

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
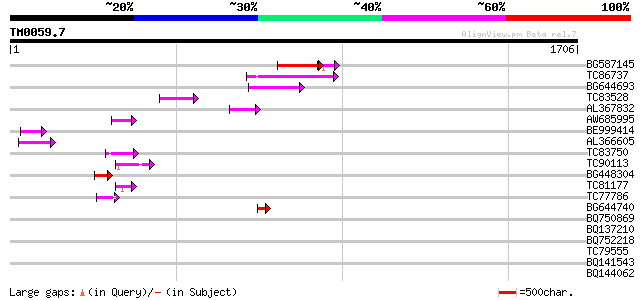
Query= TM0059.7
(1706 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 166 5e-41
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 147 4e-35
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 81 3e-15
TC83528 66 1e-10
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 58 4e-08
AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - A... 55 2e-07
BE999414 53 1e-06
AL366605 51 4e-06
TC83750 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 49 2e-05
TC90113 similar to PIR|C86301|C86301 arginine/serine-rich protei... 48 3e-05
BG448304 47 8e-05
TC81177 similar to GP|15293081|gb|AAK93651.1 unknown protein {Ar... 46 1e-04
TC77786 45 3e-04
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 44 4e-04
BQ750869 weakly similar to GP|21205303|db alpha-acetolactate syn... 43 0.001
BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall struc... 41 0.003
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 41 0.003
TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishman... 41 0.005
BQ141543 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 40 0.008
BQ144062 weakly similar to SP|P03211|EBN1_ EBNA-1 nuclear protei... 40 0.008
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 166 bits (421), Expect = 5e-41
Identities = 77/136 (56%), Positives = 106/136 (77%)
Frame = +2
Query: 807 MHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVK 866
MH DD EKTAFITDRGTYCY ++PFGLKNA +TYQR++ R+F D +G ++EVYIDD++VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 867 TPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIM 926
+ + DH L F+ L ++ M+LNP KCTFGV SG+FLGY++T +GIE+NP + AI+
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 927 NMKSPRTVKEVQQLAG 942
++ SP+ +EVQ+L G
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 46.6 bits (109), Expect = 8e-05
Identities = 25/74 (33%), Positives = 40/74 (53%), Gaps = 7/74 (9%)
Frame = +3
Query: 925 IMNMKSPRTVKEVQQLA-------GRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEA 977
I++ P VQ++A GR+AA+ RF+ ++ + LP Y LL F W +
Sbjct: 336 ILSRSPPY*TSLVQRIAERSSDSRGRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKC 515
Query: 978 DAAFTQLKEILSSP 991
+ AF QLK+ L++P
Sbjct: 516 EEAFEQLKQYLTTP 557
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 147 bits (370), Expect = 4e-35
Identities = 88/278 (31%), Positives = 145/278 (51%), Gaps = 2/278 (0%)
Frame = +1
Query: 714 MSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKD 773
MS ++ V+++ +LL G I+ + V+ V+K G R CVDY LN KD
Sbjct: 487 MSRQELLVLKKTLEDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAITKKD 663
Query: 774 PFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGL 833
+PLP I + +G + + +D + ++++ + +D+EKTAF T G + + + PFGL
