

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0008.5
(504 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
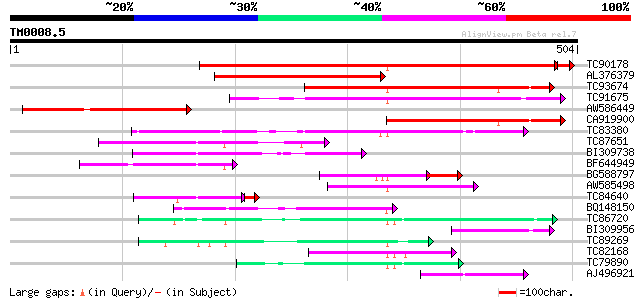
Score E
Sequences producing significant alignments: (bits) Value
TC90178 similar to GP|13958032|gb|AAK50769.1 polygalacturonase {... 560 e-161
AL376379 homologue to GP|13958032|gb polygalacturonase {Pisum sa... 288 3e-78
TC93674 similar to GP|15292729|gb|AAK92733.1 putative polygalact... 231 6e-61
TC91675 similar to GP|6624205|dbj|BAA88472.1 polygalacturonase {... 200 1e-51
AW586449 similar to PIR|B96631|B966 probable polygalacturonase F... 199 3e-51
CA919900 similar to GP|15292729|gb putative polygalacturonase PG... 172 4e-43
TC83380 weakly similar to GP|10176984|dbj|BAB10216. polygalactur... 162 4e-40
TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalact... 116 2e-26
BI309738 weakly similar to GP|15028105|gb putative polygalacturo... 109 3e-24
BF644949 similar to GP|15292729|gb putative polygalacturonase PG... 100 1e-21
BG588797 weakly similar to PIR|T08215|T08 polygalacturonase (EC ... 85 6e-21
AW585498 weakly similar to GP|10177065|db polygalacturonase-like... 97 2e-20
TC84640 similar to GP|10177065|dbj|BAB10507. polygalacturonase-l... 74 4e-17
BQ148150 weakly similar to GP|10185719|gb NTS1 protein {Nicotian... 86 4e-17
TC86720 similar to PIR|T47941|T47941 hypothetical protein F2A19.... 68 9e-12
BI309956 weakly similar to GP|7381227|gb|A polygalacturonase {Ly... 65 5e-11
TC89269 similar to PIR|T05614|T05614 hypothetical protein F9D16.... 63 2e-10
TC82168 weakly similar to GP|11762132|gb|AAG40344.1 AT3g62110 {A... 62 4e-10
TC79890 similar to GP|11762132|gb|AAG40344.1 AT3g62110 {Arabidop... 62 5e-10
AJ496921 similar to GP|10185719|gb| NTS1 protein {Nicotiana taba... 57 1e-08
>TC90178 similar to GP|13958032|gb|AAK50769.1 polygalacturonase {Pisum
sativum}, partial (94%)
Length = 1114
Score = 560 bits (1442), Expect(2) = e-161
Identities = 261/344 (75%), Positives = 297/344 (85%), Gaps = 24/344 (6%)
Frame = +3
Query: 169 KPNIVFQLDGTIIAPTNSNAWGGGLLQWLEFSKLVGITIQGKGVIDGRGSVWWQDFPYDD 228
KP I+FQ+DGTIIAPTNSNAWG GLLQWL+F+KLVG TIQG G+ DGRGSVWWQD Y+D
Sbjct: 3 KPRIIFQVDGTIIAPTNSNAWGRGLLQWLDFTKLVGFTIQGNGIFDGRGSVWWQDTQYND 182
Query: 229 PIDDEEKLIVPLNHT-QKPSLPVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSPQC 287
P+DDEEKL+VPLN+T P + +++ +G +MP++KPTA+RFYGS +PTVTGITIQNSPQC
Sbjct: 183 PLDDEEKLLVPLNNTIVSPPMQIESSLGGKMPSIKPTAIRFYGSINPTVTGITIQNSPQC 362
Query: 288 HLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHSSKMACG------------- 334
HLKFDNCNGVLVHDV+ISSPGDSPNTDGIHLQNS+DVLI+ S MACG
Sbjct: 363 HLKFDNCNGVLVHDVTISSPGDSPNTDGIHLQNSRDVLIYKSTMACGDDCISIQTGCSNV 542
Query: 335 ----------HGISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSGSVQGVL 384
