

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0181.13
(507 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
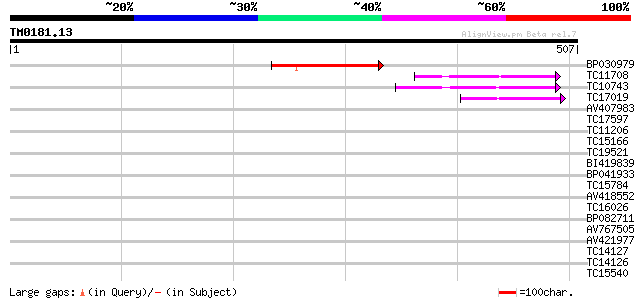
Score E
Sequences producing significant alignments: (bits) Value
BP030979 136 9e-33
TC11708 similar to UP|O82718 (O82718) Cyclin, partial (31%) 74 7e-14
TC10743 similar to UP|Q8H955 (Q8H955) B1 type cyclin, partial (21%) 68 3e-12
TC17019 similar to UP|Q40223 (Q40223) Cyclin, partial (21%) 62 2e-10
AV407983 35 0.029
TC17597 similar to UP|AZF1_YEAST (P41696) Asparagine-rich zinc f... 32 0.19
TC11206 similar to UP|Q84V88 (Q84V88) Cyclin D, partial (43%) 32 0.24
TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-bind... 32 0.32
TC19521 similar to UP|O22526 (O22526) Cation-chloride co-transpo... 32 0.32
BI419839 32 0.32
BP041933 32 0.32
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 31 0.42
AV418552 31 0.42
TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%) 31 0.42
BP082711 30 0.71
AV767505 30 0.71
AV421977 30 0.71
TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 30 0.93
TC14126 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1,... 30 0.93
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 30 0.93
>BP030979
Length = 510
Score = 136 bits (342), Expect = 9e-33
Identities = 63/102 (61%), Positives = 84/102 (81%), Gaps = 2/102 (1%)
Frame = +3
Query: 235 DPQLCASFACDIYKHLRASEA--KKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRL 292
DPQ+C + DIY++LR+ E KRP D+++KVQ+D+N +MR + +DWLVEV+EEY+L
Sbjct: 204 DPQMCLPYVSDIYEYLRSMEVDPSKRPLPDYVQKVQRDVNANMRGVXVDWLVEVSEEYKL 383
Query: 293 VPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEE 334
V DTLY V+YIDR+LS N++SRQKLQLLGVASM++ASKYEE
Sbjct: 384 VSDTLYFCVSYIDRFLSLNSLSRQKLQLLGVASMLVASKYEE 509
>TC11708 similar to UP|O82718 (O82718) Cyclin, partial (31%)
Length = 668
Score = 73.6 bits (179), Expect = 7e-14
Identities = 43/130 (33%), Positives = 69/130 (53%)
Frame = +2
Query: 363 VLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSL 422
+L L++ +T PT FL RF++A+ VP + ++ ++++EL +M Y L Y PS+
Sbjct: 5 ILGRLEWTLTVPTPFVFLTRFIKAS-----VPDEGVTNMAHFLSELGMMHYDTLMYCPSM 169
Query: 423 VAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEK 482
+AASA++ A+ L S W+ TL+ +T Y L C + L C N L + K
Sbjct: 170 IAASAVYAARCTLNKS-PAWNETLKLHTDYSEEQLMDCARLLVSFHCTVGNGKLKVVFRK 346
Query: 483 YSQHKYKYVA 492
YS + VA
Sbjct: 347 YSDPERGAVA 376
>TC10743 similar to UP|Q8H955 (Q8H955) B1 type cyclin, partial (21%)
Length = 554
Score = 68.2 bits (165), Expect = 3e-12
Identities = 48/149 (32%), Positives = 80/149 (53%), Gaps = 2/149 (1%)
Frame = +1
Query: 346 ITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYI 405
I+++ Y K ++ ME +L L++ +T PT FL R P Q+E+L ++
Sbjct: 1 ISNDAYTKHQIHSMERSILEKLQWYLTVPTPYVFLVRLSSVQ------PDKQMENLAFFL 162
Query: 406 AELSLMEY-SMLCYAPSLVAASAIFLAKFILFPSIKP-WSSTLQHYTLYQPSDLCVCVKE 463
AEL+L Y +++ Y+PS +AASAI+ A+ L + +P WS TL+ + Y + C K
Sbjct: 163 AELTLEHYQAIVSYSPSTIAASAIYAARCTL--NRRPFWSETLKRLSSYCTEQIIECAKL 336
Query: 464 LHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
L L ++ +S A+ K+S +VA
Sbjct: 337 LVSLQTSAADSKPNALYWKFSSEDKGFVA 423
>TC17019 similar to UP|Q40223 (Q40223) Cyclin, partial (21%)