Sbjct: 664 RYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGL 843
Query: 834 KNAVATYQRMMTRIFGDLMGKSVEVYIDDIIV-KTPKGGDHAADLAVVFEQLKKHNMRLN 892
A AT+QR + + + + V YIDD+++ T DH A + V +L + L+
Sbjct: 844 TGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLD 1023
Query: 893 PDKCTFGVRSGKFLGYMLT-NRGIELNPDKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFL 951
P KC F V + K++G++LT +G+ +P K AI + P +VK + G F+
Sbjct: 1024PKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFI 1203
Query: 952 PKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEILS 989
P + PL L + F W AE +AAFT+LK + +
Sbjct: 1204PGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFA 1317
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 81.3 bits (199), Expect = 3e-15
Identities = 53/168 (31%), Positives = 84/168 (49%)
Frame = +2
Query: 718 KQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPL 777
K +V++ Q +LL G I+ Y + V+ +KK +G RM +DY LN K +PL
Sbjct: 107 KLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRMSIDYPQLNNVNIKIKYPL 283
Query: 778 PSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAV 837
P ID L DN G ++ +D G +Q + +D KTAF G Y ++ FG N
Sbjct: 284 PLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRIRYGHYEILVMSFG*TNPP 463
Query: 838 ATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQLK 885
+ +M R+F D + V V+ +DI++ + +H L + + LK
Sbjct: 464 MAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRLALKVLK 607
>TC83528
Length = 555
Score = 65.9 bits (159), Expect = 1e-10
Identities = 40/121 (33%), Positives = 60/121 (49%), Gaps = 4/121 (3%)
Frame = +3
Query: 451 PIVIEALIGNG-KIRRTLIDTGSSADIMFYDAYKSLGLSVKDLLPYDHDLVGFTGDRVLP 509
P+VI+ I N + R ++ S DI+++ A+ + L L P L G G +
Sbjct: 180 PMVIKLQINNNFSVLRVFVNPMSKVDILYWSAFLKMKLQESMLKPCQGFLKGTFGKGLPV 359
Query: 510 LGYFDTCLSLGDHRICRTIKARFLVVECP---TAYNAILGRPSLNTFRAIISTHHLMLKY 566
GY D + G +TIK R+ VVE P + YN +LG PSL +A++S +KY
Sbjct: 360 KGYIDLDTTFGKGENTKTIKVRYFVVESPPSVSIYNVVLGWPSLKDLKAVLSVAEFTIKY 539
Query: 567 P 567
P
Sbjct: 540 P 542
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 57.8 bits (138), Expect = 4e-08
Identities = 30/96 (31%), Positives = 51/96 (52%)
Frame = -1
Query: 660 LPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQ 719
L + +++ +LL +FA S EDMPG+DP HR+ + PV K R+ +
Sbjct: 288 LEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEHRIPTKPECPPVR*KLRRTHPDMA 109
Query: 720 QVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANG 755
I+ + + + AG + V+Y W++N+V V K +G
Sbjct: 108 LKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
>AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - African
clawed frog, partial (5%)
Length = 516
Score = 55.5 bits (132), Expect = 2e-07
Identities = 37/81 (45%), Positives = 44/81 (53%), Gaps = 5/81 (6%)
Frame = +1
Query: 306 SRYVAVSTGALPRPRSPPPRRTPTPPRHR--DQTPPKRRSAERRDKTPPRRRSPERR--- 360
