

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.14
(75 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
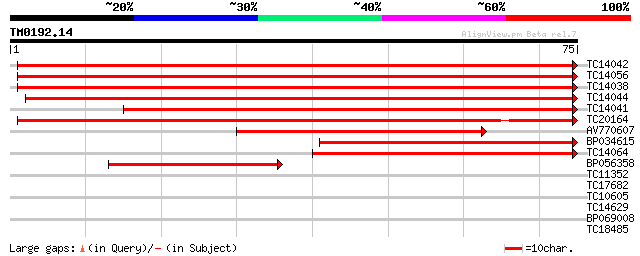
Score E
Sequences producing significant alignments: (bits) Value
TC14042 homologue to UP|Q9XEW9 (Q9XEW9) Elongation factor 1-alph... 139 8e-35
TC14056 homologue to UP|P93769 (P93769) Elongation factor-1 alph... 133 5e-33
TC14038 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%) 133 5e-33
TC14044 similar to UP|Q9LN13 (Q9LN13) T6D22.2, partial (10%) 128 2e-31
TC14041 homologue to UP|Q9XEW9 (Q9XEW9) Elongation factor 1-alph... 117 4e-28
TC20164 UP|Q8LPC4 (Q8LPC4) Elongation factor 1-alpha (Elongation... 98 3e-22
AV770607 58 4e-10
BP034615 53 1e-08
TC14064 similar to UP|EF1A_MAIZE (Q41803) Elongation factor 1-al... 51 5e-08
BP056358 45 3e-06
TC11352 27 0.93
TC17682 similar to UP|AAS18240 (AAS18240) Enolase , partial (68%) 26 1.6
TC10605 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor Tu... 25 2.1
TC14629 homologue to UP|EFT2_SOYBN (P46280) Elongation factor Tu... 25 2.1
BP069008 25 2.1
TC18485 similar to UP|Q9LEU4 (Q9LEU4) CCR4-associated factor-lik... 23 7.8
>TC14042 homologue to UP|Q9XEW9 (Q9XEW9) Elongation factor 1-alpha 1,
partial (38%)
Length = 800
Score = 139 bits (351), Expect = 8e-35
Identities = 70/74 (94%), Positives = 71/74 (95%)
Frame = +2
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSVEKK
Sbjct: 293 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVEKK 472
Query: 62 EPSGAKVTKAALKK 75
+PSGAKVTKAALKK
Sbjct: 473 DPSGAKVTKAALKK 514
>TC14056 homologue to UP|P93769 (P93769) Elongation factor-1 alpha, complete
Length = 1897
Score = 133 bits (335), Expect = 5e-33
Identities = 66/74 (89%), Positives = 69/74 (93%)
Frame = +1
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM+PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSVEKK
Sbjct: 1225 KEIEKEPKFLKNGDAGMVKMVPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVEKK 1404
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA KK
Sbjct: 1405 DPTGAKVTKAAQKK 1446
>TC14038 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%)
Length = 1844
Score = 133 bits (335), Expect = 5e-33
Identities = 66/74 (89%), Positives = 69/74 (93%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
KEIEKEPKFLKNGDAG VKM+PTKPMVVETFSEYP LGRFAVRDMRQTV GVIKSVEKK
Sbjct: 1206 KEIEKEPKFLKNGDAGMVKMLPTKPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVEKK 1385
Query: 62 EPSGAKVTKAALKK 75
+P+GAKVTKAA KK
Sbjct: 1386 DPTGAKVTKAAAKK 1427
>TC14044 similar to UP|Q9LN13 (Q9LN13) T6D22.2, partial (10%)
Length = 550
Score = 128 bits (321), Expect = 2e-31
Identities = 63/73 (86%), Positives = 68/73 (92%)
Frame = +2
Query: 3 EIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKE 62
E+EKEP+FLKNGDAG VKMIPT+PMVVETFSEYP LGRFAVRDMRQTV GVIKSVEKK+
Sbjct: 44 ELEKEPQFLKNGDAGLVKMIPTQPMVVETFSEYPPLGRFAVRDMRQTVAVGVIKSVEKKD 223
Query: 63 PSGAKVTKAALKK 75
P+GAKVTKAA KK
Sbjct: 224 PTGAKVTKAAQKK 262
>TC14041 homologue to UP|Q9XEW9 (Q9XEW9) Elongation factor 1-alpha 1,
partial (34%)
Length = 777
Score = 117 bits (293), Expect = 4e-28
Identities = 59/60 (98%), Positives = 60/60 (99%)
Frame = +1
Query: 16 AGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
+GFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK
Sbjct: 277 SGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 456
>TC20164 UP|Q8LPC4 (Q8LPC4) Elongation factor 1-alpha (Elongation
factor-1a), partial (29%)
Length = 631
Score = 97.8 bits (242), Expect = 3e-22
Identities = 49/74 (66%), Positives = 59/74 (79%)
Frame = +3
Query: 2 KEIEKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVEKK 61
K++E PK +K+GDA VKM+ +KPM VE F++YP LGRFAVRDMRQTV GVIKSVEKK
