

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.1
(267 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
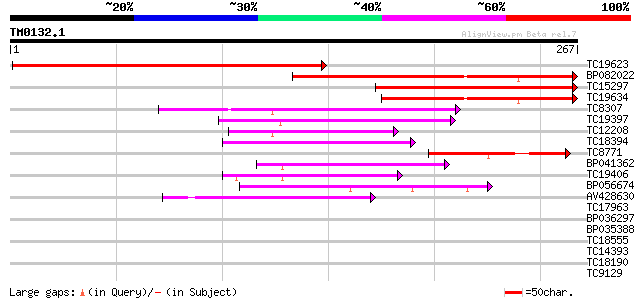
Score E
Sequences producing significant alignments: (bits) Value
TC19623 similar to PIR|T52371|T52371 homeobox protein HAT9 [impo... 292 4e-80
BP082022 233 3e-62
TC15297 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zippe... 201 1e-52
TC19634 similar to PIR|T52371|T52371 homeobox protein HAT9 [impo... 152 6e-38
TC8307 weakly similar to UP|Q9SP47 (Q9SP47) Homeodomain-leucine ... 72 1e-13
TC19397 similar to PIR|S47137|S47137 homeotic protein Athb-7 - A... 68 1e-12
TC12208 similar to UP|Q8LC03 (Q8LC03) Homeobox gene 13 protein, ... 65 9e-12
TC18394 similar to UP|O23208 (O23208) Homeodomain protein, parti... 63 5e-11
TC8771 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper p... 59 1e-09
BP041362 47 3e-06
TC19406 similar to UP|Q9FJS2 (Q9FJS2) Homeobox protein, partial ... 47 3e-06
BP056674 45 1e-05
AV428630 41 2e-04
TC17963 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zippe... 33 0.039
BP036297 32 0.11
BP035388 32 0.11
TC18555 similar to GB|AAP68307.1|31711902|BT008868 At2g33570 {Ar... 31 0.25
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 31 0.25
TC18190 weakly similar to UP|O24569 (O24569) HOX1B protein, part... 30 0.33
TC9129 homologue to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic pro... 30 0.33
>TC19623 similar to PIR|T52371|T52371 homeobox protein HAT9 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(21%)
Length = 554
Score = 292 bits (748), Expect = 4e-80
Identities = 148/148 (100%), Positives = 148/148 (100%)
Frame = +1
Query: 2 GLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNN 61
GLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNN
Sbjct: 4 GLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNN 183
Query: 62 KVYCEDPLELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGT 121
KVYCEDPLELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGT
Sbjct: 184 KVYCEDPLELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGT 363
Query: 122 NARKKLRLTKEQSALLEESFKQHSTLNP 149
NARKKLRLTKEQSALLEESFKQHSTLNP
Sbjct: 364 NARKKLRLTKEQSALLEESFKQHSTLNP 447
>BP082022
Length = 455
Score = 233 bits (593), Expect = 3e-62
Identities = 116/137 (84%), Positives = 124/137 (89%), Gaps = 3/137 (2%)
Frame = -3
Query: 134 SALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCC 193
SA+LEESFKQHSTLNPKQKQALARQLNLR RQVEVWFQNRRARTKLKQTEVDC+FLKKCC
Sbjct: 453 SAMLEESFKQHSTLNPKQKQALARQLNLRPRQVEVWFQNRRARTKLKQTEVDCDFLKKCC 274
Query: 194 ETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERLG---DGGSNIKSPFTI 250
ETLTDEN RL+KELQELKALK QPLYMPM AATLTMCPSCERLG GG++ K PF++
Sbjct: 273 ETLTDENMRLQKELQELKALK-TQPLYMPMPAATLTMCPSCERLGGVSGGGASNKIPFSM 97
Query: 251 TPKPHFFNPFTHPSAAC 267
KPHFFNPFT+PSAAC
Sbjct: 96 ASKPHFFNPFTNPSAAC 46
>TC15297 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (32%)
Length = 585
Score = 201 bits (511), Expect = 1e-52
Identities = 95/95 (100%), Positives = 95/95 (100%)
Frame = +2
