

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
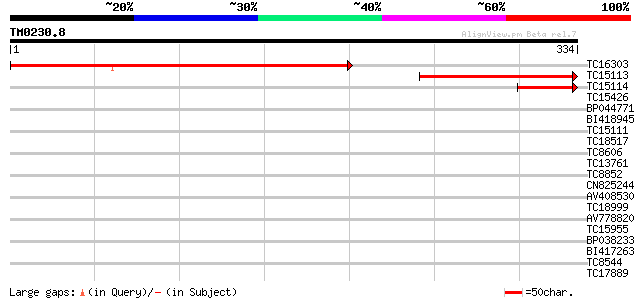
Query= TM0230.8
(334 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin at... 394 e-110
TC15113 180 3e-46
TC15114 71 2e-13
TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 k... 29 0.97
BP044771 29 0.97
BI418945 29 1.3
TC15111 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis re... 28 1.7
TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial ... 28 2.2
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 28 2.2
TC13761 weakly similar to GB|AAO63290.1|28950733|BT005226 At5g57... 28 2.8
TC8852 similar to UP|O24046 (O24046) 2-dehydro-3-deoxyphosphohep... 27 4.8
CN825244 27 4.8
AV408530 27 4.8
TC18999 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 27 6.3
AV778820 27 6.3
TC15955 27 6.3
BP038233 27 6.3
BI417263 26 8.2
TC8544 26 8.2
TC17889 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein (... 26 8.2
>TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (3%)
Length = 1235
Score = 394 bits (1012), Expect = e-110
Identities = 200/204 (98%), Positives = 201/204 (98%), Gaps = 2/204 (0%)
Frame = +1
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK- 59
DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK
Sbjct: 622 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKK 801
Query: 60 -NERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEF 118
NERSPAEIALLVENV+AELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEF
Sbjct: 802 KNERSPAEIALLVENVMAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEF 981
Query: 119 LDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIM 178
LDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIM
Sbjct: 982 LDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIM 1161
Query: 179 FLSKSDEEINVNRKLTKELVDKWS 202
FLSKSDEEINVNRKL KELVDKWS
Sbjct: 1162FLSKSDEEINVNRKLAKELVDKWS 1233
>TC15113
Length = 530
Score = 180 bits (457), Expect = 3e-46
Identities = 91/93 (97%), Positives = 91/93 (97%)
Frame = +2
Query: 242 SRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQR 301
SRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRP SKVDPEEVRARAK ASHDQQR
Sbjct: 2 SRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPPSKVDPEEVRARAKPASHDQQR 181
Query: 302 MKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
MKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY
Sbjct: 182 MKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 280
>TC15114
Length = 464
Score = 71.2 bits (173), Expect = 2e-13
Identities = 35/35 (100%), Positives = 35/35 (100%)
Frame = +1
Query: 300 QRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
QRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY
Sbjct: 1 QRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 105
>TC15426 weakly similar to UP|HS7E_DROME (P29845) Heat shock 70 kDa protein
cognate 5, partial (3%)
Length = 1216
Score = 29.3 bits (64), Expect = 0.97
Identities = 40/151 (26%), Positives = 68/151 (44%), Gaps = 5/151 (3%)
Frame = +2
Query: 36 EAPQAEEGEEDDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTAEEDAELNRQGKPA 95
+ P ++ +ED + ND+ + +E +PA +A V++ + +E+ AE ++GK
Sbjct: 230 DQPISKWDDEDVDENDVKDSWEDFDEPAPAPVAPAVKDAEKAPQKPSEKAAE--KKGK-K 400
Query: 96 INKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDF 155
+ +K+ PL + L++K Q ++ K PD NI T I K +DF