SR A + +LP PR PP R PPR ++P RRS R +P RRRSP RR
Sbjct: 175 SRRTAPAPRSLPAPRMRPPLRRLPPPRRPLPRRSPAPRRSRARLSPSPRRRRSPPRRPPP 354
Query: 361 RRSRERRRSPDRHGRDEVRRQ 381
RRSR RRS R R +RR+
Sbjct: 355 RRSRALRRSRARPRRSPMRRR 417
Score = 37.4 bits (85), Expect = 0.050
Identities = 40/143 (27%), Positives = 49/143 (33%), Gaps = 9/143 (6%)
Frame = +1
Query: 316 LPRPR-----SPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERR----RRSRER 366
LPRPR S RR PP +T P RS PP RR P R RRS
Sbjct: 106 LPRPRRRRRSSRRRRRRSPPPWRSRRTAPAPRSLPAPRMRPPLRRLPPPRRPLPRRSPAP 285
Query: 367 RRSPDRHGRDEVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLH 426
RRS R RR+ P R +R + + RPR +
Sbjct: 286 RRSRARLSPSPRRRR------------------SPPRRPPPRRSRALRRSRARPRRSPMR 411
Query: 427 PPRQKVAITFTEDDYGEDTGEED 449
R K A + E+ +D
Sbjct: 412 RRRMKPAAAENKRKRDEEENGDD 480
>BE999414
Length = 613
Score = 52.8 bits (125), Expect = 1e-06
Identities = 24/79 (30%), Positives = 39/79 (48%)
Frame = +3
Query: 32 PKVKLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLHDW 91
P++ Y+G DP EH+ + Y + C+ F +L +G M WF+NL NS+ W
Sbjct: 3 PQMDSYDGT*DPDEHMENIEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSW 182
Query: 92 NGVVTCFLAQYSSVRSIPK 110
+ F ++ R+ PK
Sbjct: 183 GDLCHEFTTHFTVSRTQPK 239
>AL366605
Length = 422
Score = 50.8 bits (120), Expect = 4e-06
Identities = 28/112 (25%), Positives = 50/112 (44%)
Frame = -3
Query: 26 PQGWARPKVKLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPS 85
P+ + P Y G + PQ H+ +V M +D L CF SL + W+ +L
Sbjct: 342 PKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLSK 163
Query: 86 NSLHDWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEA 137
N +H ++ + F + Y + + L + QK++ES + + R+ A
Sbjct: 162 NDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAA 7
>TC83750 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (68%)
Length = 772
Score = 48.5 bits (114), Expect = 2e-05
Identities = 38/101 (37%), Positives = 49/101 (47%), Gaps = 1/101 (0%)
Frame = +2
Query: 287 DECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRS-PPPRRTPTPPRHRDQTPPKRRSAE 345
D+ + D + EL G +R S G R RS PPR +P R P+ RS
Sbjct: 179 DDRRDALDAIREL--DGHFARECNSSRGGSGRRRSRSPPRFRRSPSYGRRSYSPRERSPR 352
Query: 346 RRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRRQHGSNI 386
RR +P RRRSP RRRS R P R GR++ +G+ I
Sbjct: 353 RRSPSP-RRRSPSPRRRSYSRSPPPYR-GREDATHANGNGI 469
Score = 30.0 bits (66), Expect = 8.0
Identities = 20/50 (40%), Positives = 22/50 (44%), Gaps = 2/50 (4%)
Frame = +2
Query: 317 PRPRSPPPRRTPTPPRHR--DQTPPKRRSAERRDKTPPRRRSPERRRRSR 364
PR RSP PRR PR R ++PP R R D T RSR
Sbjct: 347 PRRRSPSPRRRSPSPRRRSYSRSPPPYRG--REDATHANGNGIRAAHRSR 490
>TC90113 similar to PIR|C86301|C86301 arginine/serine-rich protein
[imported] - Arabidopsis thaliana, partial (70%)
Length = 1259
Score = 48.1 bits (113), Expect = 3e-05