Query: 173 RRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCP 232
RRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCP
Sbjct: 2 RRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCP 181
Query: 233 SCERLGDGGSNIKSPFTITPKPHFFNPFTHPSAAC 267
SCERLGDGGSNIKSPFTITPKPHFFNPFTHPSAAC
Sbjct: 182 SCERLGDGGSNIKSPFTITPKPHFFNPFTHPSAAC 286
>TC19634 similar to PIR|T52371|T52371 homeobox protein HAT9 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(32%)
Length = 544
Score = 152 bits (384), Expect = 6e-38
Identities = 75/95 (78%), Positives = 82/95 (85%), Gaps = 3/95 (3%)
Frame = +1
Query: 176 RTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCE 235
RTKLK TEVDC+FLKKCCETLTDEN RL+KELQELKALK QPLYMPM AATLTMCPSCE
Sbjct: 214 RTKLKPTEVDCDFLKKCCETLTDENMRLQKELQELKALK-TQPLYMPMPAATLTMCPSCE 390
Query: 236 RLG---DGGSNIKSPFTITPKPHFFNPFTHPSAAC 267
RLG GG++ K PF++ KPHFFNPFT+PSAAC
Sbjct: 391 RLGGVSGGGASNKIPFSMASKPHFFNPFTNPSAAC 495
>TC8307 weakly similar to UP|Q9SP47 (Q9SP47) Homeodomain-leucine zipper
protein 57, partial (51%)
Length = 1629
Score = 71.6 bits (174), Expect = 1e-13
Identities = 51/150 (34%), Positives = 73/150 (48%), Gaps = 8/150 (5%)
Frame = +1
Query: 71 LSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTN-------- 122
L + P S + S S + + R V + ++ + R D D++G
Sbjct: 457 LLQNQPPDSSLFLSTSASFLGSRSMVSFADNKL-GQTRSFFSAFDLDENGDEVMDEYFHQ 633
Query: 123 ARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQT 182
+ KK RL+ +Q LE+SF + L P +K +A+ L L+ RQV +WFQNRRAR K KQ
Sbjct: 634 SEKKRRLSVDQVQFLEKSF*VDNKLEPDRKTKIAKDLGLQPRQVAIWFQNRRARWKTKQL 813
Query: 183 EVDCEFLKKCCETLTDENRRLKKELQELKA 212
E D + L E+L L KE LKA
Sbjct: 814 EKDYDSLHSSFESLKSNYDNLLKEKDMLKA 903
>TC19397 similar to PIR|S47137|S47137 homeotic protein Athb-7 - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (35%)
Length = 723
Score = 68.2 bits (165), Expect = 1e-12
Identities = 44/114 (38%), Positives = 57/114 (49%), Gaps = 2/114 (1%)
Frame = +2
Query: 99 SCEEIEAEERLSSRVSDEDDDGTNARKK--LRLTKEQSALLEESFKQHSTLNPKQKQALA 156
S EE E ++ S + R K R + EQ LE F+ S L P++K LA
Sbjct: 173 SAEETADAETYTTSSSTQPVSMRKKRNKNARRFSDEQIKSLESMFESESRLEPRKKLQLA 352
Query: 157 RQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQEL 210
R+L L+ RQV +WFQN+RAR K KQ E + L L + LKKE Q L
Sbjct: 353 RELGLQPRQVAIWFQNKRARWKSKQLEREYSMLSSNYNNLASKYEVLKKEKQTL 514
>TC12208 similar to UP|Q8LC03 (Q8LC03) Homeobox gene 13 protein, partial
(24%)
Length = 660
Score = 65.5 bits (158), Expect = 9e-12
Identities = 36/81 (44%), Positives = 48/81 (58%), Gaps = 1/81 (1%)
Frame = +1
Query: 104 EAEERLSSRVSDEDDDGTN-ARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLR 162
+ +E L D DDG+ KK RL+ EQ LE SF+ + L P++K LA+ L L+
Sbjct: 418 KCDEVLHGGDDDLSDDGSQLGEKKKRLSLEQVKALERSFELGNKLEPERKMQLAKALGLQ 597
Query: 163 ARQVEVWFQNRRARTKLKQTE 183
RQ+ +WFQNRRAR K K E
Sbjct: 598 PRQIAIWFQNRRARWKTKHLE 660
>TC18394 similar to UP|O23208 (O23208) Homeodomain protein, partial (46%)
Length = 846
Score = 63.2 bits (152), Expect = 5e-11
Identities = 36/91 (39%), Positives = 52/91 (56%)
Frame = +2
Query: 101 EEIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLN 160