HGISIG LGKDNTRACVSNIT+RDVNMHNTMNGVRIKTWQGGSGSVQGVL
Sbjct: 543 YVHNVDCGPGHGISIGGLGKDNTRACVSNITVRDVNMHNTMNGVRIKTWQGGSGSVQGVL 722
Query: 385 FSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALAGINYEGIKGTYTVKPVHFACSDSLP 444
FSNIQVS+VQ+PI+IDQFYCDKR+C N T+AVAL GINYEGIKGTYTVKPVHFACSDSLP
Sbjct: 723 FSNIQVSQVQVPIIIDQFYCDKRNCKNQTAAVALTGINYEGIKGTYTVKPVHFACSDSLP 902
Query: 445 CVDVSLTSVELKPVQEQYHLYEPFCWQTYGQLMTPTTPPIDCIQ 488
C+DVSLTSVEL+PVQ++YHLY+PFCW+TYG+L T T PPIDC+Q
Sbjct: 903 CIDVSLTSVELQPVQDKYHLYDPFCWETYGELKTATVPPIDCLQ 1034
Score = 27.7 bits (60), Expect(2) = e-161
Identities = 11/15 (73%), Positives = 13/15 (86%)
Frame = +1
Query: 488 QIGKPSSNRIQTNHD 502
+IGKP +NRIQT HD
Sbjct: 1033 KIGKPPNNRIQTAHD 1077
>AL376379 homologue to GP|13958032|gb polygalacturonase {Pisum sativum},
partial (43%)
Length = 459
Score = 288 bits (737), Expect = 3e-78
Identities = 134/152 (88%), Positives = 141/152 (92%)
Frame = +1
Query: 183 PTNSNAWGGGLLQWLEFSKLVGITIQGKGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNH 242
PTN NAW G LQWLE +KL GITIQG GVIDGRGSVWWQDFPYD+PIDDEEKLIVPLN
Sbjct: 1 PTNPNAWSGVTLQWLECTKLEGITIQGNGVIDGRGSVWWQDFPYDNPIDDEEKLIVPLNQ 180
Query: 243 TQKPSLPVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDV 302
TQKP +PVQNEMGR+MP+ KPTALRFYGS+ PTVTGITIQNSPQCHLKFDNCNGVLVHDV
Sbjct: 181 TQKPPMPVQNEMGRKMPSNKPTALRFYGSYGPTVTGITIQNSPQCHLKFDNCNGVLVHDV 360
Query: 303 SISSPGDSPNTDGIHLQNSKDVLIHSSKMACG 334
SISSPGDSPNTDGIHLQNSKDVLIHSSK+ACG
Sbjct: 361 SISSPGDSPNTDGIHLQNSKDVLIHSSKLACG 456
>TC93674 similar to GP|15292729|gb|AAK92733.1 putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (51%)
Length = 745
Score = 231 bits (588), Expect = 6e-61
Identities = 118/247 (47%), Positives = 163/247 (65%), Gaps = 25/247 (10%)
Frame = +3
Query: 263 PTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSK 322
PT +RF+ S + + G+ IQNSPQ H+KFD C GVL+ ++SIS+P SPNTDGIHL N++
Sbjct: 6 PTMIRFFMSSNLVLRGLKIQNSPQFHVKFDGCQGVLIDELSISAPKLSPNTDGIHLGNTR 185
Query: 323 DVLIHSSKMACG-----------------------HGISIGSLGKDNTRACVSNITIRDV 359
DV I++S ++ G HGISIGSLG N+ ACVSN+T+R+
Sbjct: 186 DVGIYNSLISNGDDCISIGPGCSNVNVDGVTCAPTHGISIGSLGVHNSHACVSNLTVRNS 365
Query: 360 NMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALA 419
+ + NG+RIKTWQGG+GSV G+ F NIQ+ V+ I IDQFYC + C N TSAV +
Sbjct: 366 IIKESDNGLRIKTWQGGTGSVTGLTFDNIQMENVRNCINIDQFYCLSKECMNQTSAVYVN 545
Query: 420 GINYEGIKGTYTVK--PVHFACSDSLPCVDVSLTSVELKPVQEQYHLYEPFCWQTYGQLM 477
I+Y IKGTY V+ P+HFACSD++ C +++L+ +EL P + + + +PFCW YG+
Sbjct: 546 NISYRKIKGTYDVRTPPIHFACSDTVACTNITLSEIELLPYEGEL-VDDPFCWNAYGRQE 722
Query: 478 TPTTPPI 484
T T PP+
Sbjct: 723 TLTIPPL 743
>TC91675 similar to GP|6624205|dbj|BAA88472.1 polygalacturonase {Cucumis
sativus}, partial (65%)
Length = 1146
Score = 200 bits (508), Expect = 1e-51
Identities = 118/324 (36%), Positives = 166/324 (50%), Gaps = 25/324 (7%)