Length = 652
Score = 62.4 bits (150), Expect = 2e-10
Identities = 38/95 (40%), Positives = 53/95 (55%), Gaps = 1/95 (1%)
Frame = +2
Query: 404 YIAELSLMEYSMLC-YAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVK 462
Y+AEL +M Y ++ Y+PS++AASA++ A+ L I W+ TL+HYT Y L C K
Sbjct: 8 YLAELGMMHYPVVSSYSPSVIAASAVYAARCTLH-RIPFWTETLKHYTGYSEEHLRECAK 184
Query: 463 ELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCP 497
L L +P S L AI +K+ VA Y P
Sbjct: 185 LLVNLHTAAPESKLRAIYKKFCSSDRCAVALLYVP 289
>AV407983
Length = 417
Score = 35.0 bits (79), Expect = 0.029
Identities = 30/110 (27%), Positives = 51/110 (46%), Gaps = 1/110 (0%)
Frame = +1
Query: 8 RSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNLTNHI 67
R+S S+S ++S+ H S N+++ + N + + A ++ SHS PPL L
Sbjct: 100 RASTSASPSTSIPSPHPSPQNSSSAS*NT-----SPISASSSPSHSPSSSPPL--LPPLS 258
Query: 68 ASRNSSSQSLVPCVSKFAKTKKEAPPKVAP-ALPNVKSAAAAVVFPKVAT 116
A +SS S P S + + + P +A ALP + FP + +
Sbjct: 259 APTSSSLSSSAP--SSTSSSSASSNPSLADLALPTPPTTTTTPSFPSITS 402
>TC17597 similar to UP|AZF1_YEAST (P41696) Asparagine-rich zinc finger
protein AZF1, partial (3%)
Length = 399
Score = 32.3 bits (72), Expect = 0.19
Identities = 15/34 (44%), Positives = 21/34 (61%)
Frame = +1
Query: 2 SSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNN 35
+ N ++ S SSS +S + R + NNNN NNNN
Sbjct: 181 NQNQNQNQSSSSSPMNSPSPRSNASNNNNYNNNN 282
Score = 29.6 bits (65), Expect = 1.2
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = +1
Query: 2 SSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
S+ N ++ SSS+ + ++ +NNNN NNNN
Sbjct: 178 SNQNQNQNQSSSSSPMNSPSPRSNASNNNNYNNNN 282
Score = 28.1 bits (61), Expect = 3.5
Identities = 13/36 (36%), Positives = 23/36 (63%)
Frame = +1
Query: 1 MSSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
++SN ++ + SSS++ + S+ +NNNN NNN
Sbjct: 172 INSNQNQNQNQSSSSSPMNSPSPRSNASNNNNYNNN 279
>TC11206 similar to UP|Q84V88 (Q84V88) Cyclin D, partial (43%)
Length = 816
Score = 32.0 bits (71), Expect = 0.24
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Frame = +3
Query: 19 LAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRP-PLSNLTNHIASRNSSSQSL 77
L +R +NNNNNNNN + +VK+ KK+P L ++T + +S S
Sbjct: 78 LPRRWLLENNNNNNNNLPHSSMPFSVKSNIHLMMMMKKKPLMLLSVTPLTMTHSSCSGRR 257
Query: 78 VPCVSKFAKTKKEAP 92
+ +S+ ++T E P
Sbjct: 258 MMSLSR*SRTNVERP 302
>TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-binding protein
6 {Arabidopsis thaliana;} , partial (26%)
Length = 668
Score = 31.6 bits (70), Expect = 0.32
Identities = 14/27 (51%), Positives = 17/27 (62%)
Frame = +3
Query: 9 SSFSSSTASSLAKRHASDNNNNNNNNN 35
++ SS S A + S NNNNNNNNN
Sbjct: 318 AAISSIIGGSHANNNGSHNNNNNNNNN 398
Score = 29.3 bits (64), Expect = 1.6
Identities = 13/28 (46%), Positives = 20/28 (71%), Gaps = 3/28 (10%)
Frame = +3
Query: 12 SSSTASSLAKRHASDN---NNNNNNNNN 36
+++ +S + HA++N NNNNNNNNN
Sbjct: 315 AAAISSIIGGSHANNNGSHNNNNNNNNN 398
>TC19521 similar to UP|O22526 (O22526) Cation-chloride co-transporter,
partial (5%)
Length = 556
Score = 31.6 bits (70), Expect = 0.32
Identities = 25/85 (29%), Positives = 42/85 (49%), Gaps = 6/85 (7%)
Frame = +2
Query: 1 MSSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPL 60