Sbjct: 401 VEPVKEEPL--DPLAEKLRQQRLVEEADYKSTKELFGGGPDEK----NIDTFIPKSESDF 562
Query: 156 PIDLEQIDRR-----EQLKRSGLGKVIMFLS 181
E I R + GL K +M LS
Sbjct: 563 LEYAELISHRLRSFEKSYHYMGLLKAVMRLS 655
>BP044771
Length = 551
Score = 29.3 bits (64), Expect = 0.97
Identities = 21/92 (22%), Positives = 44/92 (47%), Gaps = 5/92 (5%)
Frame = -3
Query: 2 DQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEIND--LFKMG--- 56
+ + + N D+D + G + +EP P +A++ + DE D L +M
Sbjct: 543 ETQPLANADNDPLLSHGGKNASGHKKRHEPKMPPILNEAKKARKYDESMDAALKRMAAAV 364
Query: 57 KKKNERSPAEIALLVENVVAELEVTAEEDAEL 88
K +++++ E V+NV++ L+ + D +L
Sbjct: 363 KSQSKKTKKEDNFSVDNVISVLQAMPDLDEDL 268
>BI418945
Length = 523
Score = 28.9 bits (63), Expect = 1.3
Identities = 16/47 (34%), Positives = 22/47 (46%)
Frame = -3
Query: 222 RAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRP 268
RAP PS + P+N S + L +P+P S SSS +P
Sbjct: 164 RAPHPSPS*QNPSNPQTAAPSASTQLAFAIPEP*SYDSSSNFKPHKP 24
>TC15111 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis regulatory
protein homolog, partial (62%)
Length = 1280
Score = 28.5 bits (62), Expect = 1.7
Identities = 16/62 (25%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +2
Query: 176 VIMFLSKSDEEI-NVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPA 234
+I +SK+ +I NVN+ +T + V K + +K+ ++ ++ PF++ KK +
Sbjct: 839 LIKIMSKNSHDILNVNKAVTVQTVKKEKKRNNDKARASSTDSKLKPKKKPFKKGDFKKSS 1018
Query: 235 NK 236
K
Sbjct: 1019GK 1024
>TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (4%)
Length = 493
Score = 28.1 bits (61), Expect = 2.2
Identities = 22/74 (29%), Positives = 35/74 (46%)
Frame = -2
Query: 10 DDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIAL 69
+D+ ID G + E PGE E+GEEDD+ + + G +++E +
Sbjct: 432 EDETVIDGAGEDNDRAVAEEEEEEPGE----EDGEEDDDGDGVPDEGDEEDEEEDHGV-- 271
Query: 70 LVENVVAELEVTAE 83
V VAE+ + AE
Sbjct: 270 -VAEEVAEVVLHAE 232
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 28.1 bits (61), Expect = 2.2
Identities = 21/72 (29%), Positives = 36/72 (49%), Gaps = 1/72 (1%)
Frame = -2
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKN 60
+D++G DDD+ +D E G +E E E+ EED+E + + KK+
Sbjct: 422 EDEDGEDQEDDDDDDEDDDEEDD--GGEDEEEEGVEEEDNEDEEEDEEDEEALQPPKKRK 249
Query: 61 E-RSPAEIALLV 71
+ + AE +LL+
Sbjct: 248 K*TTLAESSLLL 213
>TC13761 weakly similar to GB|AAO63290.1|28950733|BT005226 At5g57910
{Arabidopsis thaliana;}, partial (21%)
Length = 652
Score = 27.7 bits (60), Expect = 2.8
Identities = 14/48 (29%), Positives = 23/48 (47%)
Frame = -3
Query: 185 EEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKK 232
++ NV+RK L+D W P S + +++ P RPS+ K
Sbjct: 470 QKTNVSRKCQLLLLDSWQFPSHGLSFPMQIFIVLQETLVPQHRPSIHK 327
>TC8852 similar to UP|O24046 (O24046) 2-dehydro-3-deoxyphosphoheptonate
aldolase precursor , partial (31%)
Length = 808
Score = 26.9 bits (58), Expect = 4.8
Identities = 21/74 (28%), Positives = 30/74 (40%), Gaps = 4/74 (5%)
Frame = +3
Query: 252 PQPRSGQSSS--RQHASRPEATPMDFVIRPQSKVDPEEVRA--RAKQASHDQQRMKMNKK 307
P+P+ G +SS HA+ P P+ +PQ P A A + K K
Sbjct: 219 PKPKPGPNSSIFAVHAAEPAKNPVVSTDKPQIPPQPTASTAVRNAGTGKWTVESWKSKKA 398
Query: 308 LQQLRAPKKRQLQA 321
LQ P + +L A
Sbjct: 399 LQLPEYPSQEELDA 440
>CN825244
Length = 646
Score = 26.9 bits (58), Expect = 4.8
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +2
Query: 119 LDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLE 160
+DH VL+LL WL +P +++ + IL FPI LE
