

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
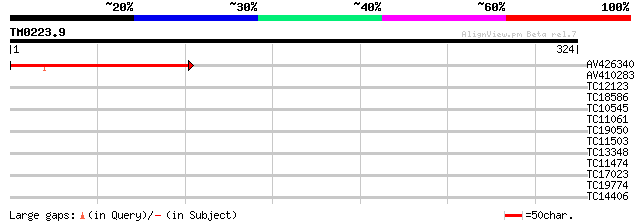
Query= TM0223.9
(324 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV426340 161 2e-40
AV410283 30 0.72
TC12123 similar to GB|AAO39931.1|28372898|BT003703 At4g28730 {Ar... 29 0.94
TC18586 homologue to UP|Q9LTG4 (Q9LTG4) Acetyl-CoA synthetase, p... 28 1.6
TC10545 homologue to UP|Q9LKH8 (Q9LKH8) NADPH-protochlorophyllid... 28 2.7
TC11061 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [va... 28 2.7
TC19050 weakly similar to UP|Q9M6T9 (Q9M6T9) Nonspecific lipid-t... 27 3.6
TC11503 similar to GB|AAO63311.1|28950775|BT005247 At3g02220 {Ar... 27 6.1
TC13348 weakly similar to UP|Q9LUJ4 (Q9LUJ4) Gb|AAF26800.1, part... 27 6.1
TC11474 27 6.1
TC17023 homologue to UP|Q9FM19 (Q9FM19) Hypersensitive-induced r... 26 8.0
TC19774 weakly similar to UP|AAR92268 (AAR92268) At1g47570, part... 26 8.0
TC14406 similar to GB|AAM61587.1|21537246|AY085029 aluminum-indu... 26 8.0
>AV426340
Length = 435
Score = 161 bits (407), Expect = 2e-40
Identities = 84/108 (77%), Positives = 92/108 (84%), Gaps = 3/108 (2%)
Frame = +3
Query: 1 MAPDLTTAATP-ITTTEQP--LATRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVA 57
MAP+L AT +T +P L RA+VTFLAGNGDYVKGVVGLAKGLRKV+S YPLVVA
Sbjct: 111 MAPELVPTATKSVTGFNRPVTLQKRAYVTFLAGNGDYVKGVVGLAKGLRKVKSAYPLVVA 290
Query: 58 ILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLR 105
+LPDVPEEHR+IL SQGCIVREI PVYPP+NQTQFA AYYVINYSKLR
Sbjct: 291 VLPDVPEEHREILESQGCIVREIEPVYPPENQTQFAMAYYVINYSKLR 434
>AV410283
Length = 426
Score = 29.6 bits (65), Expect = 0.72
Identities = 18/56 (32%), Positives = 27/56 (48%), Gaps = 12/56 (21%)
Frame = -3
Query: 54 LVVAILPDVPEEHR------------KILLSQGCIVREIVPVYPPKNQTQFAHAYY 97
L + I+P +P HR +LLSQ + E +P YP KNQ +H ++
Sbjct: 277 LELQIIP-IPSPHRTYQHNFQVKLVYSLLLSQLGFLYETIPYYPSKNQAAVSHQHH 113
>TC12123 similar to GB|AAO39931.1|28372898|BT003703 At4g28730 {Arabidopsis
thaliana;}, partial (47%)
Length = 511
Score = 29.3 bits (64), Expect = 0.94
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = +2
Query: 47 KVQSIYPLVVAILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAH 94
K+ S PL++ +L +P+ +ILL R VP++ N T++ H
Sbjct: 299 KIPSRKPLLITLLLFIPKPGARILLR*NPCSRSSVPIHWSSNWTKWVH 442
>TC18586 homologue to UP|Q9LTG4 (Q9LTG4) Acetyl-CoA synthetase, partial
(27%)
Length = 561
Score = 28.5 bits (62), Expect = 1.6
Identities = 12/34 (35%), Positives = 14/34 (40%)
Frame = -1
Query: 149 PSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLY 182
P W K + CQQCP P N+ P Y
Sbjct: 162 PMWGQCKILKLYICQQCPKNDQHPDNWVPLQTQY 61
>TC10545 homologue to UP|Q9LKH8 (Q9LKH8) NADPH-protochlorophyllide
oxidoreductase, partial (37%)
Length = 787
Score = 27.7 bits (60), Expect = 2.7
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +1
Query: 40 GLAKGLRKVQSIYPLVVAILPDVPEEHRKIL 70
G+ + R VQ +PLV A +P +PE H + L
Sbjct: 160 GMHRHNRLVQRAHPLVQAPVPSIPEVHNQRL 252
>TC11061 similar to PIR|T46895|T46895 acyl-CoA dehydrogenase [validated] -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (30%)