E+ + R + + ++ A KK +L++EQ LLE++F L ++K LA +L
Sbjct: 101 EKKQPRRRRNKKNRGGENGSAEANKKRKLSEEQVNLLEQNFGHEHKLESERKDKLAAELC 280
Query: 161 LRARQVEVWFQNRRARTKLKQTEVDCEFLKK 191
L RQV VWFQNRRAR K K+ E + LKK
Sbjct: 281 LDPRQVAVWFQNRRARWKNKKLEEEYSNLKK 373
>TC8771 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper protein,
partial (20%)
Length = 953
Score = 58.5 bits (140), Expect = 1e-09
Identities = 31/68 (45%), Positives = 42/68 (61%), Gaps = 1/68 (1%)
Frame = +1
Query: 198 DENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNIKSPFTITPKPHF 256
+ENRRL+KE+QEL+ALKL+ YM M+ TLTMCPSCER+ P + P
Sbjct: 1 EENRRLQKEVQELRALKLSPQFYMQMTPPTTLTMCPSCERVA------VPPSAVDPSTRH 162
Query: 257 FNPFTHPS 264
+P + P+
Sbjct: 163 HHPMSAPN 186
>BP041362
Length = 560
Score = 47.4 bits (111), Expect = 3e-06
Identities = 30/93 (32%), Positives = 49/93 (52%), Gaps = 2/93 (2%)
Frame = +3
Query: 117 DDDGTNARKKL--RLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
+D A+KK R T+ Q +E FK+ + KQ++ L+R+L L QV+ WFQN+R
Sbjct: 156 EDQEPRAKKKRYHRHTQHQIQEMEAFFKECPHPDDKQRKELSRELGLEPLQVKFWFQNKR 335
Query: 175 ARTKLKQTEVDCEFLKKCCETLTDENRRLKKEL 207
+ K + + L+ E L +N R ++ L
Sbjct: 336 TQMKTQHERHENTSLRTENEKLRADNMRYREAL 434
>TC19406 similar to UP|Q9FJS2 (Q9FJS2) Homeobox protein, partial (10%)
Length = 546
Score = 47.4 bits (111), Expect = 3e-06
Identities = 30/90 (33%), Positives = 47/90 (51%), Gaps = 5/90 (5%)
Frame = +1
Query: 101 EEIEA---EERLSSRVSDEDDDGTNARKKL--RLTKEQSALLEESFKQHSTLNPKQKQAL 155
EE+E+ E++ + +E + +KK R T Q +E FK+ + KQ+ L
Sbjct: 265 EEMESGSESEQVEDKSGNEQEITEQPKKKRYHRHTARQIQEMEALFKECPHPDDKQRMKL 444
Query: 156 ARQLNLRARQVEVWFQNRRARTKLKQTEVD 185
+ L L+ RQV+ WFQNRR + K +Q D
Sbjct: 445 SHDLGLKPRQVKFWFQNRRTQMKAQQDRSD 534
>BP056674
Length = 566
Score = 45.1 bits (105), Expect = 1e-05
Identities = 36/129 (27%), Positives = 61/129 (46%), Gaps = 10/129 (7%)
Frame = +1
Query: 109 LSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQL----NLRAR 164
+ R S D ++ K +R T Q LE + + + ++Q L R+ N+ +
Sbjct: 70 VQDRESSIDRHLDSSGKYVRYTAGQVEALERVYTECPKPSSLRRQQLIRECPVLANVEPK 249
Query: 165 QVEVWFQNRRARTKLKQTEVDCEF----LKKCCETLTDENRRLKKELQELKALK--LAQP 218
Q++VWFQNRR R K ++ + L + L +EN RL+K++ +L + Q
Sbjct: 250 QIKVWFQNRRCREKQRKEASRLQAVNRKLNAMNKLLMEENDRLQKQVSQLVCENGFMRQQ 429
Query: 219 LYMPMSAAT 227
L P +A T
Sbjct: 430 LQAPSAAGT 456
>AV428630
Length = 342
Score = 41.2 bits (95), Expect = 2e-04
Identities = 25/100 (25%), Positives = 48/100 (48%)
Frame = +1
Query: 73 RQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLTKE 132
+ T+ SD+ ++ R+ + D + + + + D+D ++ R T+
Sbjct: 52 QNTTSESDIAAA---VRIRGEDEFDSATKSGSENQEGGASGGDQDPRPNKKKRYHRHTQH 222
Query: 133 QSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQN 172
Q +E FK+ + KQ++ L+R+L L QV+ WFQN
Sbjct: 223 QIQEMESFFKECPHPDDKQRKELSRELGLEPLQVKFWFQN 342
>TC17963 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zipper protein
HAT5 (HD-ZIP protein 5) (HD-ZIP protein ATHB-1), partial
(24%)
Length = 738
Score = 33.5 bits (75), Expect = 0.039
Identities = 16/31 (51%), Positives = 22/31 (70%)
Frame = +1
