

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
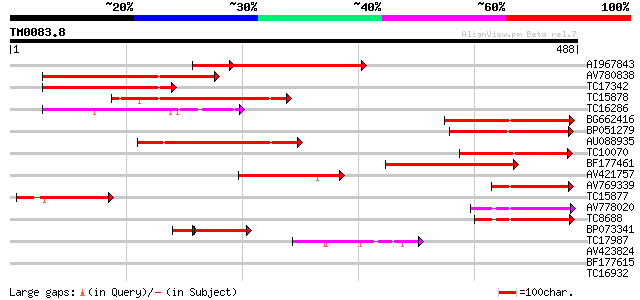
Query= TM0083.8
(488 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AI967843 246 1e-80
AV780838 184 2e-47
TC17342 similar to PIR|JC7226|JC7226 endo-1,3(4)-beta-glucanase ... 157 5e-39
TC15878 homologue to UP|Q8L6S6 (Q8L6S6) Endo-beta-1,4-glucanase ... 153 5e-38
TC16286 similar to UP|Q8GTP6 (Q8GTP6) Endo-1,4-beta-D-glucanase ... 149 1e-36
BG662416 145 1e-35
BP051279 145 2e-35
AU088935 142 9e-35
TC10070 similar to UP|O81403 (O81403) Endo-1,4-beta-glucanase pr... 136 9e-33
BF177461 110 5e-25
AV421757 105 2e-23
AV769339 90 7e-19
TC15877 similar to UP|O65186 (O65186) Cellulase, partial (12%) 84 4e-17
AV778020 76 1e-14
TC8688 homologue to UP|AAS45400 (AAS45400) Endo-1,4-beta-glucana... 75 2e-14
BP073341 50 1e-09
TC17987 similar to UP|O04890 (O04890) Endo-1,4-beta-glucanase ,... 54 4e-08
AV423824 39 0.001
BF177615 30 1.2
TC16932 similar to UP|Q94A94 (Q94A94) AT5g11880/F14F18_50, parti... 29 1.5
>AI967843
Length = 485
Score = 246 bits (628), Expect(2) = 1e-80
Identities = 117/118 (99%), Positives = 118/118 (99%)
Frame = +3
Query: 190 FKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLF 249
F+KVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLF
Sbjct: 111 FQKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLF 290
Query: 250 RATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAEN 307
RATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAEN
Sbjct: 291 RATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAEN 464
Score = 71.2 bits (173), Expect(2) = 1e-80
Identities = 34/36 (94%), Positives = 36/36 (99%)
Frame = +1
Query: 158 DTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKV 193
DTARTVYWVSPNNPGSDVAAETAAALAAASLVFK++
Sbjct: 16 DTARTVYWVSPNNPGSDVAAETAAALAAASLVFKRL 123
>AV780838
Length = 578
Score = 184 bits (468), Expect = 2e-47
Identities = 95/153 (62%), Positives = 111/153 (72%), Gaps = 1/153 (0%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
+Y +AL+K++LF +GQRSG LP DQ++ R +SGL DG DL+GGYYDAGDNVKF F
Sbjct: 122 DYHDALSKAILFCEGQRSGFLPQDQRMTGRGNSGLSDGWTYKTDLAGGYYDAGDNVKFGF 301
Query: 89 PMAFTTTMLSWSAIEYGKRM-GPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVD 147
PMAFTTTMLSWS +EYG M +L+ A A+RW TDYLLK S P R+ V VGDP+ D
Sbjct: 302 PMAFTTTMLSWSVLEYGDVMPADELRNALVAVRWATDYLLK-TVSQPNRILVQVGDPSSD 478
Query: 148 HKCWERPEDMDTARTVYWVSPNNPGSDVAAETA 180
H ERPEDMDTARTVY V N SDVA ETA
Sbjct: 479 HDGRERPEDMDTARTVYAVDAPNAASDVAGETA 577
