

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.5
(1481 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
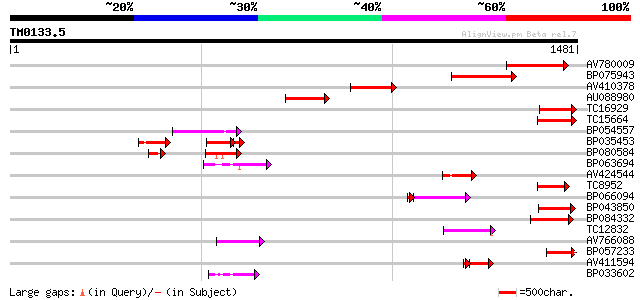
Score E
Sequences producing significant alignments: (bits) Value
AV780009 241 8e-64
BP075943 226 2e-59
AV410378 168 5e-42
AU088980 144 1e-34
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 130 2e-30
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 122 3e-28
BP054557 120 2e-27
BP035453 101 2e-27
BP080584 107 1e-23
BP063694 103 2e-22
AV424544 95 7e-20
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 94 2e-19
BP066094 86 1e-18
BP043850 83 3e-16
BP084332 83 4e-16
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 73 4e-13
AV766088 73 4e-13
BP057233 68 1e-11
AV411594 60 3e-10
BP033602 61 1e-09
>AV780009
Length = 529
Score = 241 bits (614), Expect = 8e-64
Identities = 113/163 (69%), Positives = 137/163 (83%)
Frame = -1
Query: 1297 AAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWR 1356
AA RVLRY+KG+P GLF+ A S L A+SDSD AGC DTR+S+TGY +FLG+SL+SWR
Sbjct: 526 AATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWR 347
Query: 1357 SKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNP 1416
+KKQTT SRSS EAEYRA+AATVCEVQWL YL Q L+++ PV +FCDNQSA+HIAHNP
Sbjct: 346 TKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPLPVPLFCDNQSALHIAHNP 167
Query: 1417 SYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKP 1459
++HERTKHIELDCH+VR K+ GL+HLLP+++ QLADIFTKP
Sbjct: 166 TFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKP 38
>BP075943
Length = 547
Score = 226 bits (577), Expect = 2e-59
Identities = 117/171 (68%), Positives = 138/171 (80%), Gaps = 1/171 (0%)
Frame = -3
Query: 1153 SATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGI 1212
+++GSFTALLLYVDDVLLAGNDM EI+LVK SLH +FRIKDLGEAK+FLGL IARS GI
Sbjct: 545 ASSGSFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGI 366
Query: 1213 ILNQRKYALELLSDSGLLGGKSATTPMDC-SQKLSASSGTPLSDISSYRRLIGRLLYLTT 1271
+LNQRKYAL+L+SDSG G + SQ L ++GTPL+DI SYRR++GRLLYL T
Sbjct: 365 VLNQRKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNT 186
Query: 1272 TRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTV 1322
TRPDI + VNQLSQFLSAPTD+HE L+Y+ GSPG GLFYPA+SST+
Sbjct: 185 TRPDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPASSSTL 33
>AV410378
Length = 358
Score = 168 bits (426), Expect = 5e-42
Identities = 84/120 (70%), Positives = 96/120 (80%)
Frame = +1
Query: 890 PLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVI 949
PL S N E+ V R+S R R+ P+YLQDYHCTLAA++TV SS+ G S+PLS+VI
Sbjct: 1 PLPSETLSNPEEGNVQRISNRVRRPPSYLQDYHCTLAASTTVSKVPSSA-GISYPLSKVI 177
Query: 950 SYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDK 1009
SYHKL PSY AFIMNITT EPTRYSEAVKHECWR AM+ EIEALERN+TW+LVDKPPDK
Sbjct: 178 SYHKLTPSYRAFIMNITTVSEPTRYSEAVKHECWRKAMDQEIEALERNHTWILVDKPPDK 357
>AU088980
Length = 360
Score = 144 bits (363), Expect = 1e-34
Identities = 66/115 (57%), Positives = 86/115 (74%)
Frame = +2
Query: 720 NSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQ 779
NSIVERKH+HILNV RAL+F ++LPK W A+ H+ +LIN LP+P+L QCP+Q L+
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 780 LPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDI 834
P + LKVFG+LC++STL + K D RA++CVFLGFK+GT GY+V DL T I
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDLPTTXI 346
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 130 bits (326), Expect = 2e-30
Identities = 62/98 (63%), Positives = 75/98 (76%)
Frame = +1
Query: 1383 QWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVH 1442