Length = 488
Score = 27.7 bits (60), Expect = 2.7
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = -2
Query: 240 LWRHPENVQLDKVKVVHYCAFGSKPWRFTGEE 271
LW+ N+Q +K+ +C FGS W EE
Sbjct: 148 LWKCSRNIQWW*IKIGRHCPFGSILWILRREE 53
>TC19050 weakly similar to UP|Q9M6T9 (Q9M6T9) Nonspecific lipid-transfer
protein (LTP), partial (57%)
Length = 579
Score = 27.3 bits (59), Expect = 3.6
Identities = 13/45 (28%), Positives = 17/45 (36%)
Frame = -3
Query: 134 MPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPR 178
+P N Y + C C SW Q + +C W S G R
Sbjct: 220 LPKNVPYPIAAC-CRDSWSRVPQVANAWCNSKRKLATWQSRIGER 89
>TC11503 similar to GB|AAO63311.1|28950775|BT005247 At3g02220 {Arabidopsis
thaliana;}, partial (60%)
Length = 975
Score = 26.6 bits (57), Expect = 6.1
Identities = 8/20 (40%), Positives = 12/20 (60%)
Frame = +2
Query: 162 CQQCPDKVDWPSNFGPRPPL 181
C +C D++DW +G PL
Sbjct: 140 CPRCRDQIDWKRRYGKYKPL 199
>TC13348 weakly similar to UP|Q9LUJ4 (Q9LUJ4) Gb|AAF26800.1, partial (7%)
Length = 539
Score = 26.6 bits (57), Expect = 6.1
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = +3
Query: 192 PNLETYHDLLKTC 204
P+LETYH LLK C
Sbjct: 135 PDLETYHPLLKMC 173
>TC11474
Length = 664
Score = 26.6 bits (57), Expect = 6.1
Identities = 14/43 (32%), Positives = 20/43 (45%)
Frame = +2
Query: 145 CFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGF 187
C EPS H+K++ + + K WP F RP + GF
Sbjct: 167 CGLEPSGEHSKKFKMLTFKVLITKSCWPLLFVVRPQVLALIGF 295
>TC17023 homologue to UP|Q9FM19 (Q9FM19) Hypersensitive-induced response
protein, partial (29%)
Length = 510
Score = 26.2 bits (56), Expect = 8.0
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = -2
Query: 260 FGSKPWRFTGEEEN 273
FGS+PW TGE E+
Sbjct: 101 FGSRPWYITGESEH 60
>TC19774 weakly similar to UP|AAR92268 (AAR92268) At1g47570, partial (19%)
Length = 642
Score = 26.2 bits (56), Expect = 8.0
Identities = 11/33 (33%), Positives = 16/33 (48%)
Frame = -2
Query: 130 HLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYC 162
H P Q + CF P W++T Q S+ +C
Sbjct: 380 HCIPQPHQQVLYLDSCF--PLWQNTDQNSLLFC 288
>TC14406 similar to GB|AAM61587.1|21537246|AY085029 aluminum-induced
protein-like {Arabidopsis thaliana;}, partial (66%)
Length = 806
Score = 26.2 bits (56), Expect = 8.0
Identities = 15/44 (34%), Positives = 19/44 (43%), Gaps = 2/44 (4%)
Frame = -2
Query: 174 NFGPRPPLYFNAGFF--VYEPNLETYHDLLKTCEATTPTSFAEQ 215
+F P L F+ G F V E H K C +PT+F Q
Sbjct: 727 DFQP*EHLXFHXGMFKXVQTSRCEKKHPFRKRCRGISPTAFRXQ 596
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,372,761
Number of Sequences: 28460
Number of extensions: 97863
Number of successful extensions: 473
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 473
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 473
length of query: 324
length of database: 4,897,600
effective HSP length: 90
effective length of query: 234
effective length of database: 2,336,200
effective search space: 546670800
effective search space used: 546670800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0223.9


