

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.15
(1070 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
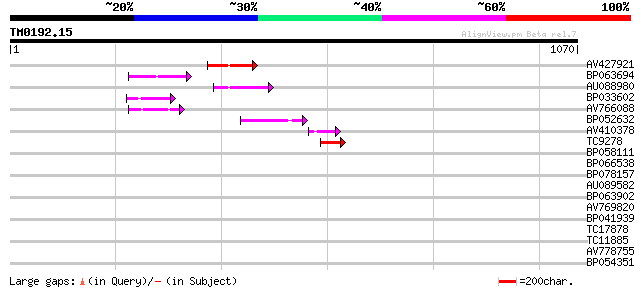
Score E
Sequences producing significant alignments: (bits) Value
AV427921 100 9e-22
BP063694 88 6e-18
AU088980 63 3e-10
BP033602 54 1e-07
AV766088 54 2e-07
BP052632 51 1e-06
AV410378 47 2e-05
TC9278 42 7e-04
BP058111 38 0.010
BP066538 32 0.013
BP078157 36 0.028
AU089582 34 0.11
BP063902 34 0.11
AV769820 32 0.53
BP041939 32 0.69
TC17878 similar to GB|AAK91488.1|15215889|AY050475 AT5g02770/F9G... 31 0.90
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 30 1.5
AV778755 30 2.6
BP054351 29 4.5
>AV427921
Length = 284
Score = 100 bits (250), Expect = 9e-22
Identities = 51/95 (53%), Positives = 63/95 (65%)
Frame = +1
Query: 374 RQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAV 433
RQ T AY PQQNGVSERK RTI+ M RS+L G+ +F EAV +++ILNR PT V
Sbjct: 1 RQ*TAAYTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFL-PEAVMWSLHILNRSPTLVV 177
Query: 434 QNKTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKR 468
QN P EAW G++P+ H R+ + Y HVPD KR
Sbjct: 178 QNMMPEEAWSGRQPAVDHFRISRCLAYAHVPDQKR 282
>BP063694
Length = 511
Score = 88.2 bits (217), Expect = 6e-18
Identities = 46/119 (38%), Positives = 66/119 (54%)
Frame = -2
Query: 224 LWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQRRVSFSTCKEWRAKDV 283
LWH R GH N ++K L++ + D+ S+ C C + +R+ F + RA V
Sbjct: 360 LWHSRLGHVNFDIIKQLHKHGCL-DVSSILPKPICCTSCQMAKSKRLVFHDNNK-RASAV 187
Query: 284 LELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVFKKFKALVEKQ 342
L+LIH D+ GP +S D YF++F DDFSR TW Y LK KS+ V +FK +E +
Sbjct: 186 LDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFMENR 10
>AU088980
Length = 360
Score = 62.8 bits (151), Expect = 3e-10
Identities = 33/113 (29%), Positives = 59/113 (52%)
Frame = +2
Query: 385 NGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCG 444
N + ERK R I+ + R+++ + W AV A Y++N P+ ++ + P +
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLW-TFAVKHAAYLINXLPSPLLKGQCPYQFLNN 178
Query: 445 KKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQT 497
P+ L+VFG++CY+ KL+ + + +FLG+ T + GY V +L T
Sbjct: 179 DSPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDLPT 337
>BP033602
Length = 533
Score = 53.9 bits (128), Expect = 1e-07
Identities = 32/94 (34%), Positives = 49/94 (52%), Gaps = 1/94 (1%)
Frame = +3
Query: 220 DDSWLWHRRFGHFNTQVLKLLYQKNMMRDL-PSLKESNEACERCLLGMQRRVSFSTCKEW 278
D LWH R GH N Q L+ L+ DL ++ S+ CE C+L +R + + + +
Sbjct: 264 DQIMLWHNRLGHPNFQYLRHLFP-----DLFKNVNCSSLECESCVLAKNQRAPYYS-QPY 425
Query: 279 RAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDD 312
A LIH+DV GP + ++ R+F+ F DD
Sbjct: 426 HASRPFYLIHSDVWGPSKITTQFGKRWFVTFIDD 527
>AV766088
Length = 501
Score = 53.5 bits (127), Expect = 2e-07
Identities = 31/107 (28%), Positives = 54/107 (49%), Gaps = 1/107 (0%)
Frame = -3
Query: 224 LWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQRRVSFSTCKEWRAKDV 283
LWH R GH ++ L+ L + L + CE C+ G R+ F +
Sbjct: 307 LWHSRLGHPSSSALRYLRSNKFISY--ELLNYSPVCESCVFGKHVRLPFVSSNNVTVMP- 137
Query: 284 LELIHTDV-CGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVF 329
+++H+D+ P+ +S+ +R+++LF DDF+ W + L KS+VF
Sbjct: 136 FDILHSDLWTSPVLSSA--GHRFYVLFLDDFTDFLWTFPLSNKSQVF 2
>BP052632
Length = 489
Score = 50.8 bits (120), Expect = 1e-06
Identities = 36/131 (27%), Positives = 65/131 (49%), Gaps = 5/131 (3%)
Frame = -3
Query: 436 KTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNL 495
KTP E W G++P+ + FG + + K ++K + I LGYS+ SK Y V+N
Sbjct: 487 KTPYELWRGRRPTVSYFHPFGCK*FILNTNDNIGKFDNKFDKRILLGYSSNSKAYIVFNS 308
Query: 496 QTKKLTISRDVEVDG---NASWNWDEEKVEKNILIPTQRPQEEVEEEAENPGEP--TSPP 550
+T+ + S +V+ D A++ +++ + ++ ++ EE N G+P TS
Sbjct: 307 RTQVVEESINVKFDD*SIEATYRLEKDFLNLDL-------SDQDEEVPTNEGQPSETSHE 149
Query: 551 QQQEQQQDLSP 561
+ Q Q +P
Sbjct: 148 EPSTQPQTTNP 116
>AV410378
Length = 358
Score = 46.6 bits (109), Expect = 2e-05
Identities = 23/60 (38%), Positives = 35/60 (58%)
Frame = +1
Query: 564 TPRRVRFLVDVYETCNLAILEPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGK 623
TP F++++ + EP + A K E W KAM++EI+ +E+N+TW LVD P K
Sbjct: 193 TPSYRAFIMNI-----TTVSEPTRYSEAVKHECWRKAMDQEIEALERNHTWILVDKPPDK 357
>TC9278
Length = 743
Score = 41.6 bits (96), Expect = 7e-04
Identities = 23/47 (48%), Positives = 29/47 (60%)
Frame = +1
Query: 587 SFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYK 633
+FEA + Q W K M EI +IEKN T EL D + +I VKWV+K
Sbjct: 574 TFEALNGQH-WRKVMAGEIHVIEKNET*ELTDLLGDQRLISVKWVHK 711
>BP058111
Length = 570
Score = 37.7 bits (86), Expect = 0.010
Identities = 22/69 (31%), Positives = 37/69 (52%)
Frame = -1
Query: 221 DSWLWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQRRVSFSTCKEWRA 280
D LWH R GH +++ L +L QKN + K+ C+ C + Q+++SF +
Sbjct: 195 DCNLWHLRLGHTSSKKLAIL-QKNF--PFITCKKITSPCDTCHMAKQKKLSFPN-SVTLS 28
Query: 281 KDVLELIHT 289
++ +LIHT
Sbjct: 27 SEIFDLIHT 1
>BP066538
Length = 396
Score = 32.0 bits (71), Expect(2) = 0.013
Identities = 16/31 (51%), Positives = 20/31 (63%)
Frame = -3
Query: 127 VESKGKGTVMVET*KGTRLISDVLLVPNLKE 157
V+ K K V VET GT+ I DVLLVP + +
Sbjct: 145 VDVKAK*VVAVETSSGTKYIPDVLLVPEINQ 53
Score = 24.3 bits (51), Expect(2) = 0.013
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = -1
Query: 98 SNHMAKDESIFKNIDDSVKVKV 119
S+HM DE +FK +D KV
Sbjct: 234 SHHMTYDEKLFKELDQPFVSKV 169
>BP078157
Length = 395
Score = 36.2 bits (82), Expect = 0.028
Identities = 19/49 (38%), Positives = 24/49 (48%)
Frame = -2
Query: 224 LWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQRRVSF 272
LWH R GH + +VLK++ LP N C C L RR+SF
Sbjct: 148 LWHSRLGHLSDKVLKVVSNLVPFTVLPDFHSHN--CNVCPLSKMRRLSF 8
>AU089582
Length = 383
Score = 34.3 bits (77), Expect = 0.11
Identities = 17/54 (31%), Positives = 31/54 (56%)
Frame = +2
Query: 352 SDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKE 405
SDRG ++TS + F G +++ A+ PQ +G SER + + +M R+ + +
Sbjct: 80 SDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQILEDMLRACVXD 241