QWL YLLQDL+V P ++CDN SA HIA NP +HERTKHIE+DCHIVRE+I +GL+H
Sbjct: 1 QWLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIH 180
Query: 1443 LLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
LLP++SS LADI+TK LSP F I +KL L +I SP
Sbjct: 181 LLPISSSEPLADIYTKALSPQNFHQICAKLGLINICSP 294
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 122 bits (307), Expect = 3e-28
Identities = 56/100 (56%), Positives = 74/100 (74%)
Frame = +1
Query: 1380 CEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQG 1439
CE+ WL Y+LQDL+V SP ++CD+QSA HIA N +HERTKH+++DCH+VREK+
Sbjct: 1 CEL*WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAK 180
Query: 1440 LVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHS 1479
L HLLP++S Q ADI TKPL PF H+ SKL + +I+S
Sbjct: 181 LFHLLPISSVDQTADILTKPLESGPFSHLVSKLGVLNIYS 300
>BP054557
Length = 538
Score = 120 bits (301), Expect = 2e-27
Identities = 70/182 (38%), Positives = 100/182 (54%), Gaps = 3/182 (1%)
Frame = -2
Query: 426 ILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPN 485
+LDTGAT H+C+ L F + + V P+ +S+PNG E A + S+ S F L VL++P+
Sbjct: 537 LLDTGATHHVCHNLDVFQALRQVHPVLLSMPNGQKEIAHMADSVYFSEKFFLAXVLYVPS 358
Query: 486 FEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPF--P 543
F FNLISV LVKSL RL D+ CL+Q+ K IG A LY + + ++ P
Sbjct: 357 FNFNLISVSALVKSLNCRLVIFDECCLLQERPTLKTIGAAEERGGLYAVIDPAVKAKHPP 178
Query: 544 SCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQ-DHVCEVCP 602
+ ++ T S C+F NLWH RLGH S K ++ FP++ Q C+ C
Sbjct: 177 AEGNIKTQVSHFNFCQFD---CNLWHLRLGHTSSKKLAILQKNFPFITCKQITSPCDTCH 7
Query: 603 IA 604
+A
Sbjct: 6 MA 1
>BP035453
Length = 550
Score = 101 bits (251), Expect(2) = 2e-27
Identities = 49/80 (61%), Positives = 58/80 (72%), Gaps = 4/80 (5%)
Frame = -1
Query: 515 DSNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTC----KSQNLVCEFSPSVQNLWHY 570
DS+A + IGT +AV+ LYILN S++P CNSV + S N+ F PSV NLWHY
Sbjct: 307 DSSAWRTIGTVKAVKGLYILNKSSVTPLSHCNSVHSTVNIPSSVNVSSHFDPSVHNLWHY 128
Query: 571 RLGHPSFVKGQSIKDLFPYV 590
RLGHPSFVKGQSI+DLFPYV
Sbjct: 127 RLGHPSFVKGQSIRDLFPYV 68
Score = 90.9 bits (224), Expect = 1e-18
Identities = 52/84 (61%), Positives = 61/84 (71%), Gaps = 2/84 (2%)
Frame = -3
Query: 337 RSANLVQTTKEDQESTSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQK-NSTSQHTASI 395
+SAN V EDQES QG + Q+ RFGFTADQY HLL LLP SE K +S+SQHTAS+
Sbjct: 548 KSANCVHIADEDQES--QGGSSGQDT-RFGFTADQYQHLLALLPTSESKSSSSSQHTASV 378
Query: 396 NSCVQMLPGKNGN-SISQISTKNG 418
NSC Q+LP K+GN S S + TKNG
Sbjct: 377 NSCAQVLPTKHGNGSTSTVPTKNG 306
Score = 39.7 bits (91), Expect(2) = 2e-27
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = -3
Query: 584 KDLFPYVQYTQDHVCEVCPIAKQKRLKFP 612
K F T+DH+C+VCP+ KQK+L FP
Sbjct: 89 KGSFSLCTCTKDHICDVCPLLKQKKLSFP 3
>BP080584
Length = 507
Score = 107 bits (268), Expect = 1e-23
Identities = 55/109 (50%), Positives = 70/109 (63%), Gaps = 15/109 (13%)
Frame = -3
Query: 512 LIQDSNACKMIGTARAVRSLYILNNDSL---------SPFPSCNSVSTCKSQ------NL 556
LI D +A K IGTARAV+ LY+L + S P+CNSVS S NL
Sbjct: 328 LIMDYSAYKRIGTARAVKGLYLLMKGNPKSTTSVTCNSVSPACNSVSVSGSDSPVVDTNL 149
Query: 557 VCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQDHVCEVCPIAK 605
C F PS+ NLWH+RLGHPSFVK QSI+D+FPY+ ++ VC++CP+AK
Sbjct: 148 ACNFIPSMHNLWHFRLGHPSFVKEQSIRDIFPYIVCNKEQVCDICPLAK 2
Score = 56.6 bits (135), Expect = 3e-08
Identities = 27/45 (60%), Positives = 32/45 (71%)
Frame = -2
Query: 363 ARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNG 407
A FG TADQYHHL+ LLP +QK+S T S+NSCVQ +P NG