Identities = 50/143 (34%), Positives = 61/143 (41%), Gaps = 26/143 (18%)
Frame = +1
Query: 318 RPRSPPPRR--------------TPTPPRHRDQTPP-KRRSAERRDKTPPRRRSPERRRR 362
R SPPPRR +P P R R PP +RRS RR ++PPRR SP RRR
Sbjct: 808 RSPSPPPRRLRSPARVSPRRMRGSPGPGRRRSPPPPPRRRSPPRRARSPPRR-SPVGRRR 984
Query: 363 SRE------RRRS---PDRHGRDEVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTRVI 413
SR R RS R GR +RR S+ D S + S+ RS R
Sbjct: 985 SRSPIRRSARSRSRSFSPRRGRPPMRRGRSSSYSDSPS------PRKVSRRSKSRSPRRP 1146
Query: 414 MS--AAGRPRPGSLHPPRQKVAI 434
+ A+ S PP +K I
Sbjct: 1147LKGRASSNSSSSSSPPPARKPXI 1215
Score = 42.4 bits (98), Expect = 0.002
Identities = 31/80 (38%), Positives = 35/80 (43%), Gaps = 20/80 (25%)
Frame = +1
Query: 321 SPPPRRTPTPPRHRDQTPPKRR------------------SAERRDKTPPRRRSPERRRR 362
+PP RR +P R R +PP RR R PPRRRSP RR R
Sbjct: 769 TPPRRRPVSPGRGRSPSPPPRRLRSPARVSPRRMRGSPGPGRRRSPPPPPRRRSPPRRAR 948
Query: 363 SRERRRSP--DRHGRDEVRR 380
S RRSP R R +RR
Sbjct: 949 S-PPRRSPVGRRRSRSPIRR 1005
Score = 41.2 bits (95), Expect = 0.003
Identities = 23/52 (44%), Positives = 27/52 (51%), Gaps = 1/52 (1%)
Frame = +1
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRR-SPERRRRSRERRR 368
RPR PRR P P R P+R + RR ++P RR SP RRR RR
Sbjct: 601 RPRESSPRRKPPPSPRRRSPVPRRAGSPRRPESPRRRADSPVRRRLDSPYRR 756
Score = 40.8 bits (94), Expect = 0.005
Identities = 47/147 (31%), Positives = 52/147 (34%), Gaps = 43/147 (29%)
Frame = +1
Query: 317 PRPRSPPPRRTPTP-----PRHR-----------------DQTPPKRRSAE----RRDKT 350
PR RSP PRR +P PR R TPP+RR R
Sbjct: 643 PRRRSPVPRRAGSPRRPESPRRRADSPVRRRLDSPYRRGAGDTPPRRRPVSPGRGRSPSP 822
Query: 351 PPRR-RSPER-------------RRRS---RERRRSPDRHGRDEVRRQHGSNIVDVGSIA 393
PPRR RSP R RRRS RRRSP R R RR S
Sbjct: 823 PPRRLRSPARVSPRRMRGSPGPGRRRSPPPPPRRRSPPRRARSPPRR----------SPV 972
Query: 394 GGWAAGGPTNNSQKRSTRVIMSAAGRP 420
G + P S + +R GRP
Sbjct: 973 GRRRSRSPIRRSARSRSRSFSPRRGRP 1053
Score = 38.5 bits (88), Expect = 0.022
Identities = 26/79 (32%), Positives = 33/79 (40%), Gaps = 14/79 (17%)
Frame = +1
Query: 317 PRPRSPPPRRTPTPPRHRDQTP-------------PKRRSAERRDKTPPRRRSPERRRRS 363
PR ++ PP + P R +T P+ S R+ PRRRSP RR
Sbjct: 499 PRQKASPPPKAVAPKRDAPRTDNAGAEVEKDGPKRPRESSPRRKPPPSPRRRSPVPRRAG 678
Query: 364 RERR-RSPDRHGRDEVRRQ 381
RR SP R VRR+
Sbjct: 679 SPRRPESPRRRADSPVRRR 735
Score = 37.0 bits (84), Expect = 0.065
Identities = 26/61 (42%), Positives = 32/61 (51%), Gaps = 7/61 (11%)
Frame = +1
Query: 317 PRPRSPP------PRRTPTPPRHRDQTPPKRRSAER-RDKTPPRRRSPERRRRSRERRRS 369
PR RSPP PRR+P R R ++P +R + R R +P R R P RR RS S
Sbjct: 916 PRRRSPPRRARSPPRRSPV-GRRRSRSPIRRSARSRSRSFSPRRGRPPMRRGRSSSYSDS 1092
Query: 370 P 370
P
Sbjct: 1093P 1095
Score = 33.1 bits (74), Expect = 0.94
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Frame = +1
Query: 312 STGALPRPRSPPP--RRTPTPPRHRDQTPPKRRSAE-RRDKTPPRRRSPER 359