+S + H SS SSS++SSL+ H +N +++ V AA + K RP L
Sbjct: 38 ISDSKHSSSSSSSSSSSSLSPIHTMSSNPEMGADDD--VEAAGADGGFRSPIGRKYRPVL 211
Query: 61 SN------LTNHIASRNSSSQSLVP 79
+N +++ +SSS S++P
Sbjct: 212 ANDRAVLEMSSIDPGSSSSSSSVIP 286
>BI419839
Length = 496
Score = 31.6 bits (70), Expect = 0.32
Identities = 17/62 (27%), Positives = 31/62 (49%)
Frame = +1
Query: 46 APAAASHSAKKRPPLSNLTNHIASRNSSSQSLVPCVSKFAKTKKEAPPKVAPALPNVKSA 105
APA S PP + L++ + +R+ SS + V++ + ++PP +PA +K
Sbjct: 196 APATPPSSDPTEPPQTTLSSVVPTRSPSSSTSRGQVNRSTASSTQSPPAASPAWTILKEL 375
Query: 106 AA 107
A
Sbjct: 376 TA 381
>BP041933
Length = 472
Score = 31.6 bits (70), Expect = 0.32
Identities = 27/93 (29%), Positives = 38/93 (40%)
Frame = -1
Query: 8 RSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSNLTNHI 67
R SF + +S + DNNNNNNNN+ + P + S+ P N
Sbjct: 448 RISFGTQFHASHLVKSQKDNNNNNNNNSITETKSRTDPKPF*NTVSS*NP*PAPN----E 281
Query: 68 ASRNSSSQSLVPCVSKFAKTKKEAPPKVAPALP 100
AS ++S+S P +S AP K P
Sbjct: 280 ASNTNNSRSNSPQISALKLIHNAAPSKPLSKFP 182
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 31.2 bits (69), Expect = 0.42
Identities = 22/105 (20%), Positives = 45/105 (41%), Gaps = 2/105 (1%)
Frame = +1
Query: 48 AAASHSAKKRPPLSNLTNHIASRNSSSQSLVPCVSKFAKTKKEAPPKVAPALPNVKSAAA 107
+++S + PP +H S N+ + + + + T K PP AP P S++
Sbjct: 58 SSSSQHSHSSPPSHTRDSHSTSTNTRAHNSPKSSATPSPTNKSPPPPPAP--PRSASSST 231
Query: 108 AVVFPKVATATASFTE--RNEDDAAAAASGARVSVPVSTSMDVSP 150
P AT +S + + +A ++ + P ++S ++ P
Sbjct: 232 TASTPTAATPPSSSPQLPSTKPNATPTSTSPSPATPSTSSSELKP 366
>AV418552
Length = 386
Score = 31.2 bits (69), Expect = 0.42
Identities = 16/36 (44%), Positives = 25/36 (69%)
Frame = +3
Query: 2 SSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNY 37
SSNN+ RS + S+ + H+S+NNN +N+NNN+
Sbjct: 168 SSNNNSRSC--NLHCSNSSN*HSSNNNNLHNSNNNW 269
>TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%)
Length = 1183
Score = 31.2 bits (69), Expect = 0.42
Identities = 23/82 (28%), Positives = 39/82 (47%), Gaps = 7/82 (8%)
Frame = +2
Query: 100 PNVKSAAAAVVFPKVATATASFTERNEDDAAAAASGARVSVPVST-------SMDVSPCK 152
P +KS++ +T + S + + ++ + S + S+P S+ S V+P
Sbjct: 587 PFIKSSSPTPT--STSTFSPSSSSSSSSSSSPSPSSSNSSLPCSSPPNSFYSSPSVTPFF 760
Query: 153 SDGCSVSMDESMSSCDSFKSPA 174
SDGCS SM ++ SF PA
Sbjct: 761 SDGCSTSMTHVFANGLSFVQPA 826
>BP082711
Length = 379
Score = 30.4 bits (67), Expect = 0.71
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +3
Query: 22 RHASDNNNNNNNNNN 36
RH NNNNNNNNNN
Sbjct: 162 RHLFYNNNNNNNNNN 206
Score = 28.9 bits (63), Expect = 2.1
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = +3
Query: 16 ASSLAKRHASDNNNNNNNNNN 36
A L + ++NNNNNNNNNN
Sbjct: 159 ARHLFYNNNNNNNNNNNNNNN 221
Score = 28.5 bits (62), Expect = 2.7
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +3
Query: 25 SDNNNNNNNNNN 36
++NNNNNNNNNN
Sbjct: 189 NNNNNNNNNNNN 224
Score = 28.5 bits (62), Expect = 2.7
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +3
Query: 25 SDNNNNNNNNNN 36
++NNNNNNNNNN
Sbjct: 195 NNNNNNNNNNNN 230
Score = 28.5 bits (62), Expect = 2.7
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +3
Query: 25 SDNNNNNNNNNN 36
++NNNNNNNNNN
Sbjct: 198 NNNNNNNNNNNN 233
Score = 28.5 bits (62), Expect = 2.7