Sbjct: 443 ASFGLTADQYHHLMALLPAQDQKDS---RTPSVNSCVQEIPTHNG 318
>BP063694
Length = 511
Score = 103 bits (258), Expect = 2e-22
Identities = 65/187 (34%), Positives = 92/187 (48%), Gaps = 9/187 (4%)
Frame = -2
Query: 506 FSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQ 565
F DD +IQ+ ++G R + LY+LN DS Q L+ SP +
Sbjct: 507 FLDDSFVIQNRKTGAVLGRXRCDQGLYVLNQDS---------------QALLTTSSPLPR 373
Query: 566 ---NLWHYRLGHPSFVKGQSIKDLFPYVQYTQDHV------CEVCPIAKQKRLKFPLSNS 616
LWH RLGH +F IK L + + C C +AK KRL F +N
Sbjct: 372 ASFELWHSRLGHVNF---DIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLVFHDNNK 202
Query: 617 TSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAF 676
+ + +IH D+ GP V S++GFSYF+ VDD+SR+TW Y LK K + ++ F F
Sbjct: 201 RASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVF 22
Query: 677 VTNQFGV 683
+ N+F V
Sbjct: 21 MENRFSV 1
>AV424544
Length = 276
Score = 95.1 bits (235), Expect = 7e-20
Identities = 49/90 (54%), Positives = 65/90 (71%)
Frame = +3
Query: 1130 LCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKF 1189
L L LG+ QS DH+L+ K SFT +L+YVDD++LAGND++EI+ VK+ L +F
Sbjct: 9 LSSYLHILGYIQSAHDHSLFTK-FRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 1190 RIKDLGEAKFFLGLAIARSQKGIILNQRKY 1219
RIKDLG K+FLGL +ARS G+ L+QRKY
Sbjct: 186 RIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 93.6 bits (231), Expect = 2e-19
Identities = 42/82 (51%), Positives = 61/82 (74%)
Frame = +3
Query: 1380 CEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQG 1439
CE WL Y L DL+++ V ++CDN+SA+H+A N +H+RT++IE+DCHIV K+ G
Sbjct: 18 CEALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFG 197
Query: 1440 LVHLLPVTSSLQLADIFTKPLS 1461
++HLL V SS Q+AD+FTK +S
Sbjct: 198 ILHLLHVPSSDQVADVFTKTIS 263
>BP066094
Length = 532
Score = 85.9 bits (211), Expect(2) = 1e-18
Identities = 48/149 (32%), Positives = 79/149 (52%), Gaps = 1/149 (0%)
Frame = +2
Query: 1056 AKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-PNQVCLLQK 1114
+K +R+L+S N LHQ++V +AFL+ + EE+Y+ P G +K + + L+K
Sbjct: 86 SKTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKK 265
Query: 1115 SLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGND 1174
SLYGLKQA R W+ L L + D TL+ K+ + +YVDD++ +
Sbjct: 266 SLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFC-KTYKDDILIVQIYVDDIIFGSAN 442
Query: 1175 MDEIKLVKHSLHQKFRIKDLGEAKFFLGL 1203
K + +F ++ +GE K+FLG+
Sbjct: 443 PSLCKEFSEMMQAEFEMRMMGELKYFLGI 529
Score = 25.8 bits (55), Expect(2) = 1e-18
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +3
Query: 1040 YTQIEGIDFMDTFSPVAK 1057
Y+Q GID+ +TF+PVA+
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
>BP043850
Length = 515
Score = 83.2 bits (204), Expect = 3e-16
Identities = 43/96 (44%), Positives = 60/96 (61%)
Frame = -1
Query: 1381 EVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGL 1440
E+ WL LL +L Q+ P + DN SA+ IA NP YHE T+HIE+DCH VRE + +
Sbjct: 512 EIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRRV 333
Query: 1441 VHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYD 1476
+ L V++S+Q+ADI TK L+ + SKL L +
Sbjct: 332 ITLPHVSTSVQIADILTKSLTRQRHNFLVSKLMLVE 225
>BP084332
Length = 368
Score = 82.8 bits (203), Expect = 4e-16
Identities = 41/113 (36%), Positives = 70/113 (61%)
Frame = +2
Query: 1360 QTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYH 1419
Q+T + S+ EAEY + A ++ W+ + L+D Q+ +++ + ++CDN +A+ ++ NP H
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILH 208
Query: 1420 ERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
R KHIE+ H +R+ + +G++ L V + Q ADIFTKPL+ F I L