S + R RSPPP R++ P R ++PP + +R PPR+ SP R
Sbjct: 103 SRSSKSRSRSPPPPRRKSSAEPARRGRSPPLPPPPQSKRPSPPPRKPSPVR 255
>BG448304
Length = 637
Score = 46.6 bits (109), Expect = 8e-05
Identities = 22/53 (41%), Positives = 33/53 (61%), Gaps = 1/53 (1%)
Frame = +2
Query: 256 LRDRPPPLLTRGDK-LDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSR 307
L PP T+ +K D ++C FH H TD+C++LKD +E LI+ GRL++
Sbjct: 50 LGSNPPKTPTQENKGTDKTKYCRFHKCHRHLTDDCIHLKDTIEILIQRGRLNQ 208
>TC81177 similar to GP|15293081|gb|AAK93651.1 unknown protein {Arabidopsis
thaliana}, partial (28%)
Length = 632
Score = 46.2 bits (108), Expect = 1e-04
Identities = 32/80 (40%), Positives = 39/80 (48%), Gaps = 16/80 (20%)
Frame = +1
Query: 318 RPRSPP-----PRRTPTPPRHRDQT---------PPKRRSAERRDKTPPRRRSPERRRRS 363
R RSPP PR + +PPRHR ++ PPKRR R RR S E + S
Sbjct: 283 RRRSPPRYSRSPRYSRSPPRHRSRSRGSRDYHSPPPKRREYSRSASPEDRRHSREGSQHS 462
Query: 364 RERR--RSPDRHGRDEVRRQ 381
RER RSP ++G R Q
Sbjct: 463 REREYSRSPPKNGDARSRSQ 522
>TC77786
Length = 1094
Score = 44.7 bits (104), Expect = 3e-04
Identities = 22/68 (32%), Positives = 37/68 (54%)
Frame = +3
Query: 261 PPLLTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPR 320
P LL + D + C +H S GH+T+EC L++ +E+LI+ +L R++ R
Sbjct: 780 PQLLK*SARTDKAK*CCYHRSRGHDTEECFQLREAIEDLIKKVKLGRFIDKG------HR 941
Query: 321 SPPPRRTP 328
PR++P
Sbjct: 942 KNSPRKSP 965
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 44.3 bits (103), Expect = 4e-04
Identities = 18/39 (46%), Positives = 26/39 (66%)
Frame = -1
Query: 747 VVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVD 785
++ V+K +G +RMC+DY NK K+ +PLP ID L D
Sbjct: 232 LLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>BQ750869 weakly similar to GP|21205303|db alpha-acetolactate synthase
{Staphylococcus aureus subsp. aureus MW2}, partial (18%)
Length = 732
Score = 42.7 bits (99), Expect = 0.001
Identities = 29/77 (37%), Positives = 36/77 (46%), Gaps = 7/77 (9%)
Frame = +3
Query: 320 RSPPPRRTPTPPRHRDQTPPKRRSAER-RDKTPPRRRSPERRRRSRER------RRSPDR 372
R P RR +P R D + P RR+A R R + RRR+P RR+ R R S
Sbjct: 336 RRPRRRRQESPGRQEDPSEPARRAAARARHQKGHRRRTPRPDRRAHARCLPDRHRPSSGS 515
Query: 373 HGRDEVRRQHGSNIVDV 389
HGR+ R H DV
Sbjct: 516 HGREPAHRHHDVRGQDV 566
>BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall structural
protein 1 precursor. [Petunia] {Petunia hybrida},
partial (13%)
Length = 990
Score = 41.2 bits (95), Expect = 0.003
Identities = 31/86 (36%), Positives = 38/86 (44%), Gaps = 5/86 (5%)
Frame = -2
Query: 312 STGALPRPRS---PPPRRTPTPPRHRDQTPPKRRSA--ERRDKTPPRRRSPERRRRSRER 366
S ALP R P PRR P P R T P+R + TPPR R P R R +
Sbjct: 857 SRAALPAQRRGLLPRPRRLPRPAACRACTSPRRLAMLPSPPRSTPPRPRRPRRARPTSLI 678
Query: 367 RRSPDRHGRDEVRRQHGSNIVDVGSI 392
R + RHG QH + +V S+
Sbjct: 677 RDASQRHG------QHSARVVSA*SV 618
Score = 30.8 bits (68), Expect = 4.7