Sbjct: 180 KKLEDTPKMIKSGDAAMVKMVASKPMCVEAFTQYPPLGRFAVRDMRQTVAVGVIKSVEKK 359
Query: 62 EPSGAKVTKAALKK 75
E G K+TK+A KK
Sbjct: 360 EVEG-KMTKSAAKK 398
>AV770607
Length = 447
Score = 57.8 bits (138), Expect = 4e-10
Identities = 27/33 (81%), Positives = 29/33 (87%)
Frame = -1
Query: 31 TFSEYPLLGRFAVRDMRQTVVTGVIKSVEKKEP 63
TFSEYP LGRFAVRDMRQTV GVI+ VEKK+P
Sbjct: 447 TFSEYPPLGRFAVRDMRQTVAVGVIRRVEKKDP 349
>BP034615
Length = 287
Score = 52.8 bits (125), Expect = 1e-08
Identities = 25/34 (73%), Positives = 31/34 (90%)
Frame = -2
Query: 42 AVRDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
AVRDMRQTV GVI+SVEKK+P+GA+VT+AA +K
Sbjct: 286 AVRDMRQTVAVGVIQSVEKKDPTGAQVTQAAAQK 185
>TC14064 similar to UP|EF1A_MAIZE (Q41803) Elongation factor 1-alpha
(EF-1-alpha), partial (8%)
Length = 523
Score = 50.8 bits (120), Expect = 5e-08
Identities = 24/35 (68%), Positives = 31/35 (88%)
Frame = +2
Query: 41 FAVRDMRQTVVTGVIKSVEKKEPSGAKVTKAALKK 75
FAVRD+R TV GVI+SVEKK+P+GA+VT+AA +K
Sbjct: 2 FAVRDLRPTVAFGVIQSVEKKDPTGAQVTQAAAQK 106
>BP056358
Length = 598
Score = 44.7 bits (104), Expect = 3e-06
Identities = 19/23 (82%), Positives = 21/23 (90%)
Frame = -1
Query: 14 GDAGFVKMIPTKPMVVETFSEYP 36
GDAG VKM+P +PMVVETFSEYP
Sbjct: 598 GDAGMVKMLPPQPMVVETFSEYP 530
>TC11352
Length = 738
Score = 26.6 bits (57), Expect = 0.93
Identities = 12/30 (40%), Positives = 15/30 (50%)
Frame = +1
Query: 1 NKEIEKEPKFLKNGDAGFVKMIPTKPMVVE 30
N IEK P F D +K IP PM+ +
Sbjct: 214 NSVIEKNPAFFNGRDIDILKRIPGFPMLTK 303
>TC17682 similar to UP|AAS18240 (AAS18240) Enolase , partial (68%)
Length = 1146
Score = 25.8 bits (55), Expect = 1.6
Identities = 14/49 (28%), Positives = 24/49 (48%)
Frame = +1
Query: 11 LKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSVE 59
L +GD V + PM +E+ S + ++ + VTG IK+V+
Sbjct: 778 LIDGDVPLVSFLKRVPMAIESKSNTEICNCLMLKVSQMGSVTGCIKTVK 924
>TC10605 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor Tu,
chloroplast precursor (EF-Tu), partial (16%)
Length = 496
Score = 25.4 bits (54), Expect = 2.1
Identities = 19/54 (35%), Positives = 31/54 (57%)
Frame = +2
Query: 5 EKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSV 58
++E K + GD VKM+ ++V E + RFA+R+ +TV GVI+S+
Sbjct: 77 DEESKMVMPGDR--VKMVVE--LIVPVACEQGM--RFAIREGGKTVGAGVIQSI 220
>TC14629 homologue to UP|EFT2_SOYBN (P46280) Elongation factor Tu,
chloroplast precursor (EF-Tu), partial (24%)
Length = 675
Score = 25.4 bits (54), Expect = 2.1
Identities = 19/54 (35%), Positives = 31/54 (57%)
Frame = +2
Query: 5 EKEPKFLKNGDAGFVKMIPTKPMVVETFSEYPLLGRFAVRDMRQTVVTGVIKSV 58
++E K + GD VKM+ ++V E + RFA+R+ +TV GVI+S+
Sbjct: 194 DEESKMVMPGDR--VKMVVE--LIVPVACEQGM--RFAIREGGKTVGAGVIQSI 337
>BP069008
Length = 398
Score = 25.4 bits (54), Expect = 2.1
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = -1
Query: 57 SVEKKEPSGAKVTKAALKK 75
SVEKK+P AKVT+A ++
Sbjct: 398 SVEKKDPPDAKVTQACCQE 342
>TC18485 similar to UP|Q9LEU4 (Q9LEU4) CCR4-associated factor-like protein,
partial (48%)
Length = 586
Score = 23.5 bits (49), Expect = 7.8
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +2
Query: 45 DMRQTVVTGVIKSVEKKEPSGAKVTKAAL 73
D+R + TG+ VE+K P+ +T +L
Sbjct: 401 DLRSRMRTGICLLVERKLPASGSLTLGSL 487
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.133 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 756,375
Number of Sequences: 28460
Number of extensions: 5583
Number of successful extensions: 34
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 34
length of query: 75
length of database: 4,897,600
effective HSP length: 51
effective length of query: 24
effective length of database: 3,446,140
effective search space: 82707360
effective search space used: 82707360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0192.14


