

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.5
(471 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
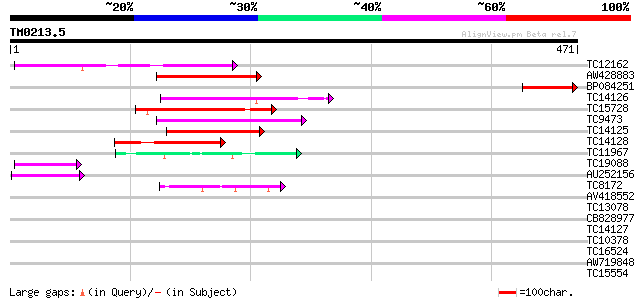
TC12162 similar to AAS09999 (AAS09999) MYB transcription factor,... 105 2e-23
AW428883 102 1e-22
BP084251 99 2e-21
TC14126 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 96 1e-20
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 94 5e-20
TC9473 similar to AAS09986 (AAS09986) MYB transcription factor, ... 94 6e-20
TC14125 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 89 2e-18
TC14128 similar to AAS09980 (AAS09980) MYB transcription factor,... 76 1e-14
TC11967 homologue to UP|AAS09978 (AAS09978) MYB transcription fa... 54 7e-08
TC19088 homologue to UP|Q8S8Z9 (Q8S8Z9) Syringolide-induced prot... 52 2e-07
AU252156 52 3e-07
TC8172 similar to AAS10008 (AAS10008) MYB transcription factor, ... 44 7e-05
AV418552 39 0.001
TC13078 similar to UP|Q7X9Y1 (Q7X9Y1) Gonidia forming protein Gl... 39 0.002
CB828977 39 0.002
TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 38 0.004
TC10378 similar to UP|Q84TG2 (Q84TG2) At3g16350, partial (26%) 37 0.007
TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC... 37 0.009
AW719848 33 0.077
TC15554 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Ar... 33 0.10
>TC12162 similar to AAS09999 (AAS09999) MYB transcription factor, partial
(56%)
Length = 751
Score = 105 bits (262), Expect = 2e-23
Identities = 65/188 (34%), Positives = 100/188 (52%), Gaps = 3/188 (1%)
Frame = +1
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG--PA 61
WTP +++ E ALA D PDRW ++A+ + G+ +++++Y+ L V IE G P
Sbjct: 244 WTPQENKLFENALAYFDKDTPDRWLRVASMIPGKTVGDVIKQYKELEEDVCVIEAGLVPV 423
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
P + D N+ +I++G + V PS ++
Sbjct: 424 PGYGADSFTLEWANNHQGY---------DIDDGFKQFYSVGGKRGASCTRPSEQER---- 564
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
+K V WT++EHR FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R
Sbjct: 565 ------KKGVPWTEEEHRLFLLGLKKYGKGDWRNISRNFVTTRTPTQVASHAQKYFIRQL 726
Query: 182 STPKRRKR 189
+ K ++R
Sbjct: 727 TGGKDKRR 750
>AW428883
Length = 430
Score = 102 bits (255), Expect = 1e-22
Identities = 46/88 (52%), Positives = 66/88 (74%), Gaps = 1/88 (1%)
Frame = +1
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
S + K + WT++EHR FL GL KYGKG W++ISR F+ ++TPTQ+ASHAQKY++RLNS
Sbjct: 109 SDQERRKGIAWTEEEHRLFLLGLDKYGKGDWRSISRNFVVTRTPTQVASHAQKYFIRLNS 288
Query: 183 TPKRRKRASIHDLT-IDDTDLVPQQNQV 209
K R+R+SIHD+T +++ D+ Q +
Sbjct: 289 MNKDRRRSSIHDITSVNNGDISAPQGPI 372
>BP084251
Length = 379
Score = 98.6 bits (244), Expect = 2e-21
Identities = 45/45 (100%), Positives = 45/45 (100%)
Frame = -3
Query: 427 VLASQQHPVPPLEVMPMQQLQERPQEHQAVNSRHFVWKPVGNRGR 471
VLASQQHPVPPLEVMPMQQLQERPQEHQAVNSRHFVWKPVGNRGR
Sbjct: 377 VLASQQHPVPPLEVMPMQQLQERPQEHQAVNSRHFVWKPVGNRGR 243
Score = 35.8 bits (81), Expect = 0.015
Identities = 18/22 (81%), Positives = 19/22 (85%), Gaps = 1/22 (4%)
Frame = -3
Query: 370 LASQQHMVPPLDA-PMQQLQER 390
LASQQH VPPL+ PMQQLQER
Sbjct: 374 LASQQHPVPPLEVMPMQQLQER 309