>BP063902
Length = 494
Score = 34.3 bits (77), Expect = 0.11
Identities = 18/53 (33%), Positives = 27/53 (49%)
Frame = -3
Query: 222 SWLWHRRFGHFNTQVLKLLYQKNMMRDLPSLKESNEACERCLLGMQRRVSFST 274
S +WH R GH + L L ++ PS S+ C+ C+LG R+ FS+
Sbjct: 189 SSVWHHRLGHPASSALNHLRNNKLIFCEPS--RSSSVCDSCVLGKHVRLPFSS 37
>AV769820
Length = 316
Score = 32.0 bits (71), Expect = 0.53
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = -2
Query: 43 CNYCKKFGHMEKYCYSKNRHQANLAEEQEQD 73
CN C K GH+ YC+ + QAN + E D
Sbjct: 261 CNVCGKKGHLSIYCWRRCEKQANERKAVEDD 169
>BP041939
Length = 392
Score = 31.6 bits (70), Expect = 0.69
Identities = 17/34 (50%), Positives = 24/34 (70%)
Frame = +1
Query: 470 KLEDKNVRGIFLGYSTKSKGYRVYNLQTKKLTIS 503
KL+ K+++ +FLGYS KGYRV +LQ T+S
Sbjct: 40 KLDPKSLKCVFLGYSRT*KGYRV-SLQILVATLS 138
>TC17878 similar to GB|AAK91488.1|15215889|AY050475 AT5g02770/F9G14_80
{Arabidopsis thaliana;}, partial (17%)
Length = 465
Score = 31.2 bits (69), Expect = 0.90
Identities = 19/45 (42%), Positives = 20/45 (44%), Gaps = 3/45 (6%)
Frame = +2
Query: 524 NILIPTQRPQE---EVEEEAENPGEPTSPPQQQEQQQDLSPESTP 565
N PTQ E EEEAE P P S PQ Q PE+ P
Sbjct: 80 NDTTPTQENPNKTLEPEEEAEEPTPPQSEPQSQPDPIAAEPETEP 214
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 30.4 bits (67), Expect = 1.5
Identities = 22/67 (32%), Positives = 30/67 (43%), Gaps = 14/67 (20%)
Frame = +2
Query: 4 RRESSRNFSKNSQDKN-----------PPCSIC*RLG---HAEADCRYRDKPQCNYCKKF 49
RR+S R FS+++ KN P +IC G H ++C K C CK+
Sbjct: 305 RRDSRRGFSRDNLCKNCKRPGHFARECPNVAICHNCGLPGHIASECT--TKSLCWNCKEP 478
Query: 50 GHMEKYC 56
GHM C
Sbjct: 479 GHMASSC 499
>AV778755
Length = 537
Score = 29.6 bits (65), Expect = 2.6
Identities = 26/74 (35%), Positives = 35/74 (47%), Gaps = 5/74 (6%)
Frame = +1
Query: 950 R*HEKHFRLCFLPWIGNIFLDV-*EASYSCTINI*S*VCGSC*SNK----SSYMASKNTR 1004
+*HE++ R+CFL IGN + *E * N+ S + A++NT
Sbjct: 250 K*HEENLRICFLSGIGNSLSHLR*ELQ---------------*HNQQQK*SEWRATENT* 384
Query: 1005 RYGRKTR*VYYTQL 1018
RY RKT YY L
Sbjct: 385 RYERKTGCAYYNLL 426
>BP054351
Length = 431
Score = 28.9 bits (63), Expect = 4.5
Identities = 25/88 (28%), Positives = 40/88 (45%), Gaps = 6/88 (6%)
Frame = -1
Query: 182 YDKHDKRVEIAQVKMQKSNISFPLNFKYVA-----NIAMKAQVDDSWLWHRRFGHFNTQV 236
YD+ ++ IA KM+ F F+ A ++ + D LWH R GH N Q
Sbjct: 248 YDRISGKM-IASAKMKDGLYYFEDTFRNKAAQGLCGMSCTSVRDQIMLWHNRLGHPNFQ- 75
Query: 237 LKLLYQKNMMRDL-PSLKESNEACERCL 263
Y +++ DL ++ S+ CE C+
Sbjct: 74 ----YLRHLFPDLFKNVNCSSLECESCV 3
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,143,602
Number of Sequences: 28460
Number of extensions: 271796
Number of successful extensions: 1682
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 1648
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1672
length of query: 1070
length of database: 4,897,600
effective HSP length: 100
effective length of query: 970
effective length of database: 2,051,600
effective search space: 1990052000
effective search space used: 1990052000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0192.15