Identities = 10/12 (83%), Positives = 12/12 (99%)
Frame = +3
Query: 25 SDNNNNNNNNNN 36
++NNNNNNNNNN
Sbjct: 192 NNNNNNNNNNNN 227
>AV767505
Length = 536
Score = 30.4 bits (67), Expect = 0.71
Identities = 18/66 (27%), Positives = 31/66 (46%)
Frame = -1
Query: 234 MDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLV 293
M P + + AC I H+ S+++ M+KV +N + AIL + + A V
Sbjct: 377 MLPSILSLSACAIMSHMHNSDSQINTKASSMKKVTWQLNKELGAILAENVGFEANNGSSV 198
Query: 294 PDTLYL 299
T+Y+
Sbjct: 197 ESTIYI 180
>AV421977
Length = 535
Score = 30.4 bits (67), Expect = 0.71
Identities = 23/85 (27%), Positives = 29/85 (34%), Gaps = 12/85 (14%)
Frame = +3
Query: 27 NNNNNNNNN------NYVVGA-----AAVKAPAAASHSAKKRPP-LSNLTNHIASRNSSS 74
NNNNNNNNN N+ V + + P SH P + S +S
Sbjct: 36 NNNNNNNNNINKSEFNFCVSSTPTQQTTLTPPNVVSHPTNPNPR*FRRIITRFTSSSSGD 215
Query: 75 QSLVPCVSKFAKTKKEAPPKVAPAL 99
P F PP + P L
Sbjct: 216 PDTQPAFLHFHHPLHPPPPLLQPRL 290
>TC14127 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(21%)
Length = 556
Score = 30.0 bits (66), Expect = 0.93
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = +2
Query: 26 DNNNNNNNNNNYVVGAAAVKAPAAASHSAKKR 57
D NNNNN+NN V+ A A AA S+ K+
Sbjct: 236 DANNNNNSNNKDVIAADYASADNAAPPSSGKQ 331
Score = 27.3 bits (59), Expect = 6.0
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = +2
Query: 27 NNNNNNNNNNYVVGAAAVKAPAAASHSAKKR 57
NNNNN+NN + + A AA S K+R
Sbjct: 242 NNNNNSNNKDVIAADYASADNAAPPSSGKQR 334
>TC14126 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(44%)
Length = 1500
Score = 30.0 bits (66), Expect = 0.93
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = +2
Query: 26 DNNNNNNNNNNYVVGAAAVKAPAAASHSAKKR 57
D NNNNN+NN V+ A A AA S+ K+
Sbjct: 245 DANNNNNSNNKDVIAADYASADNAAPPSSGKQ 340
Score = 27.3 bits (59), Expect = 6.0
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = +2
Query: 27 NNNNNNNNNNYVVGAAAVKAPAAASHSAKKR 57
NNNNN+NN + + A AA S K+R
Sbjct: 251 NNNNNSNNKDVIAADYASADNAAPPSSGKQR 343
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 30.0 bits (66), Expect = 0.93
Identities = 16/35 (45%), Positives = 22/35 (62%)
Frame = +3
Query: 2 SSNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNN 36
SSNN+ S+F S +SS + S NN+ N +NNN
Sbjct: 63 SSNNNSSSNFHKSNSSSFPR--CSSNNSLNFSNNN 161
Score = 27.7 bits (60), Expect = 4.6
Identities = 25/84 (29%), Positives = 34/84 (39%)
Frame = +1
Query: 72 SSSQSLVPCVSKFAKTKKEAPPKVAPALPNVKSAAAAVVFPKVATATASFTERNEDDAAA 131
+S+ + + T AP A A +AAA FP A T S T AAA
Sbjct: 7 TSATAAAAATTTATTTTYPAPTTTAAATSTSPTAAA---FPDAAATTPS-TSATTTLAAA 174
Query: 132 AASGARVSVPVSTSMDVSPCKSDG 155
AA+ A ++ S S +DG
Sbjct: 175 AAATASAVAAITASTTSSTAATDG 246
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,456,895
Number of Sequences: 28460
Number of extensions: 115445
Number of successful extensions: 1447
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 968
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1216
length of query: 507
length of database: 4,897,600
effective HSP length: 94
effective length of query: 413
effective length of database: 2,222,360
effective search space: 917834680
effective search space used: 917834680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0181.13