>TC14126 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(44%)
Length = 1500
Score = 95.9 bits (237), Expect = 1e-20
Identities = 54/150 (36%), Positives = 88/150 (58%), Gaps = 6/150 (4%)
Frame = +2
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ ++ V WT++EH+ FL GL+K GKG W+ ISR ++ ++TPTQ+ASHAQKY+LR ++ +
Sbjct: 350 ERKRGVPWTEEEHKLFLVGLQKVGKGDWRGISRNYVKTRTPTQVASHAQKYFLRRSNLNR 529
Query: 186 RRKRASIHDLTIDDTDLV-----PQQNQVPQLNA-PMQELSPQQNGVQLDAPMQQVSSQQ 239
RR+R+S+ D+T D + QNQ ++ PM + + +G + P+ Q +
Sbjct: 530 RRRRSSLFDITTDTVSAIQVEEEQVQNQDTMFHSQPMCPETSKISGFPMMPPVYQFGVNE 709
Query: 240 NQVDALDFPMQQLSFQQNGDPPLNFPHQVF 269
+ PM++LS Q P N P +F
Sbjct: 710 S-------PMEELSLGQGNMKP-NVPTNLF 775
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 94.0 bits (232), Expect = 5e-20
Identities = 50/120 (41%), Positives = 77/120 (63%), Gaps = 3/120 (2%)
Frame = +1
Query: 105 PEPVPNDPS--SSQKITKKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFL 161
PE P D + +S + + ++ KR V WT++EH+ FL GL K GKG W+ ISR F+
Sbjct: 370 PESNPADAAGYASDDVVHPSARARDRKRGVPWTEEEHKLFLLGLHKVGKGDWRGISRNFV 549
Query: 162 PSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSP 221
++TPTQ+ASHAQKY+LR ++ +RR+R+S+ D+T TD V + + + + QE+ P
Sbjct: 550 KTRTPTQVASHAQKYFLRRHNHNRRRRRSSLFDIT---TDTVMESSTIMEEEQDQQEMVP 720
>TC9473 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(50%)
Length = 887
Score = 93.6 bits (231), Expect = 6e-20
Identities = 43/124 (34%), Positives = 73/124 (58%)
Frame = +3
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
S + +K WT++EHR FL GL+K GKG W+ I+R ++ S+TPTQ+ASHAQKY++R ++
Sbjct: 450 SSRERKKGTPWTEEEHRMFLLGLQKLGKGDWRGIARNYVISRTPTQVASHAQKYFIRQSN 629
Query: 183 TPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQV 242
+R++R+S+ D+ DD P Q +Q + N + P+ + +
Sbjct: 630 VSRRKRRSSLFDIVADDASDTPMVEQDFLSANQLQTETEGNNPLPAPPPIDEECESMDST 809
Query: 243 DALD 246
+++D
Sbjct: 810 NSID 821
>TC14125 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(40%)
Length = 601
Score = 88.6 bits (218), Expect = 2e-18
Identities = 38/81 (46%), Positives = 59/81 (71%)
Frame = +3
Query: 131 VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRRKRA 190
V WT++EH+ FL GL+K GKG W+ ISR ++ ++TPTQ+ASHAQKY+LR ++ +RR+R+
Sbjct: 333 VPWTEEEHKLFLVGLQKVGKGDWRGISRNYVKTRTPTQVASHAQKYFLRRSNLNRRRRRS 512
Query: 191 SIHDLTIDDTDLVPQQNQVPQ 211
S+ D+T D + + + Q
Sbjct: 513 SLFDITTDTVSAIQVEEEQVQ 575
>TC14128 similar to AAS09980 (AAS09980) MYB transcription factor, partial
(36%)
Length = 423
Score = 76.3 bits (186), Expect = 1e-14
Identities = 38/92 (41%), Positives = 56/92 (60%)
Frame = +1
Query: 88 SPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKK 147
+PN +E A D P P+P K++ ++ V WT++EH+ FL GL+K
Sbjct: 160 APNNKEHAAGYASADEAPAPLP----------VKLANPDRKRGVPWTEEEHKLFLLGLQK 309
Query: 148 YGKGQWQTISREFLPSKTPTQIASHAQKYYLR 179
GKG W+ IS+ F+ ++T TQ+ASHAQKY+LR
Sbjct: 310 VGKGDWRGISKNFVKTRTSTQVASHAQKYFLR 405
>TC11967 homologue to UP|AAS09978 (AAS09978) MYB transcription factor,
partial (32%)
Length = 664
Score = 53.5 bits (127), Expect = 7e-08
Identities = 45/163 (27%), Positives = 66/163 (39%), Gaps = 9/163 (5%)
Frame = +3
