

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
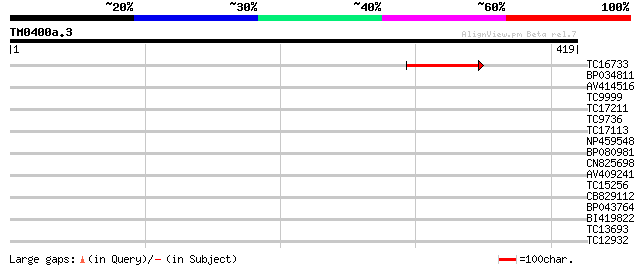
Query= TM0400a.3
(419 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16733 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, ... 111 3e-25
BP034811 30 0.97
AV414516 30 0.97
TC9999 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transport... 28 2.8
TC17211 similar to UP|PRT2_SEPOF (P80002) Spermatid-specific pro... 28 3.7
TC9736 homologue to UP|Q42558 (Q42558) Acyl-(acyl carrier protei... 27 4.8
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 27 4.8
NP459548 PSII 47KDa protein [Lotus japonicus] 27 4.8
BP080981 27 6.3
CN825698 27 6.3
AV409241 27 6.3
TC15256 homologue to UP|AX28_SOYBN (P13089) Auxin-induced protei... 27 6.3
CB829112 27 8.2
BP043764 27 8.2
BI419822 27 8.2
TC13693 similar to PIR|T00933|T00933 RNA-binding protein homolog... 27 8.2
TC12932 27 8.2
>TC16733 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, partial
(26%)
Length = 629
Score = 111 bits (277), Expect = 3e-25
Identities = 54/57 (94%), Positives = 55/57 (95%)
Frame = +3
Query: 294 LLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIQSESLREASFQ 350
LLK KVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGI SESLREAS+Q
Sbjct: 3 LLKXKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQIQYGFGIXSESLREASYQ 173
>BP034811
Length = 578
Score = 29.6 bits (65), Expect = 0.97
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = -3
Query: 4 KEVKPPPPPLVVLVPPLFDFP 24
+E+ P PPP++ ++ P FDFP
Sbjct: 525 EEIWPSPPPVLCVLSPSFDFP 463
>AV414516
Length = 424
Score = 29.6 bits (65), Expect = 0.97
Identities = 16/31 (51%), Positives = 18/31 (57%), Gaps = 5/31 (16%)
Frame = +3
Query: 1 WFRKEVKPPPPPLVVL-VPPLFDF----PPL 26
WF E +PPP LV L PPLF+ PPL
Sbjct: 315 WFNPEARPPPSSLVCLSSPPLFNAEARPPPL 407
>TC9999 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transporter,
partial (43%)
Length = 656
Score = 28.1 bits (61), Expect = 2.8
Identities = 13/22 (59%), Positives = 14/22 (63%), Gaps = 2/22 (9%)
Frame = +3
Query: 6 VKPPPPPLVVL--VPPLFDFPP 25
+KPPPPP L PP F FPP
Sbjct: 162 LKPPPPPPSPLSTAPPPFKFPP 227
>TC17211 similar to UP|PRT2_SEPOF (P80002) Spermatid-specific protein T2
[Contains: Sperm protamine SP2], partial (31%)
Length = 540
Score = 27.7 bits (60), Expect = 3.7
Identities = 25/67 (37%), Positives = 28/67 (41%), Gaps = 7/67 (10%)
Frame = +2
Query: 261 FELQTSVDDAIATNNISDSTFRIGASWQANKNF-------LLKAKVGPRISSMALAFKSW 313
FE V IAT I+ TF GAS KN LLK KV P S +L+ W
Sbjct: 164 FERNAIVGGRIATVTIAGETFEAGASILHPKNLHAVDYTKLLKLKVKPP-GSDSLSLGIW 340
Query: 314 WKPSFTF 320
F F
Sbjct: 341 DGNKFVF 361
>TC9736 homologue to UP|Q42558 (Q42558) Acyl-(acyl carrier protein)
thioesterase , partial (49%)
Length = 1058
Score = 27.3 bits (59), Expect = 4.8
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = -3
Query: 7 KPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGK 43
KPP P + L PP PP + ES D V GK
Sbjct: 402 KPPEEPCLDLCPPRLTCPPPSLAPAAPESCEDEVTGK 292
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 27.3 bits (59), Expect = 4.8
Identities = 12/26 (46%), Positives = 15/26 (57%), Gaps = 3/26 (11%)
Frame = +1