Identities = 16/42 (38%), Positives = 20/42 (47%)
Frame = -1
Query: 306 SRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERR 347
SRY AV++ P +PPP R P R Q P+ RR
Sbjct: 759 SRYAAVASALNAPPPAPPPTRPPYVTHTRRQPTPRTALRTRR 634
Score = 30.4 bits (67), Expect = 6.1
Identities = 23/78 (29%), Positives = 28/78 (35%)
Frame = -1
Query: 304 RLSRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRS 363
RL+R A S + P P PPP + PR P + + PP P R
Sbjct: 861 RLAR-CAPSAASRPAPTPPPPAASRRLPRVHVAAPSRYAAVASALNAPPPAPPPTRPPYV 685
Query: 364 RERRRSPDRHGRDEVRRQ 381
RR P RRQ
Sbjct: 684 THTRRQPTPRTALRTRRQ 631
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/64 (40%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = -3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKR-RSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRD 376
R R P ++ PR + P +R R RR K PR+R +RRSR RRR P R R
Sbjct: 584 RRRRRPGKKLQRRPRRPGRQPIRRPRRRRRRPKRRPRKRRGRLKRRSRRRRRRPKRRPR- 408
Query: 377 EVRR 380
+RR
Sbjct: 407 RIRR 396
Score = 35.4 bits (80), Expect = 0.19
Identities = 25/65 (38%), Positives = 33/65 (50%), Gaps = 7/65 (10%)
Frame = -3
Query: 318 RPRSP--PPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERR-RRSRERRRSPDR-- 372
RPR P P R P R R + P++R + ++ RRR P+RR RR R R + R
Sbjct: 548 RPRRPGRQPIRRPRRRRRRPKRRPRKRRGRLKRRSRRRRRRPKRRPRRIRRSRIAKTRNQ 369
Query: 373 --HGR 375
HGR
Sbjct: 368 LCHGR 354
Score = 33.5 bits (75), Expect = 0.72
Identities = 23/54 (42%), Positives = 25/54 (45%), Gaps = 2/54 (3%)
Frame = -3
Query: 324 PRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERR--RRSRERRRSPDRHGR 375
PRR R R + PKRR K R R P R+ RR R RRR P R R
Sbjct: 635 PRRGRKRRRRRLRRRPKRRRRRPGKKLQRRPRRPGRQPIRRPRRRRRRPKRRPR 474
Score = 31.6 bits (70), Expect = 2.7
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = -3
Query: 341 RRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRR 380
RR +RR + RR RRR ++ +R P R GR +RR
Sbjct: 632 RRGRKRRRRRLRRRPKRRRRRPGKKLQRRPRRPGRQPIRR 513
>TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishmania major},
partial (1%)
Length = 1410
Score = 40.8 bits (94), Expect = 0.005
Identities = 32/76 (42%), Positives = 36/76 (47%), Gaps = 10/76 (13%)
Frame = +3
Query: 318 RPRS-PPPRRTPTPPRHRDQTPPKRRSAERRDKTPPR----RRSPERRRRSRERRRSPDR 372
RPR PPPRR RHR++ K R A RR + PPR RR R RRRS R
Sbjct: 516 RPRGHPPPRRRRAHRRHRNRALHKLRPAPRRRRGPPRLPQLRRGRHFSRVHLRRRRSFRR 695
Query: 373 HGRD-----EVRRQHG 383
G + RR HG
Sbjct: 696 GGLEPGHCRNRRRGHG 743
Score = 35.0 bits (79), Expect = 0.25
Identities = 35/123 (28%), Positives = 49/123 (39%), Gaps = 6/123 (4%)
Frame = +2
Query: 306 SRYVAVSTGALPRPRSPP-----PRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERR 360
SR + +T P P S P P R+P+ R + R+ PP P RR
Sbjct: 521 SRPSSPATTTRPPPTSKPSPTQTPSRSPSSARTPSTSSTPTRAPLLSSPPPPPPFFPARR 700