Query: 89 PNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNQN--EKRVLWTDDEHRNFLRGLK 146
P + P P P P ++++ +KKI + R WT+ EH FL L+
Sbjct: 132 PGVNSLPPP-------PPPPSTTAAAAEDPSKKIRKPYTITKSRESWTEQEHDKFLEALQ 290
Query: 147 KYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST-------PKRRKRASIHDLTIDD 199
+ + W+ I F+ SKT QI SHAQKY+L++ P R KR + H
Sbjct: 291 LFDR-DWKKIEA-FIGSKTVIQIRSHAQKYFLKVQKNGTSEHVPPPRPKRKAAH------ 446
Query: 200 TDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQV 242
P + P+ Q P Q+ P SS + V
Sbjct: 447 ----PYPQKAPKNAPTSQATGPLQSSSAFIEPAYIYSSDSSSV 563
>TC19088 homologue to UP|Q8S8Z9 (Q8S8Z9) Syringolide-induced protein 1-3-1B,
partial (26%)
Length = 353
Score = 52.4 bits (124), Expect = 2e-07
Identities = 27/56 (48%), Positives = 33/56 (58%), Gaps = 1/56 (1%)
Frame = +3
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
WT D+ E AL VP+D PDRWE+IA V G+ AE+ E Y LV V I+ G
Sbjct: 183 WTRYHDKLFERALLAVPEDLPDRWEKIAEQVPGKSAAEVREHYEALVHDVFEIDSG 350
>AU252156
Length = 350
Score = 51.6 bits (122), Expect = 3e-07
Identities = 27/62 (43%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +2
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SW+ ++A E ALA PDRW +A AV G+ P E+ + Y LLV V IE G
Sbjct: 155 SGSWSAKDNKAFERALAVYDKVTPDRWYNVAHAVGGKTPEEVKKHYELLVQDVKQIESGQ 334
Query: 61 AP 62
P
Sbjct: 335 VP 340
>TC8172 similar to AAS10008 (AAS10008) MYB transcription factor, partial
(30%)
Length = 1353
Score = 43.5 bits (101), Expect = 7e-05
Identities = 36/113 (31%), Positives = 51/113 (44%), Gaps = 8/113 (7%)
Frame = +1
Query: 125 NQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISR--EFLPSKTPTQIASHAQKYYLRLNS 182
NQ +K WT +E RG++KYG G+W+ I + EF PS T K + LN
Sbjct: 106 NQKQK---WTAEEEDALHRGVQKYGAGKWKNILKDPEFAPSLTSRSNIDLKDK-WRNLNV 273
Query: 183 TPKR----RKRASIHDLTIDDTDLVPQQNQVPQLN--APMQELSPQQNGVQLD 229
+ + R S LT T + QQN P + A + +P QN + D
Sbjct: 274 VTGQGSNIKSRTSKPKLTAPSTPVPNQQNATPSVQNVATPKVPTPSQNSAEKD 432
>AV418552
Length = 386
Score = 39.3 bits (90), Expect = 0.001
Identities = 35/110 (31%), Positives = 48/110 (42%), Gaps = 7/110 (6%)
Frame = +2
Query: 354 PLNFPMQQLQEIQQ--SGLASQQHMVPPLDAPMQ----QLQERQHTLGDSGLTS-QQLMV 406
P + Q +Q+ QQ S QQ ++P L Q QL + Q L LT QQ +
Sbjct: 44 PSSQEQQLMQQHQQMLSYFQPQQQLLPHLQLQQQLQQQQLAQLQQQLSQLQLTLLQQQQL 223
Query: 407 PPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEVMPMQQLQERPQEHQAV 456
L QQL + + L L QH +++P QQ Q+ PQ Q V
Sbjct: 224 TQLQ--QQQLAQLQQQLAQLQLTQHQHTQLQQQLLPQQQFQQLPQHQQMV 367
>TC13078 similar to UP|Q7X9Y1 (Q7X9Y1) Gonidia forming protein GlsA, partial
(9%)
Length = 564
Score = 38.9 bits (89), Expect = 0.002
Identities = 17/33 (51%), Positives = 22/33 (66%)
Frame = +3
Query: 3 DSWTPAQDRALELALATVPDDAPDRWEQIAAAV 35
D W+ Q+RAL AL P +A RWE++AAAV
Sbjct: 162 DIWSAVQERALVQALKAFPKEASQRWERVAAAV 260
>CB828977
Length = 528
Score = 38.5 bits (88), Expect = 0.002
Identities = 37/119 (31%), Positives = 53/119 (44%), Gaps = 15/119 (12%)
Frame = +1
Query: 125 NQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISR--EFLPSKTPTQIASHAQKYYLRLN- 181
NQ +K WT +E RG++KYG G+W+ I + EF PS T K + LN