Frame = +1
Query: 196 WLEFSKLVGITIQGKGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMG 255
WL+FSKL QG GVIDG GS WW + + + P +
Sbjct: 25 WLDFSKLKKAVFQGSGVIDGSGSKWWA-----------------ASCKKNKTNPCKGA-- 147
Query: 256 RQMPAVKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDG 315
PTAL S S V G+TIQN+ Q H C+ V + V ++SPGDSPNTDG
Sbjct: 148 -------PTALTIDTSSSIKVKGLTIQNNQQMHFTISRCDSVRILGVKVASPGDSPNTDG 306
Query: 316 IHLQNSKDVLIHSSKMACG-----------------------HGISIGSLGKDNTRACVS 352
IH+ S +V++ K+ G HGISIGSLGKDN+ V+
Sbjct: 307 IHISESTNVIVQDCKIGTGDDCISIVNASSNIKMKRIFCGPGHGISIGSLGKDNSTGVVT 486
Query: 353 NITIRDVNMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDK-RSCTN 411
+ + + T NG+RIKTWQGG+G V+GV F N++V V PI+IDQFYCD SC N
Sbjct: 487 KVILDTAFLKGTTNGLRIKTWQGGAGYVKGVRFQNVRVENVSNPILIDQFYCDSPTSCQN 666
Query: 412 HTSAVALAGINYEGIKG-TYTVKPVHFACSDSLPCVDVSLTSVELKPVQEQYHLYEPFCW 470
+SA+ ++ I Y+ + G T + K + F CSD++PC ++ L++V+L ++Q E +C
Sbjct: 667 QSSALEISEIMYQNVSGTTNSAKAIKFDCSDTVPCSNLVLSNVDL---EKQDGTVETYCH 837
Query: 471 QTYGQLMTPTTPPIDCIQIGKPSS 494
G P +C+ +S
Sbjct: 838 SAQGFGYGVVHPSAECLNSSDKTS 909
>AW586449 similar to PIR|B96631|B966 probable polygalacturonase F8A5.12
[imported] - Arabidopsis thaliana, partial (13%)
Length = 448
Score = 199 bits (505), Expect = 3e-51
Identities = 99/151 (65%), Positives = 111/151 (72%), Gaps = 1/151 (0%)
Frame = +3
Query: 12 FSYMLLIVLLIWSSNFETCIARRGKHWRQSRTVSASLFKKKGNSHGNSHNQHGGGGSSSH 71
F+YM LI +LIWSSNFE CIARRGKHWRQ+R ASL KKKG +HGN H+ +GGG
Sbjct: 9 FTYMFLIAILIWSSNFEECIARRGKHWRQNREEIASLLKKKGKNHGNGHSHNGGG----- 173
Query: 72 SKPKPPPPTHKKTPSSPLFPKPKVD-PPSTPPSNDYNGGSSTMFNVLDFGAKGDGRTDDT 130
SK K PPP SP P P+ + PSTPP Y GGSST FNVLDFGAKGDGR+DDT
Sbjct: 174 SKSKSPPPQKSIPSLSPPLPPPQEEVTPSTPPPQTYIGGSSTTFNVLDFGAKGDGRSDDT 353
Query: 131 KAFQATCAMACKVEASTMVVPAAYVFYVGPI 161
KAF+AT ACKVEASTM++PA Y FYVGPI
Sbjct: 354 KAFEATWGAACKVEASTMLIPADYTFYVGPI 446
>CA919900 similar to GP|15292729|gb putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (33%)
Length = 752
Score = 172 bits (435), Expect = 4e-43
Identities = 82/161 (50%), Positives = 113/161 (69%), Gaps = 2/161 (1%)
Frame = -2
Query: 336 GISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQL 395
GISIGSLG N++ACVSN+T+RD + + NG+RIKTWQGG GSV + F NIQ+ V
Sbjct: 751 GISIGSLGVHNSQACVSNLTVRDTVIRESDNGLRIKTWQGGMGSVSNLKFENIQMENVGN 572
Query: 396 PIVIDQFYCDKRSCTNHTSAVALAGINYEGIKGTYTVK--PVHFACSDSLPCVDVSLTSV 453
I+IDQ+YC + C N TSAV + ++Y+ IKGTY V+ P+HFACSD++ C +++L+ V
Sbjct: 571 CILIDQYYCLTKECLNQTSAVHVNDVSYKNIKGTYDVRTAPIHFACSDTVACTNITLSEV 392
Query: 454 ELKPVQEQYHLYEPFCWQTYGQLMTPTTPPIDCIQIGKPSS 494
EL P + + L +PFCW YG T T PPI+C++ G P +
Sbjct: 391 ELLPFEGEL-LDDPFCWNAYGAQETLTIPPINCLREGDPET 272
>TC83380 weakly similar to GP|10176984|dbj|BAB10216. polygalacturonase-like
