

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0246c.3
(286 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
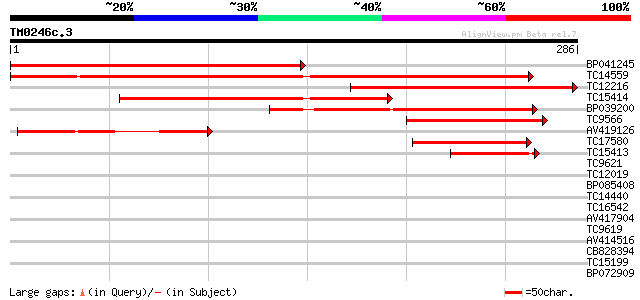
BP041245 303 2e-83
TC14559 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associat... 268 6e-73
TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Ar... 228 7e-61
TC15414 weakly similar to UP|O81655 (O81655) Senescence-associat... 140 2e-34
BP039200 128 1e-30
TC9566 weakly similar to UP|Q9FIQ5 (Q9FIQ5) Senescence-associate... 86 7e-18
AV419126 75 1e-14
TC17580 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associat... 61 2e-10
TC15413 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated prot... 50 6e-07
TC9621 similar to UP|Q9SIW2 (Q9SIW2) At2g16390 protein, partial ... 35 0.015
TC12019 29 1.1
BP085408 27 3.1
TC14440 similar to UP|V711_ARATH (O49377) Vesicle-associated mem... 27 4.0
TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, pa... 27 4.0
AV417904 27 5.2
TC9619 similar to UP|SPE1_PEA (Q43075) Arginine decarboxylase (... 27 5.2
AV414516 26 6.8
CB828394 26 6.8
TC15199 26 6.8
BP072909 26 6.8
>BP041245
Length = 516
Score = 303 bits (777), Expect = 2e-83
Identities = 149/149 (100%), Positives = 149/149 (100%)
Frame = +1
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG
Sbjct: 70 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 249
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG
Sbjct: 250 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 429
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMG 149
WLEERVASHSYWGKIASCVRDSKVCAKMG
Sbjct: 430 WLEERVASHSYWGKIASCVRDSKVCAKMG 516
>TC14559 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (92%)
Length = 1125
Score = 268 bits (686), Expect = 6e-73
Identities = 122/264 (46%), Positives = 183/264 (69%)
Frame = +2
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
++ SN +IG+LN LT +LSIPIL G+WLS +AN T+C ++L+ P+I +GV +++VSLAG
Sbjct: 122 VKFSNSVIGILNLLTLVLSIPILIAGVWLSKQAN-TECERWLEKPVIALGVFLLIVSLAG 298
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
GAC R ++L+ +YL+VMF++I +L F IFA+VVT+KG+G + N+GY +Y L YS
Sbjct: 299 LIGACCRVSWLLWVYLLVMFVLIVLLFAFTIFAFVVTNKGAGEALSNKGYKEYRLGGYSD 478
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV + + W +I SC++ KVC+ S + + D FY L+++QSGCCKP
Sbjct: 479 WLQKRVNNDNNWNRIRSCLQSGKVCSDF---KSKFLNDTVDKFYTENLSALQSGCCKPSD 649
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DC + Y+N T W + N DC W+NDQ LC+ C SCKAG+L ++K W+KV+V+
Sbjct: 650 DCNFTYENPTTWTKPANATYTNTDCDAWNNDQTILCFNCQSCKAGLLQNIKTDWKKVAVV 829
Query: 241 NIVVMIILVIVYIIAYAAYRNNKR 264
NI+ +I L+IVY + A+RN++R
Sbjct: 830 NIIFLIFLIIVYSVGCCAFRNSRR 901
>TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;}, partial (30%)
Length = 570
Score = 228 bits (582), Expect = 7e-61
Identities = 108/114 (94%), Positives = 109/114 (94%)
Frame = +2
Query: 173 SGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKK 232
S + TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKK
Sbjct: 5 SARARAATDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKK 184
Query: 233 SWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 286
SWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL
Sbjct: 185 SWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 346
>TC15414 weakly similar to UP|O81655 (O81655) Senescence-associated protein
5, partial (47%)
Length = 448
Score = 140 bits (353), Expect = 2e-34
Identities = 64/138 (46%), Positives = 93/138 (67%)
Frame = +1
Query: 56 VSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYL 115