Query: 125 KKLRLTKEQSALLEESFKQHSTLNPKQKQAL 155
KK RLT EQ LLE+SF++ + L P++K L
Sbjct: 646 KKRRLTSEQVNLLEKSFEEENKLEPERKTQL 738
>BP036297
Length = 569
Score = 32.0 bits (71), Expect = 0.11
Identities = 16/60 (26%), Positives = 32/60 (52%)
Frame = +3
Query: 119 DGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTK 178
+G N K+ + Q LLE+++ + + L+ +L L RQ+++WF +RR + +
Sbjct: 159 EGENKVKRKMKSASQLELLEKTYAVEAYPGEVLRAELSEKLGLSDRQLQMWFCHRRLKDR 338
>BP035388
Length = 557
Score = 32.0 bits (71), Expect = 0.11
Identities = 27/109 (24%), Positives = 48/109 (43%), Gaps = 10/109 (9%)
Frame = +3
Query: 132 EQSALLEESFKQHSTLNPKQKQALARQL-----NLRARQVEVWFQNRRARTKLKQTEVDC 186
EQ +LE F PK + R+L + V WFQNRR+R++ +Q ++
Sbjct: 123 EQILILESIFNSGMVNPPKDETVRIRKLLEKFGTVGDANVFYWFQNRRSRSRRRQRQMQA 302
Query: 187 EFLKKCCETLTDENRRLKKELQE-----LKALKLAQPLYMPMSAATLTM 230
++ + +N ++++L E A+ Q P AA ++M
Sbjct: 303 SLVE---QRNNLQNNNMQQQLLEGAPAAASAIPCDQAQGHPNLAAAISM 440
>TC18555 similar to GB|AAP68307.1|31711902|BT008868 At2g33570 {Arabidopsis
thaliana;}, partial (19%)
Length = 759
Score = 30.8 bits (68), Expect = 0.25
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +1
Query: 25 TSKETTTTTTTSSPKPTVMKPYSSKEP 51
T K TTTTT +SP P V+ P S P
Sbjct: 256 TQKTATTTTTATSPPPPVLPPPSPPPP 336
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 30.8 bits (68), Expect = 0.25
Identities = 22/78 (28%), Positives = 36/78 (45%)
Frame = +1
Query: 7 ASNPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNNKVYCE 66
+S+ LH++L LSL+ + T T T + KPT +KP S+ + T + +
Sbjct: 1 SSSSHLHISLYLSLSLKTKQEWQQTETETETEKPT-LKPVSNPLSTSTTWKNSSESSPAS 177
Query: 67 DPLELSRQTSPHSDVVSS 84
P +R S S S+
Sbjct: 178TPTATARSPSTSSTTSSA 231
>TC18190 weakly similar to UP|O24569 (O24569) HOX1B protein, partial (10%)
Length = 981
Score = 30.4 bits (67), Expect = 0.33
Identities = 15/38 (39%), Positives = 22/38 (57%)
Frame = +3
Query: 137 LEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
L +SFK + + K++LA +L L QV+ WF N R
Sbjct: 480 LHKSFKDNQYPDRATKESLAEELGLTFFQVDKWFGNAR 593
>TC9129 homologue to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic protein 13
(Fragment), partial (22%)
Length = 604
Score = 30.4 bits (67), Expect = 0.33
Identities = 27/89 (30%), Positives = 33/89 (36%)
Frame = +2
Query: 144 HSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRL 203
H + K LARQ L QV WF N R R E + +K E + R
Sbjct: 218 HPYPSDADKHLLARQTGLSRNQVSNWFINARVRLWKPMVEEMYQQERKEVECAEERERN- 394
Query: 204 KKELQELKALKLAQPLYMPMSAATLTMCP 232
Q + AQ P S+AT T P
Sbjct: 395 ----QSNSGQQQAQTPTTPPSSATSTAPP 469
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,366,789
Number of Sequences: 28460
Number of extensions: 54825
Number of successful extensions: 720
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 598
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 663
length of query: 267
length of database: 4,897,600
effective HSP length: 89
effective length of query: 178
effective length of database: 2,364,660
effective search space: 420909480
effective search space used: 420909480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0132.1