protein {Arabidopsis thaliana}, partial (21%)
Length = 1220
Score = 162 bits (409), Expect = 4e-40
Identities = 115/379 (30%), Positives = 177/379 (46%), Gaps = 26/379 (6%)
Frame = +2
Query: 109 GSSTMFNVLDFGAKGDGRTDDTKAFQATCAMACKVE-ASTMVVPAAYVFYVGP-ISFSGP 166
G T+ NV+ +GAKGDGRTDD+KAF C +T+VVP + F+V P ++FSGP
Sbjct: 80 GEDTL-NVIHYGAKGDGRTDDSKAFVDAFKALCGGRYGNTLVVPNGHSFFVRPTLNFSGP 256
Query: 167 YCKPNIVFQLDGTIIAPTNSNAWGGGL-LQWLEFSKLVGITIQGKGVIDGRGSVWWQDFP 225
NI ++ G I+AP S+ WG L WL F + G+T+ G GVI+G G W
Sbjct: 257 CYSKNINIKIMGNILAPKRSD-WGRECSLMWLHFFNISGLTLDGSGVINGNGEGW----- 418
Query: 226 YDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSP 285
+ EK I P + PTAL+F + G+T N P
Sbjct: 419 -----ESREKGI------------------GGCPRI-PTALQFDKCDGLQINGLTHINGP 526
Query: 286 QCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHS----------------- 328
H+ + + + + I+SP S NTDGI L + V IH
Sbjct: 527 GAHIAVIDSQDITISHIHINSPKKSHNTDGIDLTRTIGVNIHDIQIESGDDCIAVKGGSQ 706
Query: 329 ----SKMACG--HGISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSGSVQG 382
S + CG HGIS+GSLG + V + ++++ + + V+IKTW GG G +
Sbjct: 707 FVNVSNVTCGPGHGISVGSLGGHGSEEFVQHFSVKNCTFNGADSAVKIKTWPGGKGYAKH 886
Query: 383 VLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALAGINYEGIKGTYTVKPVHFACSDS 442
++F +I +++ P+ IDQ Y R+ H AV ++ I + I GT +
Sbjct: 887 IIFEDIIINQTNYPVFIDQHY--MRTPEQH-QAVKISNITFSNIYGTCIGEDAVVLDCAK 1057
Query: 443 LPCVDVSLTSVELKPVQEQ 461
+ C +++L + + + +
Sbjct: 1058IGCYNITLNQINITSINRK 1114
>TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (35%)
Length = 822
Score = 116 bits (291), Expect = 2e-26
Identities = 75/210 (35%), Positives = 101/210 (47%), Gaps = 5/210 (2%)
Frame = +3
Query: 80 THKKTPSSPLFPKPKVDPPSTPPSNDYNGGSST-MFNVLDFGAKGDGRTDDTKAFQATCA 138
T +K SS L PS P ND SS +F+V FGA GDG DDT AF+
Sbjct: 261 TKQKKVSSLLPSNSSPSVPSDPYPNDPRDSSSDCIFDVTSFGAIGDGSADDTPAFKKAWK 440
Query: 139 MACKVEASTMVVPAAYVFYVGPISFSGPYCKPNIVFQLDGTIIAPTNSNAW--GGGLLQW 196
AC VE+ ++VP Y F + I FSGP CKP +VFQ+DG ++AP ++W QW
Sbjct: 441 AACAVESGILLVPENYTFMITSIIFSGP-CKPGLVFQVDGMLMAPDGPDSWPEADSRNQW 617
Query: 197 LEFSKLVGITIQGKGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGR 256
L F KL +++ G G I+G G WW P P + G+
Sbjct: 618 LVFYKLDQMSLNGTGTIEGNGDQWW----------------------DLPCKPHRGPDGK 731
Query: 257 QM--PAVKPTALRFYGSFSPTVTGITIQNS 284
+ P P +RF+ S + V G+ I+NS
Sbjct: 732 TLSGPCGSPAMIRFFMSSNLYVNGLKIENS 821
>BI309738 weakly similar to GP|15028105|gb putative polygalacturonase
{Arabidopsis thaliana}, partial (31%)
Length = 768
Score = 109 bits (272), Expect = 3e-24
Identities = 69/210 (32%), Positives = 100/210 (46%), Gaps = 2/210 (0%)
Frame = +2
Query: 110 SSTMFNVLDFGAKGDGRTDDTKAFQATCAMACKVEASTMVV-PAAYVFYVGPISFSGPYC 168
S ++ DFGA+GDG +DT+AF +AC + +V P F V PI GP C