VSLAG GAC R ++L+ +YL+VMFL+I ++ F IFA+ VT+KG+G + RGY +Y L
Sbjct: 37 VSLAGLVGACCRVSWLLWVYLLVMFLLIVLVFAFTIFAFAVTNKGAGEALSGRGYREYRL 216
Query: 116 QDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGC 175
DYS WL++RV S W +I SC+ K+C++ + + + A+ FY L+++QSGC
Sbjct: 217 GDYSNWLQKRVNSTGNWNRIRSCLGSGKLCSEF---RNRFVNDTAEQFYAENLSALQSGC 387
Query: 176 CKPPTDCGYIYQNETVWN 193
CKP DCG+ YQ +VWN
Sbjct: 388 CKPSNDCGFTYQGPSVWN 441
>BP039200
Length = 553
Score = 128 bits (322), Expect = 1e-30
Identities = 58/135 (42%), Positives = 86/135 (62%)
Frame = -1
Query: 132 WGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETV 191
W I SC+ + +CA++ N A F+ +LT +QSGCCKPPT C Y + N T
Sbjct: 553 WDHIRSCLSQTNMCAEL-----NQSFRMAQDFFNARLTPMQSGCCKPPTQCAYTFVNPTY 389
Query: 192 WNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIV 251
W + +A+ DC +WSNDQ LCY CDSCKAG+LA+L+K W++ V+ I+ +++L++V
Sbjct: 388 W-ISPINTAADMDCLQWSNDQTQLCYGCDSCKAGLLANLRKEWKRAXVVLIITLVVLIVV 212
Query: 252 YIIAYAAYRNNKRMD 266
Y++ A+RN K D
Sbjct: 211 YMVGCCAFRNAKTED 167
>TC9566 weakly similar to UP|Q9FIQ5 (Q9FIQ5) Senescence-associated protein
5-like protein, partial (28%)
Length = 675
Score = 85.9 bits (211), Expect = 7e-18
Identities = 35/71 (49%), Positives = 54/71 (75%)
Frame = +1
Query: 201 ANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYR 260
A+PDC+ W+NDQ LCY C++CKAG+L +L+K WRK ++I IV +++L+ VYIIA +A++
Sbjct: 43 ADPDCNLWNNDQNQLCYNCNACKAGLLGNLRKEWRKANIILIVAVVVLICVYIIACSAFK 222
Query: 261 NNKRMDNDEPY 271
N + D + Y
Sbjct: 223 NAQTEDLFDRY 255
>AV419126
Length = 230
Score = 75.5 bits (184), Expect = 1e-14
Identities = 43/98 (43%), Positives = 60/98 (60%)
Frame = +2
Query: 5 NHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGA 64
N+LIG+LNFLT LLSIPIL G+WLS + TDC ++L+ P+I +GV +
Sbjct: 2 NNLIGLLNFLTLLLSIPILVTGVWLSKQP-GTDCERWLERPIIALGVFL----------- 145
Query: 65 CYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSG 102
L+VMFL+I ++ F IFA+ VT+KG+G
Sbjct: 146 -----------LLVMFLLIVLVFAFTIFAFAVTNKGAG 226
>TC17580 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (12%)
Length = 526
Score = 61.2 bits (147), Expect = 2e-10
Identities = 21/60 (35%), Positives = 38/60 (63%)
Frame = +3
Query: 204 DCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNK 263
DC W N Q LCY C++CK G L +++ WR +++ N +V+ ++ +Y++ A +NN+
Sbjct: 30 DCKTWDNKQDKLCYDCNACKGGFLPNIRNQWRHLTIFNGLVLALVTTIYVLGCYAIKNNR 209
>TC15413 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated protein-like,
partial (15%)
Length = 564
Score = 49.7 bits (117), Expect = 6e-07
Identities = 21/45 (46%), Positives = 34/45 (74%)
Frame = +3
Query: 223 KAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDN 267
+AG+L ++K W+KV+V+N++ ++ L+IVY I A+RNN R DN
Sbjct: 48 QAGLLQNIKTDWKKVAVVNVIFLVFLIIVYSIGCCAFRNN-RNDN 179
>TC9621 similar to UP|Q9SIW2 (Q9SIW2) At2g16390 protein, partial (10%)
Length = 588
Score = 35.0 bits (79), Expect = 0.015
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Frame = +1
Query: 198 LMSANPDCSKWSNDQGFLC--YRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIA 255
L N DC +W N G LC Y CD + L WR V I + ++I+ ++ ++
Sbjct: 367 LKQNNVDCRQWPN*LGCLCSLYTCDPS-----SLLLCEWRDVQSIKLS*LVIIAVILVVF 531
Query: 256 Y 256
Y
Sbjct: 532 Y 534
>TC12019
Length = 510
Score = 28.9 bits (63), Expect = 1.1
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +3
Query: 139 VRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCK 177
+ C K ++DS G P P+ L ++TS GCCK
Sbjct: 231 ITSRSACQKRRQLDSCGAPSPS---LLDRVTSFGLGCCK 338
>BP085408
Length = 350
Score = 27.3 bits (59), Expect = 3.1