>TC17342 similar to PIR|JC7226|JC7226 endo-1,3(4)-beta-glucanase - garden
pea {Pisum sativum;} , partial (24%)
Length = 499
Score = 157 bits (396), Expect = 5e-39
Identities = 74/115 (64%), Positives = 90/115 (77%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NYR+AL KS++F +GQRSG+LP +Q++ WR DSGL DG +VDL GGYYDAGDN+KF F
Sbjct: 155 NYRDALTKSIIFFEGQRSGKLPSNQRMSWRRDSGLSDGSTMHVDLVGGYYDAGDNIKFGF 334
Query: 89 PMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGD 143
PMAFTTTMLSWS +E+G M +L AR AIRW TDYLL+ AT+ P +YV VGD
Sbjct: 335 PMAFTTTMLSWSVLEFGGMMKGELPNAREAIRWATDYLLQ-ATAYPDTIYVQVGD 496
>TC15878 homologue to UP|Q8L6S6 (Q8L6S6) Endo-beta-1,4-glucanase , partial
(24%)
Length = 786
Score = 153 bits (387), Expect = 5e-38
Identities = 80/157 (50%), Positives = 107/157 (67%), Gaps = 2/157 (1%)
Frame = +3
Query: 88 FPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPN 145
F ++FT MLSWS IEY K+M +L A A++WGTDY +K A P LY VGD N
Sbjct: 324 FSVSFTA-MLSWSIIEYDKQMASSGELDHAMEAVKWGTDYFIK-AHPEPNVLYGEVGDGN 497
Query: 146 VDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTA 205
DH CW+RPEDM T R Y + P+NPGSD+A ETAAA+AAAS+VF+ +P+YS +L+R A
Sbjct: 498 TDHYCWQRPEDMTTDRRAYRIDPSNPGSDLAGETAAAMAAASIVFRHSNPSYSSVLLRHA 677
Query: 206 QKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELL 242
+++ FA +Y+G Y S+ + +Y S SG+ DELL
Sbjct: 678 YQLFDFADKYRGKYDSSI-TVAQKYYRSISGYNDELL 785
>TC16286 similar to UP|Q8GTP6 (Q8GTP6) Endo-1,4-beta-D-glucanase , partial
(47%)
Length = 1076
Score = 149 bits (375), Expect = 1e-36
Identities = 94/193 (48%), Positives = 114/193 (58%), Gaps = 19/193 (9%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANV-----DLSGGYYDAGDN 83
NY AL K+L+F Q+SG+LP + WR SG+ DG+ V DL GGYYDAGD
Sbjct: 509 NYTLALHKALMFFNAQKSGKLPKHNNVSWRGTSGMQDGKGDGVSPTIKDLVGGYYDAGDA 688
Query: 84 VKFNFPMAFTTTMLSWSAIEY-GK-RMGPQLKEARAAIRWGTDYLLKCATSTPGRL---- 137
+KFNFP +F TMLSWS IEY GK +L + I+WGTDY LK ++ +
Sbjct: 689 IKFNFPASFAITMLSWSVIEYSGKYEAAGELNHVKDIIKWGTDYFLKTFNNSADTISTIA 868
Query: 138 -YVGVGD-------PNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLV 189
VG GD PN DH CW RPEDMD R V+ + SD+AAE AAALAAAS+V
Sbjct: 869 AQVGSGDTSGGSTTPN-DHYCWMRPEDMDYDRP---VTECHSCSDLAAEMAAALAAASIV 1036
Query: 190 FKKVDPTYSKLLV 202
FK + YSK LV
Sbjct: 1037FKD-NKAYSKKLV 1072
>BG662416
Length = 449
Score = 145 bits (366), Expect = 1e-35
Identities = 69/112 (61%), Positives = 82/112 (72%)
Frame = +3
Query: 375 SVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFH 434
S+AK QVDYILG NP +MSYMV +G N+PK +HHRGSSLPS PQ I C+ GFQ +
