positives = 20/24 (82%)
Frame = +2
Query: 1426 GLYLPNPVFSHGQLYVAISRVKTR 1449
G+YLP P+F HG LYVA+SRV +R
Sbjct: 83 GMYLPQPIF*HG*LYVALSRVTSR 154
>BG586303 weakly similar to PIR|A96586|A96 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (3%)
Length = 698
Score = 37.7 bits (86), Expect = 0.034
Identities = 17/27 (62%), Positives = 20/27 (73%)
Frame = -3
Query: 489 DRPDLACRVFRMKLDQLMKNLKKGKFF 515
DR DL RVF+MKLD+LM + KG FF
Sbjct: 663 DRHDLKSRVFKMKLDELMSDFNKGIFF 583
>TC80146
Length = 476
Score = 36.2 bits (82), Expect = 0.097
Identities = 14/33 (42%), Positives = 25/33 (75%)
Frame = -3
Query: 1408 LSFAMTINKSQGKTLSHVGLYLPNPVFSHGQLY 1440
+ + TINKS+ ++LS++ +YL P+FSH ++Y
Sbjct: 465 IPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>BQ122250
Length = 439
Score = 31.6 bits (70), Expect = 2.4
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 4/53 (7%)
Frame = +1
Query: 528 QKRGLPHAHILLWLSTKDKLHSIELI----DSVICAELPDPKVYPLLYQCVSN 576
Q RG + HI LW K+K+HS+ + + C E P + P+L QCVSN
Sbjct: 292 QFRGHLNNHIFLWNCHKNKMHSLFVSPCDHSVIACLEYP---IAPML-QCVSN 438
>TC90959 similar to GP|2117310|emb|CAB09116.1 hypothetical serine-rich
protein (52 consecutive serines); contains WSC domain;
possibly involved, partial (16%)
Length = 961
Score = 31.2 bits (69), Expect = 3.1
Identities = 12/32 (37%), Positives = 20/32 (62%)
Frame = -3
Query: 832 QRGCTSFEDLRSVNNHVHDTFRQACLALSLLK 863
+R CTSF DLR++ N +H R+ C +++
Sbjct: 740 RRYCTSFADLRNLQNWIH*NLRKECWIKKMMR 645
>BF648378 similar to GP|9294677|dbj| gene_id:F14O13.28~pir||T22638~similar to
unknown protein {Arabidopsis thaliana}, partial (8%)
Length = 651
Score = 30.8 bits (68), Expect = 4.1
Identities = 18/49 (36%), Positives = 26/49 (52%), Gaps = 12/49 (24%)
Frame = +2
Query: 1171 SSRLWRFC------------KVLTLTENMRLFSNSETSDVEKIKVFAEW 1207
SS LWRFC +V +T+N+ S+SET D+ IK+ E+
Sbjct: 71 SSYLWRFCISEDYFVVVGVVRV*EMTQNIFDGSDSETEDISNIKIDEEY 217
>TC80083 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (4%)
Length = 1269
Score = 30.8 bits (68), Expect = 4.1
Identities = 23/63 (36%), Positives = 32/63 (50%), Gaps = 4/63 (6%)
Frame = +3
Query: 933 EHLKQLCLMEIEKLLMINGRSLKDFDNMPCVDSNILIQYGNILLFN----ELNFDTVEMS 988
EH L L +++L + + RSLK F + C +NI NI L N EL F +S
Sbjct: 426 EHFPPLGLASLKELNLSSCRSLKSFPELLCKMTNI----DNIWLCNTSIGELPFSFQNLS 593
Query: 989 KLH 991
+LH
Sbjct: 594 ELH 602
>TC79621 MtN3
Length = 1117
Score = 26.2 bits (56), Expect(2) = 9.4
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +3
Query: 533 PHAHILLWLSTKDKLHSI 550
PH H L+W+ +D LH I
Sbjct: 324 PHYHKLIWMCGRDHLHHI 377
Score = 21.6 bits (44), Expect(2) = 9.4
Identities = 9/20 (45%), Positives = 11/20 (55%)
Frame = +1
Query: 574 VSNYMVHGPCGVNRTNSPCM 593
V+NY VHGP V C+
Sbjct: 469 VTNYAVHGPLRVQVLGWVCV 528
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,567,452
Number of Sequences: 36976
Number of extensions: 728913
Number of successful extensions: 3289
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 3256
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3285
length of query: 1482
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1374
effective length of database: 5,021,319
effective search space: 6899292306
effective search space used: 6899292306
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0060.4


