

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.4
(606 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
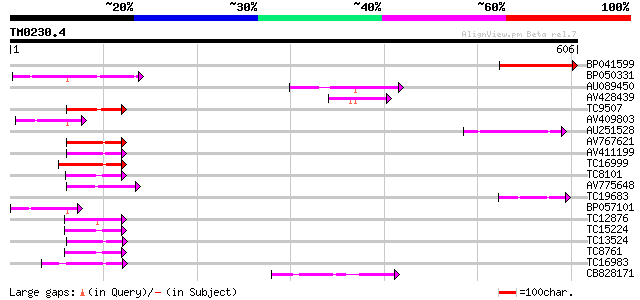
BP041599 176 7e-45
BP050331 70 1e-12
AU089450 69 2e-12
AV428439 61 5e-10
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 59 2e-09
AV409803 56 2e-08
AU251528 52 3e-07
AV767621 50 1e-06
AV411199 49 2e-06
TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein (AT4g391... 49 2e-06
TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial ... 49 2e-06
AV775648 47 7e-06
TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein (AT2G052... 47 7e-06
BP057101 46 2e-05
TC12876 45 3e-05
TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 45 3e-05
TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%) 45 4e-05
TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein P... 45 4e-05
TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravit... 45 4e-05
CB828171 44 1e-04
>BP041599
Length = 540
Score = 176 bits (447), Expect = 7e-45
Identities = 83/83 (100%), Positives = 83/83 (100%)
Frame = -3
Query: 524 EMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELL 583
EMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELL
Sbjct: 538 EMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELL 359
Query: 584 EDCWELDPASLPSYLLTIGGIDN 606
EDCWELDPASLPSYLLTIGGIDN
Sbjct: 358 EDCWELDPASLPSYLLTIGGIDN 290
>BP050331
Length = 519
Score = 69.7 bits (169), Expect = 1e-12
Identities = 53/160 (33%), Positives = 80/160 (49%), Gaps = 20/160 (12%)
Frame = +3
Query: 4 SKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT--- 60
S + E RL +AE + Q + K + + A AQ P LEG +++ + +L A
Sbjct: 45 SGSKSEGERLLEIAEQQXQ-KRDLKGSRQLALIAQENEPLLEGSDQILAIIDVLEAAEKP 221
Query: 61 ---------------DWYTVLGVE--PFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEE 103
D+Y +L V+ + N +++QY++L+LLLHPDKN + + S
Sbjct: 222 LTTTASTTTNNRNNLDFYAILQVDHRDSQDLNLIKRQYRRLALLLHPDKN---LFSFSHH 392
Query: 104 AFKLVGEAFRVLSDSALKKGYDAELRKKEAPTFWTACSAC 143
F LV A+ +LSD A K+ YDA L +FWTAC C
Sbjct: 393 TFDLVSHAWALLSDPAQKEIYDAGLGCAPG-SFWTACPYC 509
>AU089450
Length = 383
Score = 68.9 bits (167), Expect = 2e-12
Identities = 46/130 (35%), Positives = 63/130 (48%), Gaps = 8/130 (6%)
Frame = +3
Query: 300 RGLEIGEVRT-LKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEE 358
R L I + RT ++ ++E + SK + V E K + EA +
Sbjct: 24 RKLLIEKARTEIRKKLEEMKLASKTVAAVNERE----------KSQAEAGQVKSGSHTNT 173
Query: 359 CGDANGGELE-------VMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALI 411
D +G E M V D DF+DFDKDR E F+ Q+WA D++DGMPR Y LI
Sbjct: 174 VLDVSGNHFEHGKIGPVSMTVPDPDFHDFDKDRSEECFRPKQIWAL*DEEDGMPRLYCLI 353
Query: 412 DETVSANPFE 421
E +S NPF+
Sbjct: 354 REVISFNPFK 383
>AV428439
Length = 292
Score = 61.2 bits (147), Expect = 5e-10
Identities = 34/79 (43%), Positives = 46/79 (58%), Gaps = 11/79 (13%)
Frame = +3
Query: 341 IKEKPEASGKRKRLELEECGDA----NGGELE-------VMAVVDSDFYDFDKDRVERSF 389
+KE+ +A +++ E C A G ELE + V DSDF+DFDKDR E F
Sbjct: 54 LKEREKAQADVGQVKRETCRRAVLSVPGLELEHSKAGPISITVPDSDFHDFDKDRAEECF 233
Query: 390 KKGQVWAAYDDDDGMPRHY 408
+ Q+WA YD++DGMPR Y
Sbjct: 234 RPKQIWALYDEEDGMPRLY 290
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 58.9 bits (141), Expect = 2e-09
Identities = 30/65 (46%), Positives = 41/65 (62%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
++Y +LG+E VRK Y+KLSL +HPDKN A +EEAFK V +AF+ L +
Sbjct: 517 NYYEILGLEKSCTVEDVRKSYRKLSLKVHPDKNK---APGAEEAFKAVSKAFQCLGNEES 687
Query: 121 KKGYD 125
K+ YD
Sbjct: 688 KRKYD 702
>AV409803
Length = 413
Score = 55.8 bits (133), Expect = 2e-08