Sbjct: 209 SRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 367
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 72.8 bits (177), Expect = 4e-13
Identities = 48/144 (33%), Positives = 75/144 (51%), Gaps = 9/144 (6%)
Frame = +2
Query: 1134 LQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKD 1193
+ LG+ + +DH Y K+ F LLLYVDD+L+ G + D ++ +K L ++F +KD
Sbjct: 14 IMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDMKD 193
Query: 1194 LGEAKFFLGLAIARSQKG--IILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGT 1251
LG A LG+ I R +K I L+Q+ Y ++L + +TP+ + KLS SS
Sbjct: 194 LGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLS-SSMI 370
Query: 1252 PLSDIS-------SYRRLIGRLLY 1268
P S+ Y +G L+Y
Sbjct: 371 PSSEAERMEMSRVPYASAVGSLMY 442
>AV766088
Length = 501
Score = 72.8 bits (177), Expect = 4e-13
Identities = 41/128 (32%), Positives = 61/128 (47%), Gaps = 2/128 (1%)
Frame = -3
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKD--LFPYVQYTQDHVC 598
P CNS L F+ Q+LWH RLGHPS + ++ Y VC
Sbjct: 385 PLMRCNSPGDLYPVTLSFPFAGIAQSLWHSRLGHPSSSALRYLRSNKFISYELLNYSPVC 206
Query: 599 EVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVY 658
E C K RL F SN+ + F ++H D+W VLS G +++ +DD++ + W +
Sbjct: 205 ESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRFYVLFLDDFTDFLWTF 29
Query: 659 LLKSKDEV 666
L +K +V
Sbjct: 28 PLSNKSQV 5
>BP057233
Length = 473
Score = 67.8 bits (164), Expect = 1e-11
Identities = 35/79 (44%), Positives = 50/79 (62%)
Frame = -2
Query: 1402 MFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLS 1461
++CDN SA +A NP H R+KHIE+D H +R+++ Q V + V ++ Q+AD TKPLS
Sbjct: 445 LWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIADCLTKPLS 266
Query: 1462 PAPFRHIFSKLRLYDIHSP 1480
F + KL + IHSP
Sbjct: 265 HTRFSQLRDKLGV--IHSP 215
>AV411594
Length = 244
Score = 60.5 bits (145), Expect(2) = 3e-10
Identities = 32/65 (49%), Positives = 46/65 (70%)
Frame = +2
Query: 1200 FLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSY 1259
FL IA+S+KGI+L++RKYAL+LL D+ +G K P+D S KL+++ G LSD++ Y
Sbjct: 50 FLRT*IAKSKKGILLSRRKYALQLLDDTRNMGCKPPPFPLDPSIKLNSTDGELLSDVTLY 229
Query: 1260 RRLIG 1264
R IG
Sbjct: 230 IRFIG 244
Score = 22.7 bits (47), Expect(2) = 3e-10
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +3
Query: 1185 LHQKFRIKDLGEAKFFLGL 1203
L + F++K L + K+FLGL
Sbjct: 6 LAKSFKLKVLDDMKYFLGL 62
>BP033602
Length = 533
Score = 61.2 bits (147), Expect = 1e-09
Identities = 41/136 (30%), Positives = 64/136 (46%), Gaps = 3/136 (2%)
Frame = +3
Query: 520 KMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVK 579
KMIG+A+ LY + F + + C + C LWH RLGHP+F
Sbjct: 156 KMIGSAKMKDGLYYFEDT----FRNKAAQGLC---GMSCTSVRDQIMLWHNRLGHPNF-- 308
Query: 580 GQSIKDLFPYVQYT---QDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVL 636
Q ++ LFP + CE C +AK +R + + F +IH D+WGP +
Sbjct: 309 -QYLRHLFPDLFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFYLIHSDVWGPSKIT 485
Query: 637 SLNGFSYFLTIVDDYS 652
+ G +F+T +DD++
Sbjct: 486 TQFGKRWFVTFIDDHT 533
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,196,996
Number of Sequences: 28460
Number of extensions: 431028
Number of successful extensions: 3161
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 3062
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3130
length of query: 1481
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1379
effective length of database: 1,994,680
effective search space: 2750663720
effective search space used: 2750663720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0133.5