Sbjct: 218 SRRFLSIADFGARGDGFHNDTQAFLKVWEIACSLSGFINIVFPYQKTFLVTPIDIGGP-C 394
Query: 169 KPNIVFQLDGTIIAPTNSNAWGG-GLLQWLEFSKLVGITIQGKGVIDGRGSVWWQDFPYD 227
+ I ++ G I+AP N + W G +W+ F + ++++G G IDG G WW
Sbjct: 395 RSKITLRILGAIVAPRNPDVWHGLNKRKWIYFHGVNHLSVEGGGRIDGMGQEWWSRS--- 565
Query: 228 DPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSPQC 287
+N T P LP PTAL F+ S V +T+ NS +
Sbjct: 566 ----------CKIN-TTNPCLPA------------PTALTFHKCKSLKVRNLTVLNSQKM 676
Query: 288 HLKFDNCNGVLVHDVSISSPGDSPNTDGIH 317
H+ F +C V+ + + +P SPNTDGIH
Sbjct: 677 HIAFTSCMRVVASRLKVLAPALSPNTDGIH 766
>BF644949 similar to GP|15292729|gb putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (22%)
Length = 673
Score = 100 bits (249), Expect = 1e-21
Identities = 57/143 (39%), Positives = 74/143 (50%), Gaps = 3/143 (2%)
Frame = +1
Query: 63 HGGGGSSSHSKPKPPPPTHKK-TPSSPLFPKPKVDPPSTPPSNDYNGGSSTMFNVLDFGA 121
H G H KPK P +P SP FP P+ P + S+ +F+V FGA
Sbjct: 256 HNVEGRYHHKKPKKTSPAPSDPSPPSPSFPSDPYPYPNDPGESP----SNCVFDVRSFGA 423
Query: 122 KGDGRTDDTKAFQATCAMACKVEASTMVVPAAYVFYVGPISFSGPYCKPNIVFQLDGTII 181
GDG DDT AF+A AC V++ ++ P Y F + FSGP CKP +VFQ+DGT++
Sbjct: 424 VGDGDADDTAAFRAAWKAACAVDSGVLLAPENYCFKITSTIFSGP-CKPGLVFQIDGTLM 600
Query: 182 APTNSNAW--GGGLLQWLEFSKL 202
AP N W QWL F +L
Sbjct: 601 APDGPNCWPEADSKSQWLVFYRL 669
>BG588797 weakly similar to PIR|T08215|T08 polygalacturonase (EC 3.2.1.15) 3
precursor - muskmelon, partial (35%)
Length = 631
Score = 85.1 bits (209), Expect(2) = 6e-21
Identities = 47/123 (38%), Positives = 68/123 (55%), Gaps = 23/123 (18%)
Frame = +2
Query: 276 VTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDV----------- 324
V + ++S Q H+ F+ C V V ++ + +P DSPNTDGIH+ ++++
Sbjct: 8 VKNLRFKDSQQMHVVFERCFNVFVSNLIVRAPEDSPNTDGIHVAETQNIDIINCDIGTGD 187
Query: 325 ----LIHSSK------MACG--HGISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKT 372
++ SK + CG HGISIGSLG DN+ A VSN+ + + T NGVRIKT
Sbjct: 188 DCISIVSGSKNVRAIDITCGPGHGISIGSLGADNSEAEVSNVVVNRAALTGTTNGVRIKT 367
Query: 373 WQG 375
W G
Sbjct: 368 WPG 376
Score = 33.9 bits (76), Expect(2) = 6e-21
Identities = 14/32 (43%), Positives = 21/32 (64%)
Frame = +3
Query: 371 KTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQF 402
K QGGSG + + F NI++ V P++IDQ+
Sbjct: 363 KHGQGGSGYARNIKFMNIKMQNVTNPVIIDQY 458
>AW585498 weakly similar to GP|10177065|db polygalacturonase-like protein
{Arabidopsis thaliana}, partial (26%)
Length = 583
Score = 96.7 bits (239), Expect = 2e-20
Identities = 53/158 (33%), Positives = 82/158 (51%), Gaps = 24/158 (15%)
Frame = +3
Query: 283 NSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHSSKMACG-------- 334
N + H+ +C ++ ++ + +PG+SPNTDGI + S+D+ + +S +A G
Sbjct: 15 NPSRSHITLTSCKKGIISNIRLIAPGESPNTDGIDISASRDIQVLNSFIATGDDCIAISA 194
Query: 335 ---------------HGISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSGS 379