Identities = 29/84 (34%), Positives = 47/84 (55%), Gaps = 8/84 (9%)
Frame = +1
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------ 60
++EA R K +AE K A K+A +A L+P LEG+ + ++ L + +
Sbjct: 160 KDEAARAKEIAEKKFSLREYA-GAKKFALKALNLYPALEGVPQFLSILNVYISAENKING 336
Query: 61 --DWYTVLGVEPFANSNAVRKQYK 82
DWY +LGV+PFA+ +RK+Y+
Sbjct: 337 QMDWYGILGVDPFADEETIRKKYR 408
>AU251528
Length = 388
Score = 52.0 bits (123), Expect = 3e-07
Identities = 36/110 (32%), Positives = 57/110 (51%)
Frame = +3
Query: 486 KGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKR 545
KG +WA+Y + D + ADE D+V L +++E +G+++ L KV G+KTVF
Sbjct: 3 KGDIWAIYRNWSPDWN-ELTADEVMHKFDVVEVLEDFSEEHGVTIIPLVKVAGFKTVF-H 176
Query: 546 QDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLP 595
IR + ++ M SHQ+P+ + E + C LDPA+ P
Sbjct: 177 HHIDPREIRVIPREEMLRFSHQVPSCLLTGL-EAQNAPKGCRVLDPAATP 323
>AV767621
Length = 560
Score = 49.7 bits (117), Expect = 1e-06
Identities = 26/65 (40%), Positives = 40/65 (61%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +L V+ A+ + ++K Y+KL++ HPDKN + A E FK + EA+ VLSD
Sbjct: 61 DYYKILQVDRGASDDDLKKAYRKLAMKWHPDKNPNNKKEA-EAKFKQISEAYDVLSDPQK 237
Query: 121 KKGYD 125
+ YD
Sbjct: 238 RAVYD 252
>AV411199
Length = 423
Score = 49.3 bits (116), Expect = 2e-06
Identities = 25/65 (38%), Positives = 39/65 (59%)
Frame = +2
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +L V+ A ++K Y++L++ HPDKN + A E FKL+ E++ VLSD
Sbjct: 68 DYYEILEVDHHATDEELKKAYRRLAMKWHPDKNPDNKNDA-ETKFKLISESYEVLSDPQK 244
Query: 121 KKGYD 125
+ YD
Sbjct: 245 RAIYD 259
>TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein
(AT4g39150/T22F8_50), partial (35%)
Length = 648
Score = 49.3 bits (116), Expect = 2e-06
Identities = 27/73 (36%), Positives = 45/73 (60%)
Frame = +3
Query: 53 SLTILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAF 112
S ++ + +Y VLGV A++ ++K Y + ++HPDKN AA E F+++GEA+
Sbjct: 102 SQEMVKESAYYDVLGVNVDASAADIKKAYYIKARIVHPDKNPGDPKAA--ENFQMLGEAY 275
Query: 113 RVLSDSALKKGYD 125
+VLSD ++ YD
Sbjct: 276 QVLSDPEKREAYD 314
>TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial (90%)
Length = 1737
Score = 48.9 bits (115), Expect = 2e-06
Identities = 26/66 (39%), Positives = 38/66 (57%)
Frame = +1
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
T +Y VLGV A+ + ++K Y+K ++ HPDK E FK +G+A+ VLSD
Sbjct: 181 TKFYEVLGVPKSASEDEIKKAYRKAAMKNHPDK------GGDPEKFKELGQAYEVLSDPE 342
Query: 120 LKKGYD 125
K+ YD
Sbjct: 343 KKELYD 360
>AV775648
Length = 371
Score = 47.4 bits (111), Expect = 7e-06
Identities = 28/79 (35%), Positives = 42/79 (52%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D Y ++GV ANS+ ++K Y KLSL HPD T+ S++ V A+ +L D A
Sbjct: 139 DCYDLVGVSQNANSSEIKKAYYKLSLKHHPD---TNPDPESKKLLVKVANAYEILKDEAT 309
Query: 121 KKGYDAELRKKEAPTFWTA 139
++ YD + E + TA
Sbjct: 310 REQYDYAIAHPEEVFYNTA 366
>TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein
(AT2G05250/F5G3.15), partial (9%)
Length = 555
Score = 47.4 bits (111), Expect = 7e-06
Identities = 28/77 (36%), Positives = 43/77 (55%)
Frame = +1
Query: 523 NEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPEL 582
+E G+ + L K+ G+KTV+K + AI+++ + M SHQ+P+ E L
Sbjct: 16 SEELGVCVTPLIKLSGFKTVYK-SNADKSAIKWIPRREMLRFSHQVPSWLLK--GEASNL 186
Query: 583 LEDCWELDPASLPSYLL 599
E CW+LDPA+ P LL
Sbjct: 187 PERCWDLDPAATPDDLL 237
>BP057101
Length = 546
Score = 46.2 bits (108), Expect = 2e-05
Identities = 29/85 (34%), Positives = 46/85 (54%), Gaps = 8/85 (9%)
Frame = +3
Query: 1 MAPSKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT 60
M +EEAL+ +AE + + A YA +A+ L PE+EGIS+MV + + A+
Sbjct: 294 MEVQANKEEALQALEIAERRF-AQRDFAGAKSYAVKAKTLCPEVEGISQMVATFEVYIAS 470