Sbjct: 18 SLAKTQVDYILGNNPRKMSYMVDFGENYPKHVHHRGSSLPSKHTQPQHISCNDGFQ-YLS 194
Query: 435 SSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAG 486
S + NPN LVGAIVGGP+ D F DDR++Y SEP TYIN VG +A+FAG
Sbjct: 195 SGSANPNTLVGAIVGGPDSHDSFSDDRNNYQQSEPATYINAPFVGAVAFFAG 350
>BP051279
Length = 488
Score = 145 bits (365), Expect = 2e-35
Identities = 68/107 (63%), Positives = 82/107 (76%)
Frame = -3
Query: 379 RQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNP 438
RQVDYILG+NP+ +SYMVGYG +P+RIHHRGSSLPS+ HPQ I C G F++S+NP
Sbjct: 486 RQVDYILGDNPMGLSYMVGYGNYYPQRIHHRGSSLPSIKDHPQFIACKEG-SVFYNSTNP 310
Query: 439 NPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
NPN+ GA+VGGP + D + DDR D+ SEPTTYIN VG LAYFA
Sbjct: 309 NPNVHEGAVVGGPGEDDFYVDDRVDFRKSEPTTYINAPFVGVLAYFA 169
>AU088935
Length = 683
Score = 142 bits (359), Expect = 9e-35
Identities = 68/142 (47%), Positives = 94/142 (65%)
Frame = +2
Query: 111 QLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNN 170
+L+ AI+WGTDY +K TS P L+ VGD + DH CW+R EDM T+R + + +N
Sbjct: 209 ELEHTMEAIKWGTDYFIKAHTS-PNVLWAEVGDGDTDHYCWQRAEDMTTSRQAFKIDESN 385
Query: 171 PGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPF 230
PGSD+A ETAAA+AAAS+VFKK +P YS LL+ AQ++++F +Y Y ++G V +
Sbjct: 386 PGSDLAGETAAAMAAASIVFKKTNPHYSHLLLHHAQQLFEFGDKYXRKYDATIG-VVKKY 562
Query: 231 YCSYSGFKDELLWGAAWLFRAT 252
Y S G+ D LLWG WL+ T
Sbjct: 563 YASVXGYXDXLLWGXXWLYNTT 628
Score = 33.5 bits (75), Expect = 0.080
Identities = 18/45 (40%), Positives = 24/45 (53%), Gaps = 4/45 (8%)
Frame = +3
Query: 273 FSWDDKYAGAHVLLSRRALLNGDKN----FDQYRQEAENFMCKIL 313
FSWD KYAG VL S+ + K +YR +AE ++C L
Sbjct: 54 FSWDVKYAGLQVLASKFLMEEKHKKHVNILKKYRSKAEFYICSCL 188
>TC10070 similar to UP|O81403 (O81403) Endo-1,4-beta-glucanase precursor ,
partial (20%)
Length = 573
Score = 136 bits (342), Expect = 9e-33
Identities = 66/98 (67%), Positives = 75/98 (76%), Gaps = 1/98 (1%)
Frame = +1
Query: 388 NPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAI 447
NPL MSYMVGYG +P+RIHHRGSSLPS+A HP IGC G Q FF S NPNPN+LVGA+
Sbjct: 1 NPLGMSYMVGYGARYPQRIHHRGSSLPSVATHPAHIGCKAGSQYFF-SPNPNPNVLVGAV 177
Query: 448 VGGP-NQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
VGGP N +D FPD R + SEPTTYIN +VG LA+F
Sbjct: 178 VGGPTNNTDSFPDSRPFFQESEPTTYINAPLVGLLAFF 291
>BF177461
Length = 439
Score = 110 bits (275), Expect = 5e-25
Identities = 49/115 (42%), Positives = 70/115 (60%)
Frame = +1
Query: 324 TQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDY 383
T GGL++ +NLQY +S FLL Y+ Y++A K NC + P L S AK Q DY