Identities = 16/67 (23%), Positives = 27/67 (39%), Gaps = 3/67 (4%)
Frame = -1
Query: 208 WSNDQGFLCYRCDSCKAGVLAS---LKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNKR 264
W N++ F YR +SC A + WRK + + +V I+ Y
Sbjct: 284 WVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGN 105
Query: 265 MDNDEPY 271
++ + PY
Sbjct: 104 INGESPY 84
>TC14440 similar to UP|V711_ARATH (O49377) Vesicle-associated membrane
protein 711 (AtVAMP711) (v-SNARE synaptobrevin 7C)
(AtVAMP7C), partial (28%)
Length = 704
Score = 26.9 bits (58), Expect = 4.0
Identities = 9/21 (42%), Positives = 16/21 (75%)
Frame = +1
Query: 234 WRKVSVINIVVMIILVIVYII 254
WR V + +++IILVI+Y++
Sbjct: 85 WRNVKLTIALILIILVIIYVV 147
>TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, partial (60%)
Length = 724
Score = 26.9 bits (58), Expect = 4.0
Identities = 20/69 (28%), Positives = 32/69 (45%), Gaps = 3/69 (4%)
Frame = +1
Query: 160 ADVFYLRKLTSVQSGCCKPPT---DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLC 216
+DV YL+KL +P +C I + +G+ + S P + + + G C
Sbjct: 205 SDVHYLKKLRCADFIVKEPMVIGHECAGIIEA-----VGAEVKSLVPG-DRVAIEPGISC 366
Query: 217 YRCDSCKAG 225
+RCD CK G
Sbjct: 367 WRCDHCKLG 393
>AV417904
Length = 413
Score = 26.6 bits (57), Expect = 5.2
Identities = 15/49 (30%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Frame = +3
Query: 21 PILGGGIWLSSRANNT----DCLKFLQWPLIIIGVSIMVVSLAGFAGAC 65
P + G W + + L+ L +PLIIIG+ + +S+AG C
Sbjct: 33 PFILGATWAKMAMSRSFLDYQVLERLLFPLIIIGI*LETMSIAGVKMGC 179
>TC9619 similar to UP|SPE1_PEA (Q43075) Arginine decarboxylase (ARGDC)
(ADC) , partial (13%)
Length = 795
Score = 26.6 bits (57), Expect = 5.2
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = +1
Query: 28 WLSSRANNTDCLKFLQWPLIIIGVSIMVV 56
WL+SR +T CL +L+ L+ + +MVV
Sbjct: 283 WLASRVPSTKCLIWLRLSLVASMLLVMVV 369
>AV414516
Length = 424
Score = 26.2 bits (56), Expect = 6.8
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 2/43 (4%)
Frame = +3
Query: 13 FLTFLLSIPILGGGIWLSSRANNTDCLKF--LQWPLIIIGVSI 53
F+ FLLS+P G SS ++ CL F L W L ++ VS+
Sbjct: 60 FICFLLSLPWS*SGEACSSSSSCRRCLMFHLLSWVLHLLLVSL 188
>CB828394
Length = 542
Score = 26.2 bits (56), Expect = 6.8
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 1/28 (3%)
Frame = -2
Query: 85 VLIGFIIFAYVVTDKGSGRR-VLNRGYM 111
V F A++ TD+GSGRR V N+GY+
Sbjct: 379 VRTAFAAGAFLGTDEGSGRRKVKNQGYI 296
>TC15199
Length = 835
Score = 26.2 bits (56), Expect = 6.8
Identities = 19/66 (28%), Positives = 31/66 (46%)
Frame = +1
Query: 85 VLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKV 144
VL +F +++ KGS +RV+ +D+ + W + YW SC +S +
Sbjct: 556 VLPILFLFVFILGRKGSYKRVIG---VDFPESPF--WFSFGDGNTMYWP*YFSCNLESII 720
Query: 145 CAKMGR 150
AKM R
Sbjct: 721 VAKMPR 738
>BP072909
Length = 472
Score = 26.2 bits (56), Expect = 6.8
Identities = 15/54 (27%), Positives = 26/54 (47%), Gaps = 12/54 (22%)
Frame = +3
Query: 231 KKSWRKVSVINIVVMIILVIV------------YIIAYAAYRNNKRMDNDEPYG 272
KKS +++ VINI +M ++ +I YRN+ R+ N++ G
Sbjct: 90 KKSLKEIHVINITLMHAYIVQKTNSNIESRRGGLLIFLVCYRNSTRLGNEKSQG 251
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,247,242
Number of Sequences: 28460
Number of extensions: 100699
Number of successful extensions: 600
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 591
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 593
length of query: 286
length of database: 4,897,600
effective HSP length: 89
effective length of query: 197
effective length of database: 2,364,660
effective search space: 465838020
effective search space used: 465838020
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0246c.3