Sbjct: 188 MDHSVLSLLLPWLH----FKIPYLHLVRVVRMILC*FPIMLE 301
>AV408530
Length = 432
Score = 26.9 bits (58), Expect = 4.8
Identities = 16/48 (33%), Positives = 27/48 (55%)
Frame = -2
Query: 67 IALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQL 114
I++ NVV EL+ E +AE + GKP + + ++TEV S ++
Sbjct: 188 ISVCGRNVVWELKEEEEHEAEAEQHGKP*MTD-TNVVVVTEVTSNSEI 48
>TC18999 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(6%)
Length = 425
Score = 26.6 bits (57), Expect = 6.3
Identities = 16/39 (41%), Positives = 20/39 (51%)
Frame = +3
Query: 231 KKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPE 269
K P+NK PG QSR S ++ S QSS S P+
Sbjct: 141 KDPSNKDPGDQSRTSS---EVADDLSSQSSKVLEGSAPQ 248
>AV778820
Length = 459
Score = 26.6 bits (57), Expect = 6.3
Identities = 13/38 (34%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Frame = +3
Query: 31 PSSPGEAPQAEEGEEDDEIND-LFKMGKKKNERSPAEI 67
P+ E P +E+G++DD N L + G SPA +
Sbjct: 9 PTMKMEEPSSEDGDDDDFCNSILSRFGNSTARESPAPL 122
>TC15955
Length = 1042
Score = 26.6 bits (57), Expect = 6.3
Identities = 25/79 (31%), Positives = 41/79 (51%), Gaps = 5/79 (6%)
Frame = +1
Query: 150 KILNDFPI---DLEQIDRR--EQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRP 204
K+L D + DL+ ++R E LK S L K+ +S+EEI V+ + +LVD S+
Sbjct: 484 KMLEDVQLADHDLDNMEREVAEVLKASEL-KLQEATKQSEEEIQVHARKLFKLVDSVSQY 660
Query: 205 IFNKSTRFEDMRNIEDERA 223
+ T+ +M+ E A
Sbjct: 661 QEHVGTKVSEMKRDLSETA 717
>BP038233
Length = 508
Score = 26.6 bits (57), Expect = 6.3
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = -2
Query: 58 KKNERSPAEIALLVENVVAELEVTAEEDAELNRQ 91
K+ PA I +++ V + LE+T EE AEL+ Q
Sbjct: 333 KETLNHPANIRNVLDYVASMLEITKEELAELSYQ 232
>BI417263
Length = 539
Score = 26.2 bits (56), Expect = 8.2
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = +2
Query: 282 KVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQAT 322
K DPE++ K+ + + MK + L + R KKRQLQ T
Sbjct: 248 KKDPEQL----KKQIDNLEMMKADGALDKARKHKKRQLQDT 358
>TC8544
Length = 1278
Score = 26.2 bits (56), Expect = 8.2
Identities = 18/61 (29%), Positives = 30/61 (48%), Gaps = 1/61 (1%)
Frame = +2
Query: 235 NKAPGMQSRDSDLDLDLPQP-RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAK 293
N P + DSD D P + + S+ A+ E+ ++ + SKV+ EEV+A K
Sbjct: 347 NPNPQRWTHDSDSDSITTDPVKETEDSASAAAASSESEEVEVGVGFDSKVEGEEVKAIEK 526
Query: 294 Q 294
+
Sbjct: 527 E 529
>TC17889 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein
(AT5g19430/F7K24_180), partial (20%)
Length = 541
Score = 26.2 bits (56), Expect = 8.2
Identities = 15/52 (28%), Positives = 23/52 (43%)
Frame = +2
Query: 256 SGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKK 307
SG SSSRQ + A R + E +A +A H Q +++ +K
Sbjct: 134 SGASSSRQPKKKEAAKSAATTGRRAKRALKREAADKAAEAKHQQHLIRLGRK 289
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.133 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,231,117
Number of Sequences: 28460
Number of extensions: 49265
Number of successful extensions: 236
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 233
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 235
length of query: 334
length of database: 4,897,600
effective HSP length: 91
effective length of query: 243
effective length of database: 2,307,740
effective search space: 560780820
effective search space used: 560780820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0230.8