Sbjct: 67 TPGGLVYVHDWNNLQYASSAAFLLAVYSNYLNAAKSQLNCPEGQIQPQELLSFAKSQADY 246
Query: 384 ILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNP 438
ILG+NP +MSY++GYGPN+P +HHR S+ S++ + C GF+ ++ P
Sbjct: 247 ILGKNPKEMSYLIGYGPNYPSHVHHRDXSIASISVLHSEVSCVQGFEAWYRRGEP 411
>AV421757
Length = 286
Score = 105 bits (262), Expect = 2e-23
Identities = 50/95 (52%), Positives = 68/95 (70%), Gaps = 4/95 (4%)
Frame = +2
Query: 198 SKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKY 257
S L+R A +V+QFA +++GSYS++L VCPFYCSYSG++DELLWGAAWL +AT Y
Sbjct: 2 SNTLIRRAIRVFQFADKHRGSYSNALKPIVCPFYCSYSGYEDELLWGAAWLHKATKNPMY 181
Query: 258 YNLIKT----LGADDQPDIFSWDDKYAGAHVLLSR 288
N I+ LGA + + F WD+K+ GA +LLS+
Sbjct: 182 LNYIQVNGQILGAAEFDNTFGWDNKHVGARILLSK 286
>AV769339
Length = 551
Score = 90.1 bits (222), Expect = 7e-19
Identities = 42/71 (59%), Positives = 50/71 (70%)
Frame = -2
Query: 415 SLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYIN 474
S+ HP I C GF +S +PNPN+LVGA+VGGP+Q D FPD+RSDY SEP TYIN
Sbjct: 550 SIGVHPGKIQCSEGFS-VMNSQSPNPNVLVGAVVGGPDQHDMFPDERSDYEQSEPATYIN 374
Query: 475 GAVVGPLAYFA 485
+VG LAY A
Sbjct: 373 APLVGALAYLA 341
>TC15877 similar to UP|O65186 (O65186) Cellulase, partial (12%)
Length = 398
Score = 84.3 bits (207), Expect = 4e-17
Identities = 44/89 (49%), Positives = 58/89 (64%), Gaps = 6/89 (6%)
Frame = +1
Query: 7 SFMSLLFLIPMLLFVGNVQCNP------NYREALAKSLLFLQGQRSGRLPPDQQLKWRSD 60
S L+ + P++L + C+P +Y AL+KS+LF + QRSG LP Q++ WRS
Sbjct: 142 SMEKLICMFPLILLL----CSPFAFAGHDYGLALSKSILFYEAQRSGVLPSHQRVTWRSH 309
Query: 61 SGLFDGRLANVDLSGGYYDAGDNVKFNFP 89
SGL DG+ + VDL GGYYDAGDNVKF P
Sbjct: 310 SGLQDGKASGVDLVGGYYDAGDNVKFGLP 396
>AV778020
Length = 465
Score = 76.3 bits (186), Expect = 1e-14
Identities = 37/91 (40%), Positives = 55/91 (59%)
Frame = -3
Query: 397 GYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDG 456
G+G F + +HHRG+S+P+ + C GG + + +S PNP L GA+VGGP++ D
Sbjct: 463 GFGNKFARHVHHRGASIPN---DKKPYSCHGGMK-WRDTSKPNPYTLTGAMVGGPDRFDP 296
Query: 457 FPDDRSDYSHSEPTTYINGAVVGPLAYFAGI 487
FPD RS+Y+++EPT N +V L I
Sbjct: 295 FPDSRSNYNYTEPTLAGNAGIVAALVSLTNI 203
>TC8688 homologue to UP|AAS45400 (AAS45400) Endo-1,4-beta-glucanase ,
partial (18%)
Length = 750
Score = 75.1 bits (183), Expect = 2e-14
Identities = 37/86 (43%), Positives = 55/86 (63%)
Frame = +3
Query: 401 NFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDD 460
+FPK +HHRG+S+P + C GG++ + +S PNPN LVGA+V GP++ DGF D
Sbjct: 18 HFPKHVHHRGASIPK---NKVKYSCKGGWK-WRDTSKPNPNTLVGAMVAGPDKHDGFHDV 185