Query: 361 RRSRERRRSPDRHGRDEVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTR-VIMSAAGR 419
+R +SP R + ++ +A AAG T+ + R TR SAA R
Sbjct: 701 PGTRSLPKSPPRPRPARLPSARRASPRGGRPLAAAAAAGATTSTTASRPTRSARTSAARR 880
Query: 420 PRP 422
RP
Sbjct: 881 GRP 889
Score = 34.7 bits (78), Expect = 0.32
Identities = 20/50 (40%), Positives = 25/50 (50%)
Frame = +3
Query: 323 PPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDR 372
PP+R R R PP+RR A RR R R+ + R + RRR P R
Sbjct: 489 PPQRALRQRRPRGHPPPRRRRAHRRH----RNRALHKLRPAPRRRRGPPR 626
Score = 33.5 bits (75), Expect = 0.72
Identities = 26/83 (31%), Positives = 34/83 (40%), Gaps = 3/83 (3%)
Frame = -3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRER---RRSPDRHG 374
R RS R P + P+RR + PPRR+ RRR +RE+ RRS G
Sbjct: 799 RQRSATTRACPPGAWQSGRPWPRRRFRQ*PGSRPPRRKERRRRRWTREKWRPRRS*GSRG 620
Query: 375 RDEVRRQHGSNIVDVGSIAGGWA 397
RR G ++ WA
Sbjct: 619 GPRRRRGAGRSLCRARFRCRRWA 551
>BQ141543 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (15%)
Length = 1271
Score = 40.0 bits (92), Expect = 0.008
Identities = 29/91 (31%), Positives = 33/91 (35%), Gaps = 8/91 (8%)
Frame = -3
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKRRSAERR--------DKTPPRRRSPERRRRSRERRR 368
P P PPPRR TPP PP RR+ + D T PRR P R
Sbjct: 690 PPPARPPPRRPHTPPATSPPPPPSRRARTWQKPGRCAPYDLTVPRRAPPLRLPSISSPPH 511
Query: 369 SPDRHGRDEVRRQHGSNIVDVGSIAGGWAAG 399
P +R GS I G + G G
Sbjct: 510 FPPSPSTPRLRFGAGSRIFIGGGVCGSVQPG 418
>BQ144062 weakly similar to SP|P03211|EBN1_ EBNA-1 nuclear protein. [strain
B95-8 Human herpesvirus 4] {Epstein-barr virus},
partial (8%)
Length = 1390
Score = 40.0 bits (92), Expect = 0.008
Identities = 28/97 (28%), Positives = 39/97 (39%), Gaps = 1/97 (1%)
Frame = -1
Query: 275 FCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVA-VSTGALPRPRSPPPRRTPTPPRH 333
+ H H+ D L L + + R++ A PRPR R+P PPR
Sbjct: 850 YTSIHRHSAHHIDASLRLAQRDPSPT*SHRITTARARARPPPRPRPRRRHRARSPPPPRS 671
Query: 334 RDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSP 370
PP +RS+ + PP +P RR R SP
Sbjct: 670 PCGPPPPKRSSATSFRLPPPPSTPARREALLPRADSP 560
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.443
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,445,137
Number of Sequences: 36976
Number of extensions: 886812
Number of successful extensions: 10363
Number of sequences better than 10.0: 200
Number of HSP's better than 10.0 without gapping: 5692
Number of HSP's successfully gapped in prelim test: 279
Number of HSP's that attempted gapping in prelim test: 1475
Number of HSP's gapped (non-prelim): 8288
length of query: 1706
length of database: 9,014,727
effective HSP length: 110
effective length of query: 1596
effective length of database: 4,947,367
effective search space: 7895997732
effective search space used: 7895997732
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0059.7