Query: 61 --------DWYTVLGVEPFANSNAV 77
D+Y +LG++P A+ AV
Sbjct: 471 EIKHNGELDYYAILGLKPSADKEAV 545
>TC12876
Length = 820
Score = 45.4 bits (106), Expect = 3e-05
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 4/71 (5%)
Frame = -1
Query: 59 ATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDK----NNTHVAAASEEAFKLVGEAFRV 114
+ D+Y VLG+ + +R YKKL+L HPD+ N +++ F+ + EA+ V
Sbjct: 241 SNDFYAVLGLNKECTESELRNAYKKLALKWHPDRCSASGNLKFVEEAKKKFQSIQEAYSV 62
Query: 115 LSDSALKKGYD 125
LSD+ + YD
Sbjct: 61 LSDANKRLMYD 29
>TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (71%)
Length = 1031
Score = 45.1 bits (105), Expect = 3e-05
Identities = 24/67 (35%), Positives = 38/67 (55%)
Frame = +3
Query: 59 ATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
+T +Y +LGV A+ + ++K YKK ++ HPDK E FK + +A+ VLSD
Sbjct: 168 STRYYEILGVPKNASQDDLKKAYKKAAIKNHPDK------GGDPEKFKELAQAYEVLSDP 329
Query: 119 ALKKGYD 125
++ YD
Sbjct: 330 EKREIYD 350
>TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%)
Length = 485
Score = 44.7 bits (104), Expect = 4e-05
Identities = 24/66 (36%), Positives = 37/66 (55%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D Y VLGV + ++ Y+K++L HPDKN AA + FK V ++ +LSD
Sbjct: 292 DPYEVLGVSRSSTDQEIKTAYRKMALKYHPDKNANDPKAA--DMFKEVTFSYNILSDPDK 465
Query: 121 KKGYDA 126
++ YD+
Sbjct: 466 RRQYDS 483
>TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (74%)
Length = 1050
Score = 44.7 bits (104), Expect = 4e-05
Identities = 23/67 (34%), Positives = 38/67 (56%)
Frame = +3
Query: 59 ATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
+T +Y +LGV A+ + ++K YKK ++ HPDK E FK + +A+ VL+D
Sbjct: 111 STRYYDILGVSKTASQDDLKKAYKKAAIKNHPDK------GGDPEKFKELAQAYEVLNDP 272
Query: 119 ALKKGYD 125
++ YD
Sbjct: 273 EKREIYD 293
>TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravity (ARG1
protein), partial (42%)
Length = 716
Score = 44.7 bits (104), Expect = 4e-05
Identities = 29/92 (31%), Positives = 46/92 (49%)
Frame = +2
Query: 35 KRAQRLFPELEGISEMVTSLTILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNN 94
+R + ++EG S V D Y VL V + ++ Y+KL+L HPDKN
Sbjct: 179 ERGAAMGSKMEGTSSPVIR------RDPYEVLSVSRDSTDQEIKTAYRKLALKYHPDKNV 340
Query: 95 THVAAASEEAFKLVGEAFRVLSDSALKKGYDA 126
+ A+ E FK V ++ +LSD ++ YD+
Sbjct: 341 NNPEAS--ELFKEVAYSYSILSDPEKRRQYDS 430
>CB828171
Length = 570
Score = 43.5 bits (101), Expect = 1e-04
Identities = 35/137 (25%), Positives = 57/137 (41%), Gaps = 1/137 (0%)
Frame = +3
Query: 281 EEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRG-SEVGEERSLKKNVKL 339
+ E +T ++ K N G E TLK+ A K + G+ KN+K
Sbjct: 57 QPEGFQTHDKTPLKHDTNNHGNE-----TLKVRRSTRAFSKKTHLGDAGDCSDTDKNMKD 221
Query: 340 AIKEKPEASGKRKRLELEECGDANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYD 399
++ +P+ G + ++C V + YDF K++ + Q+WA Y
Sbjct: 222 SVFSEPD--GSDRAFSKKDCS------------VGASCYDFKKEKSREMLQCDQIWAIYG 359
Query: 400 DDDGMPRHYALIDETVS 416
D D MP +YA I + S
Sbjct: 360 DRDNMPHYYAQIKKIES 410
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.134 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,193,818
Number of Sequences: 28460
Number of extensions: 115595
Number of successful extensions: 664
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 631
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 649
length of query: 606
length of database: 4,897,600
effective HSP length: 96
effective length of query: 510
effective length of database: 2,165,440
effective search space: 1104374400
effective search space used: 1104374400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0230.4