HGISIGSLG V ++ +++ + T+ GVRIKT QGG G
Sbjct: 195 GSSVIKITAITCGPGHGISIGSLGARGDTDIVEDVHVKNCTLTETLTGVRIKTKQGGGGF 374
Query: 380 VQGVLFSNIQVSEVQLPIVIDQFYC-DKRSCTNHTSAV 416
+ + F NI+ PI IDQFYC ++ C N T A+
Sbjct: 375 ARRITFENIKFVRAHNPIWIDQFYCVNQMVCRNMTKAI 488
>TC84640 similar to GP|10177065|dbj|BAB10507. polygalacturonase-like protein
{Arabidopsis thaliana}, partial (12%)
Length = 567
Score = 73.9 bits (180), Expect(2) = 4e-17
Identities = 43/104 (41%), Positives = 62/104 (59%), Gaps = 4/104 (3%)
Frame = +2
Query: 111 STMFNVLDFGAKGDGRTDDTKAFQATCAMACKVEAST--MVVPAAYVFYVGPISFSGPYC 168
+T FNVL +GA GDG+T+D+ AF C ++ T +++PAA F + PI+F GP C
Sbjct: 89 TTTFNVLQYGAVGDGKTNDSPAFLKAWKDVCNSKSGTSRLIIPAAKTFLLRPIAFRGP-C 265
Query: 169 KPNIVF-QLDGTIIAP-TNSNAWGGGLLQWLEFSKLVGITIQGK 210
K N ++ +L G IIAP T S G + W+ FS + G+ I K
Sbjct: 266 KSNYIYIELSGNIIAPKTKSEYSGSPINTWIGFSFVNGLIISWK 397
Score = 32.0 bits (71), Expect(2) = 4e-17
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = +3
Query: 209 GKGVIDGRGSVWWQ 222
GKG +DGRGS+WW+
Sbjct: 393 GKGTVDGRGSMWWK 434
>BQ148150 weakly similar to GP|10185719|gb NTS1 protein {Nicotiana tabacum},
partial (31%)
Length = 669
Score = 85.5 bits (210), Expect = 4e-17
Identities = 61/223 (27%), Positives = 102/223 (45%), Gaps = 24/223 (10%)
Frame = +2
Query: 146 STMVVPAAYVFYVGPISFSGPYCKPN-IVFQLDGTIIAPTNSNAWGGGLLQWLEFSKLVG 204
+T+++PA F G F+GP P I ++ G A T+ + + +W +F + G
Sbjct: 5 ATVLIPAG-TFVTGQTLFAGPCTSPKPITVEIVGKAQATTDLSEYYSP--EWFQFENIDG 175
Query: 205 ITIQGKGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPT 264
+ ++G GV DG+GS+ W PLN ++ + A P+
Sbjct: 176 LVLKGSGVFDGQGSISW-----------------PLNDCKQ---------NKGNCASLPS 277
Query: 265 ALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDV 324
+L+F + V +T NS Q H C+ + ++ I++PG+SPNTDG+H+ +S +
Sbjct: 278 SLKFDKIKNAIVQDVTSLNSMQFHFHLHGCSNISFTNLHITAPGNSPNTDGMHISSSDFI 457
Query: 325 LIHSSKMAC-----------------------GHGISIGSLGK 344
+ +S +A GHGIS+GSLGK
Sbjct: 458 TVTNSVIATGDDCISVGHSTSNITISGITCGPGHGISVGSLGK 586
>TC86720 similar to PIR|T47941|T47941 hypothetical protein F2A19.90 -
Arabidopsis thaliana, partial (85%)
Length = 1719
Score = 67.8 bits (164), Expect = 9e-12
Identities = 88/428 (20%), Positives = 157/428 (36%), Gaps = 55/428 (12%)
Frame = +2
Query: 115 NVLDFGAKGDGRTDDTKAFQATCAMACKVEA---STMVVPAAYVFYVGPISFSGPYCKPN 171
++ DFG GDG T +TKAFQ+ + + + S + VPA + G S + +
Sbjct: 275 SLTDFGGVGDGNTSNTKAFQSAISHLSQYGSQGGSQLYVPAGK-WLTGSFSLTSHF---T 442
Query: 172 IVFQLDGTIIAPTNSNAWG---------------GGLLQWLEF-SKLVGITIQG-KGVID 214
+ D ++A + W G + L F + L + + G G ID
Sbjct: 443 LYLDRDAVLLASQDITEWPVLEPLPSYGRGRDAPAGRFRSLIFGTNLTDVIVTGGNGTID 622
Query: 215 GRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPTALRFYGSFSP 274
G+G+ WWQ F H +K + +P + S S