Query: 8 PPPPPLVVLVPPLF---DFPPLAARN 30
PPPPP PPLF PP++ +N
Sbjct: 760 PPPPPFFPPFPPLFPPPGRPPISTKN 837
>NP459548 PSII 47KDa protein [Lotus japonicus]
Length = 1527
Score = 27.3 bits (59), Expect = 4.8
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +2
Query: 102 RILRMRSCAYYPRYGFGAFGV 122
RIL R+CA++P G G+ G+
Sbjct: 296 RILCFRACAFWPLSGIGSIGI 358
>BP080981
Length = 427
Score = 26.9 bits (58), Expect = 6.3
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = -3
Query: 296 KAKVGPRISSMALAFKSWWKPSFT 319
KA+ P ISS+ L F +W SFT
Sbjct: 191 KARAVPLISSIGLHFDPYWHSSFT 120
>CN825698
Length = 704
Score = 26.9 bits (58), Expect = 6.3
Identities = 14/25 (56%), Positives = 14/25 (56%), Gaps = 1/25 (4%)
Frame = +1
Query: 8 PPPPPLVVLVPPL-FDFPPLAARNR 31
PPPPP L PP D PP AR R
Sbjct: 145 PPPPPPPSLPPPSDSDLPPPPARRR 219
>AV409241
Length = 282
Score = 26.9 bits (58), Expect = 6.3
Identities = 11/21 (52%), Positives = 12/21 (56%)
Frame = +2
Query: 8 PPPPPLVVLVPPLFDFPPLAA 28
PPPPP + PP PP AA
Sbjct: 122 PPPPPPATVSPPPTPSPPAAA 184
>TC15256 homologue to UP|AX28_SOYBN (P13089) Auxin-induced protein AUX28,
partial (51%)
Length = 664
Score = 26.9 bits (58), Expect = 6.3
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -1
Query: 4 KEVKPPPPPLVVLVPPL 20
K PPPPPL++ PPL
Sbjct: 532 KLAPPPPPPLLLFSPPL 482
>CB829112
Length = 234
Score = 26.6 bits (57), Expect = 8.2
Identities = 12/18 (66%), Positives = 13/18 (71%)
Frame = +3
Query: 8 PPPPPLVVLVPPLFDFPP 25
PPPPPL +L PPL PP
Sbjct: 75 PPPPPL-LLPPPLLPPPP 125
>BP043764
Length = 479
Score = 26.6 bits (57), Expect = 8.2
Identities = 14/38 (36%), Positives = 17/38 (43%)
Frame = +3
Query: 1 WFRKEVKPPPPPLVVLVPPLFDFPPLAARNRMLESSYD 38
W ++ PPPPP PP PPL R E + D
Sbjct: 291 WLTSKLSPPPPP-----PPPPPPPPLPPPERETEIAED 389
>BI419822
Length = 380
Score = 26.6 bits (57), Expect = 8.2
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = -3
Query: 8 PPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVF 41
PPPPP PP PPL R+ +S+ V+F
Sbjct: 378 PPPPPYPPPPPPYLPPPPLLPRD---*ASFTVIF 286
>TC13693 similar to PIR|T00933|T00933 RNA-binding protein homolog
At2g42240 - Arabidopsis thaliana {Arabidopsis
thaliana;}, partial (53%)
Length = 662
Score = 26.6 bits (57), Expect = 8.2
Identities = 8/12 (66%), Positives = 11/12 (91%)
Frame = +3
Query: 8 PPPPPLVVLVPP 19
PPPPP++ +VPP
Sbjct: 60 PPPPPIMTVVPP 95
>TC12932
Length = 451
Score = 26.6 bits (57), Expect = 8.2
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = +1
Query: 270 AIATNNISDSTFRIGASWQANKNFLLKAKVGP 301
A+ T I + R+ + N+ F LKA +GP
Sbjct: 193 ALITGRIEQTHLRLSTELRRNQGFTLKASMGP 288
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,386,849
Number of Sequences: 28460
Number of extensions: 107117
Number of successful extensions: 983
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 890
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 960
length of query: 419
length of database: 4,897,600
effective HSP length: 93
effective length of query: 326
effective length of database: 2,250,820
effective search space: 733767320
effective search space used: 733767320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0400a.3


