

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0249.1
(68 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
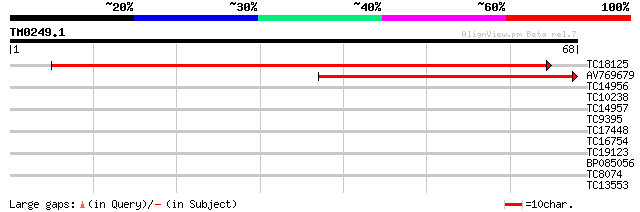
Sequences producing significant alignments: (bits) Value
TC18125 similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X carboxypep... 83 1e-17
AV769679 63 9e-12
TC14956 36 0.002
TC10238 34 0.005
TC14957 31 0.040
TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1... 29 0.20
TC17448 homologue to UP|O23628 (O23628) Histone H2A.F/Z (At3g545... 26 1.7
TC16754 25 2.2
TC19123 25 2.8
BP085056 25 2.8
TC8074 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein L... 24 4.9
TC13553 similar to UP|ARG1_ARATH (P46637) Arginase , partial (42%) 23 8.3
>TC18125 similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X carboxypeptidase,
partial (22%)
Length = 585
Score = 82.8 bits (203), Expect = 1e-17
Identities = 38/60 (63%), Positives = 47/60 (78%)
Frame = +3
Query: 6 LKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDVR 65
LK+ SNIIF NGL DPW+GG VL+NIS+++V++V KEGAHH DLR ST DP WL + R
Sbjct: 147 LKKFGSNIIFSNGLLDPWTGGSVLQNISESIVSLVTKEGAHHIDLRASTGNDPDWLVEQR 326
>AV769679
Length = 303
Score = 63.2 bits (152), Expect = 9e-12
Identities = 29/31 (93%), Positives = 29/31 (93%)
Frame = -1
Query: 38 AIVAKEGAHHRDLRYSTKEDPKWLKDVRIKE 68
AIVAKEGAHH DLRYSTKEDPKWLKDVR KE
Sbjct: 303 AIVAKEGAHHTDLRYSTKEDPKWLKDVRKKE 211
>TC14956
Length = 490
Score = 35.8 bits (81), Expect = 0.002
Identities = 17/23 (73%), Positives = 18/23 (77%)
Frame = +2
Query: 46 HHRDLRYSTKEDPKWLKDVRIKE 68
HH +LR STKED K LKDVR KE
Sbjct: 11 HHTNLRNSTKEDSKCLKDVRKKE 79
>TC10238
Length = 942
Score = 34.3 bits (77), Expect = 0.005
Identities = 13/20 (65%), Positives = 19/20 (95%)
Frame = +1
Query: 49 DLRYSTKEDPKWLKDVRIKE 68
+LR+STK+DP+WLKD+R +E
Sbjct: 235 NLRFSTKKDPEWLKDLREQE 294
>TC14957
Length = 582
Score = 31.2 bits (69), Expect = 0.040
Identities = 14/19 (73%), Positives = 15/19 (78%)
Frame = +2
Query: 46 HHRDLRYSTKEDPKWLKDV 64
HH +LR STKED K LKDV
Sbjct: 272 HHTNLRNSTKEDSKCLKDV 328
>TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 ,
partial (17%)
Length = 590
Score = 28.9 bits (63), Expect = 0.20
Identities = 15/42 (35%), Positives = 20/42 (46%)
Frame = -2
Query: 5 DLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAH 46
D R S I+ YNG ++PW+ K + LV KE H
Sbjct: 487 DQCR*KSCILLYNGKKEPWNNNNTNK*DGRKLVQTRKKESKH 362
>TC17448 homologue to UP|O23628 (O23628) Histone H2A.F/Z (At3g54560),
partial (88%)
Length = 597
Score = 25.8 bits (55), Expect = 1.7
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +1
Query: 24 SGGGVLKNISKTLVAIVAKEGAHHRD 49
+GGGV+ +I K+L+ AKE H D
Sbjct: 301 AGGGVIPHIHKSLINKTAKE*KDHFD 378
>TC16754
Length = 541
Score = 25.4 bits (54), Expect = 2.2
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +2
Query: 7 KRSASNIIFYNGLRDPW 23
K + S IIF NG +DPW
Sbjct: 29 KVAGSKIIFANGSQDPW 79
>TC19123
Length = 783
Score = 25.0 bits (53), Expect = 2.8
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +3
Query: 7 KRSASNIIFYNGLRDPW 23
K + S I+F NG +DPW
Sbjct: 282 KIAGSRIVFTNGSQDPW 332
>BP085056
Length = 465
Score = 25.0 bits (53), Expect = 2.8
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -2
Query: 15 FYNGLRDPWSGGGVLKNI 32
F GLRDPW G V+K +
Sbjct: 89 FLLGLRDPWFGWVVVKTV 36
>TC8074 similar to UP|AAQ72789 (AAQ72789) 60S ribosomal protein L5, partial
(96%)
Length = 1086
Score = 24.3 bits (51), Expect = 4.9
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = +1
Query: 6 LKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAI 39
L R I+F R+PW G + + K L+A+
Sbjct: 442 LLRPQQAIVFLARSREPWMGVWIFLTVIKDLLAL 543
>TC13553 similar to UP|ARG1_ARATH (P46637) Arginase , partial (42%)
Length = 539
Score = 23.5 bits (49), Expect = 8.3
Identities = 10/29 (34%), Positives = 18/29 (61%)
Frame = +1
Query: 34 KTLVAIVAKEGAHHRDLRYSTKEDPKWLK 62
+T ++IVA+ G H+ +Y+ K P L+
Sbjct: 37 RTGMSIVARRGIHYMQKQYAGKVSPASLE 123
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,160,460
Number of Sequences: 28460
Number of extensions: 10028
Number of successful extensions: 41
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 41
length of query: 68
length of database: 4,897,600
effective HSP length: 44
effective length of query: 24
effective length of database: 3,645,360
effective search space: 87488640
effective search space used: 87488640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0249.1