Sbjct: 623 GQGAFWWQQF-----------------HRKK------------LKYTRPYLIELMFSDSI 715
Query: 275 TVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHSSKMACG 334
++ +T+ NSP ++ + +++ ++I +P SPNTDGI+ + + I + G
Sbjct: 716 QISNLTLLNSPSWNVHPVYSSNIIIQGITIIAPISSPNTDGINPDSCTNTKIEDCYIVSG 895
Query: 335 ------------HGISIG---------------------SLGKDNTRACVSNITIRDVNM 361
+GI G +LG + + + ++ D+
Sbjct: 896 DDCVAVKSGWDEYGIKFGWPTKQLVIRRLTCISPFSATIALGSEMSGG-IQDVRAEDITA 1072
Query: 362 HNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAV-ALAG 420
T +GVRIKT G G V+ + + ++ + Y +A+ +A
Sbjct: 1073IRTESGVRIKTAVGRGGYVKDIYVKRFTMHTMKWAFKMTGDYNSHADTHFDPNALPEIAN 1252
Query: 421 INYEGIKGTYTVKPVHFACSDSLPCVDVSLTSVEL-KPVQEQYHLYEPFCWQTYGQLMTP 479
INY + F + P + + +V L V+ + + C G
Sbjct: 1253INYRDVVAENVTIAARFQGISNDPFKGICIANVTLGMAVKAKKRSWT--CTDIEGMTSGV 1426
Query: 480 TTPPIDCI 487
T PP D +
Sbjct: 1427TPPPCDLL 1450
>BI309956 weakly similar to GP|7381227|gb|A polygalacturonase {Lycopersicon
esculentum}, partial (21%)
Length = 512
Score = 65.5 bits (158), Expect = 5e-11
Identities = 36/94 (38%), Positives = 52/94 (55%), Gaps = 2/94 (2%)
Frame = +3
Query: 393 VQLPIVIDQFYCDKR-SCTNHTSAVALAGINYEGIKGT-YTVKPVHFACSDSLPCVDVSL 450
V PI+IDQ+YCD R C N TSAV + I++ I+GT T + + FACSD+ PC + L
Sbjct: 15 VSNPIIIDQYYCDSRHPCKNQTSAVQVGNISFINIQGTSATEETIKFACSDASPCEGLYL 194
Query: 451 TSVELKPVQEQYHLYEPFCWQTYGQLMTPTTPPI 484
++ L+ +CWQ +G PP+
Sbjct: 195 ENIFLRSYFGGN--TRSYCWQAHGSAQGYVYPPV 290
>TC89269 similar to PIR|T05614|T05614 hypothetical protein F9D16.290 -
Arabidopsis thaliana, partial (60%)
Length = 923
Score = 63.2 bits (152), Expect = 2e-10
Identities = 67/290 (23%), Positives = 114/290 (39%), Gaps = 28/290 (9%)
Frame = +1
Query: 115 NVLDFGAKGDGRTDDTKAFQATC--AMACKVEASTMVVPAAYVFYVGPISFSGP---YCK 169
++ DFG GD +T ++KAF+A + T++ V+ + + Y
Sbjct: 163 SITDFGGVGDSKTLNSKAFRAAIYRIQHLRRRGGTLLYVPPGVYLTDSFNLTSHMTLYLA 342
Query: 170 PNIVFQL-----DGTIIAPTNSNAWG-----GGLLQWLEFSKLVGITIQGK-GVIDGRGS 218
V + + +IAP S G G + ++ + + I G+ G IDG+G
Sbjct: 343 AGAVIKATQRLRNWPLIAPLPSYGRGRELPGGRYISFIHGDGVRDVIITGENGTIDGQGD 522
Query: 219 VWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPTALRFYGSFSPTVTG 278
VWW + R + +P + F S + ++
Sbjct: 523 VWWNMWRQ-----------------------------RTLQFTRPNLVEFLNSRNIIISN 615
Query: 279 ITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHSSKMACG---- 334
+ +NSP ++ C+ V+V V+I +P DSPNTDGI +S +V I S ++ G
Sbjct: 616 VIFKNSPFWNIHPVYCSNVVVRYVTILAPRDSPNTDGIDPDSSSNVCIEDSYISTGDDLV 795
Query: 335 --------HGISIGSLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGG 376
+GIS G D ITIR + + G+ + + G
Sbjct: 796 AVKSGWDAYGISYGRPSND--------ITIRRITGSSPFAGIALGSETSG 921
>TC82168 weakly similar to GP|11762132|gb|AAG40344.1 AT3g62110 {Arabidopsis
thaliana}, partial (33%)
Length = 587
Score = 62.4 bits (150), Expect = 4e-10