Sbjct: 58 NQKQK---WTAEEEDALHRGVQKYGAGKWKNILKDPEFAPSLTSRSNIDLKDK-WRNLNV 225
Query: 182 ----------STPKRRKRASIHDLTIDDT--DLVPQQNQVPQLNAPMQELSPQQNGVQL 228
T K + A T D T D+ P QN P++ P Q S + + V++
Sbjct: 226 GTGQGSNVKSRTLKPKLPAPCAVTTPDPTVQDVAPVQNATPKI--PSQNSSEKDHDVKV 396
>TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(21%)
Length = 556
Score = 37.7 bits (86), Expect = 0.004
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = +2
Query: 131 VLWTDDEHRNFLRGLKKYGKGQWQTI 156
V WT++EH+ FL GL+K GKG W+ I
Sbjct: 479 VPWTEEEHKLFLVGLQKVGKGDWRGI 556
>TC10378 similar to UP|Q84TG2 (Q84TG2) At3g16350, partial (26%)
Length = 442
Score = 37.0 bits (84), Expect = 0.007
Identities = 18/46 (39%), Positives = 28/46 (60%)
Frame = +2
Query: 106 EPVPNDPSSSQKITKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKG 151
E + +DP+ + + + +K V WT++EHR FL GL+K GKG
Sbjct: 308 EYLSDDPAHASTFANR--RGERKKGVPWTEEEHRLFLIGLQKLGKG 439
>TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC19_9
{Arabidopsis thaliana;}, partial (41%)
Length = 997
Score = 36.6 bits (83), Expect = 0.009
Identities = 19/43 (44%), Positives = 27/43 (62%), Gaps = 1/43 (2%)
Frame = +1
Query: 4 SWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMER 45
+W+ +D AL AL P DAP RWE++A AV G+ A ++R
Sbjct: 631 NWSNGEDIALLNALKAFPKDAPMRWEKVAVAVPGKSKAACVKR 759
>AW719848
Length = 614
Score = 33.5 bits (75), Expect = 0.077
Identities = 20/72 (27%), Positives = 38/72 (52%), Gaps = 3/72 (4%)
Frame = +2
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLL--VLHVDNIEEGPA 61
WT + E A++ D PDRW ++AA + G+ +++++++ L +L ++ GP
Sbjct: 356 WTREDNEKFESAVSIYDKDTPDRWLKVAAMIPGKTVFDVIKKFKELEDILGIE-AGHGPI 532
Query: 62 PDFIVDRVPEPV 73
P + R P V
Sbjct: 533 PATVRVRGPNHV 568
>TC15554 similar to GB|AAO24533.1|27808506|BT003101 At2g41420 {Arabidopsis
thaliana;}, partial (77%)
Length = 679
Score = 33.1 bits (74), Expect = 0.10
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 1/58 (1%)
Frame = +1
Query: 278 LNVQQLSFQQNGDPPLNFPH-QVYPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQ 334
+N+ QQ PP+ P Q YP ++ P + G PP +P Q YPQQ
Sbjct: 73 INMSYYDHQQQ--PPVGVPPPQGYPSKDAYPPPGYPAQGYPPAGYPPQGYPQQGYPQQ 240
Score = 28.5 bits (62), Expect = 2.5
Identities = 24/86 (27%), Positives = 33/86 (37%), Gaps = 1/86 (1%)
Frame = +1
Query: 309 LNVQQLSFQQNGDPPLNFPH-QVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQ 367
+N+ QQ PP+ P Q YP ++ P + G+PP +P Q
Sbjct: 73 INMSYYDHQQQ--PPVGVPPPQGYPSKDAYPPPGYPAQGYPPAGYPPQGYPQQ------- 225
Query: 368 SGLASQQHMVPPLDAPMQQLQERQHT 393
PP A QQ Q+RQ T
Sbjct: 226 ----GYPQQYPPQYA--QQPQQRQET 285
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,620,910
Number of Sequences: 28460
Number of extensions: 145034
Number of successful extensions: 786
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 616
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 771
length of query: 471
length of database: 4,897,600
effective HSP length: 94
effective length of query: 377
effective length of database: 2,222,360
effective search space: 837829720
effective search space used: 837829720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0213.5