Identities = 46/164 (28%), Positives = 79/164 (48%), Gaps = 32/164 (19%)
Frame = +3
Query: 266 LRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVL 325
L S + V+ +T +NSP + C+ V++ ++I +P ++PNTDGI +S +V
Sbjct: 33 LELMNSENVLVSNLTFRNSPFWTIHPVYCSNVVIKGMTILAPLNAPNTDGIDPDSSTNVC 212
Query: 326 IHSSKMACG------------HGISIG--------SLGKDNTRAC------------VSN 353
I + + G +GI++ S T C +SN
Sbjct: 213 IEDNYIESGDDLVAIKSGWDQYGIAVAKPSTNIIVSRVSRTTPTCSGVGIGSEMSGGISN 392
Query: 354 ITIRDVNMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPI 397
ITI ++++ N+ GVRIK+ G G ++ V SNI++ V++PI
Sbjct: 393 ITIENLHVWNSAAGVRIKSDNGRGGYIKNVSISNIRMERVKIPI 524
>TC79890 similar to GP|11762132|gb|AAG40344.1 AT3g62110 {Arabidopsis
thaliana}, partial (55%)
Length = 1151
Score = 62.0 bits (149), Expect = 5e-10
Identities = 59/235 (25%), Positives = 91/235 (38%), Gaps = 33/235 (14%)
Frame = +3
Query: 202 LVGITIQGK-GVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPA 260
L + I G G IDG+GS+WW F L+HT
Sbjct: 30 LTDVIITGNNGTIDGQGSIWWSKFRNK-----------TLDHT----------------- 125
Query: 261 VKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQN 320
+P + S ++ T NSP + C+ V V +V+I P SPNTDGI +
Sbjct: 126 -RPHLVELINSTEVLISNATFLNSPFWTIHPVYCSNVTVQNVTIIVPFGSPNTDGIDPDS 302
Query: 321 SKDVLIHSSKMACG------------HGISIG--------------------SLGKDNTR 348
S +V I ++ G +GIS G ++G + +
Sbjct: 303 SDNVCIEDCYISTGDDLISIKSGWDEYGISFGRPSTNISIHRLTGRTTSAGIAIGSEMSG 482
Query: 349 ACVSNITIRDVNMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFY 403
VS + D+ + ++ + +RIKT G G V+ V SN+ + V + I Y
Sbjct: 483 G-VSEVYAEDIYIFDSKSAIRIKTSPGRGGYVRNVYISNMTLINVDIAIRFTGLY 644
>AJ496921 similar to GP|10185719|gb| NTS1 protein {Nicotiana tabacum},
partial (9%)
Length = 508
Score = 57.4 bits (137), Expect = 1e-08
Identities = 35/98 (35%), Positives = 52/98 (52%), Gaps = 2/98 (2%)
Frame = +3
Query: 366 NGVRIKTWQGGS-GSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALAGINYE 424
NG RIKTW G + + +++ ++ + +VQ PIVIDQ Y K+ TS ++ ++
Sbjct: 45 NGARIKTWIGTAPAEAKNIVYEDLIMKDVQNPIVIDQSY-GKKERVPSTSVWKISDAHFR 221
Query: 425 GIKGTYTVK-PVHFACSDSLPCVDVSLTSVELKPVQEQ 461
IKGT PV CS PC ++ + VEL V Q
Sbjct: 222 RIKGTTVSNIPVSLQCSSKNPCENIEVADVELTYVGPQ 335
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,181,050
Number of Sequences: 36976
Number of extensions: 358109
Number of successful extensions: 6454
Number of sequences better than 10.0: 283
Number of HSP's better than 10.0 without gapping: 3397
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5374
length of query: 504
length of database: 9,014,727
effective HSP length: 100
effective length of query: 404
effective length of database: 5,317,127
effective search space: 2148119308
effective search space used: 2148119308
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0008.5