Query: 461 RSDYSHSEPTTYINGAVVGPLAYFAG 486
R +Y+++EPT N +V L +G
Sbjct: 186 RMNYNYTEPTIAGNAGLVAALVALSG 263
>BP073341
Length = 416
Score = 50.4 bits (119), Expect(2) = 1e-09
Identities = 25/49 (51%), Positives = 35/49 (71%)
Frame = +1
Query: 160 ARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKV 208
+R Y + +PGSD+AAETAAA AAA++ FK + +YS LL+ AQ+V
Sbjct: 190 SRAAYKIDEQHPGSDLAAETAAAWAAAAIAFKPYNSSYSSLLLVHAQQV 336
Score = 28.9 bits (63), Expect(2) = 1e-09
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = +3
Query: 141 VGDPNVDHKCWERPEDMDTAR 161
VGD DH WER EDM T +
Sbjct: 132 VGDGASDHSGWERAEDMTTVK 194
>TC17987 similar to UP|O04890 (O04890) Endo-1,4-beta-glucanase , partial
(25%)
Length = 501
Score = 54.3 bits (129), Expect = 4e-08
Identities = 45/130 (34%), Positives = 62/130 (47%), Gaps = 17/130 (13%)
Frame = +2
Query: 244 GAAWLFRATNAVKYYNLIKTLGADDQ-------PD--IFSWDDKYAGAHVLLSR-RALLN 293
G AW++ AT Y L T G PD + SWD+K AGA VLLSR R L+
Sbjct: 71 GGAWMYFATGNNSYLKLATTPGLAKHAGAFWGGPDYGVLSWDNKLAGAQVLLSRLRLFLS 250
Query: 294 GDKNFDQ----YRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSN---LQYVTSITFL 346
+++ + + MC LP +S T+GGL+ +L LQYV + FL
Sbjct: 251 PGYPYEEILSAFHNQTSIIMCSYLP--VFTSFNRTKGGLI-QLNHGRPQPLQYVVNAAFL 421
Query: 347 LTTYAKYMSA 356
T Y+ Y+ A
Sbjct: 422 ATLYSDYLDA 451
>AV423824
Length = 181
Score = 39.3 bits (90), Expect = 0.001
Identities = 20/41 (48%), Positives = 27/41 (65%)
Frame = +2
Query: 11 LLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPP 51
LL L+ + G+ +Y +AL KS+LF +GQRSGRLPP
Sbjct: 74 LLLLLSPIAAAGH-----DYHDALRKSILFFEGQRSGRLPP 181
>BF177615
Length = 347
Score = 29.6 bits (65), Expect = 1.2
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = -2
Query: 194 DPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVC 228
DP Y +L R Q + A G Y D+LGS +C
Sbjct: 322 DPAYGQLYGRNPQSAHA*A*YSMGIYHDNLGSDIC 218
>TC16932 similar to UP|Q94A94 (Q94A94) AT5g11880/F14F18_50, partial (30%)
Length = 616
Score = 29.3 bits (64), Expect = 1.5
Identities = 14/23 (60%), Positives = 18/23 (77%)
Frame = -2
Query: 310 CKILPNSPSSSTQYTQGGLMFKL 332
C+IL PSSSTQY+ GG +F+L
Sbjct: 399 CRILLTDPSSSTQYS-GGRIFRL 334
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,173,538
Number of Sequences: 28460
Number of extensions: 137011
Number of successful extensions: 612
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 591
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 595
length of query: 488
length of database: 4,897,600
effective HSP length: 94
effective length of query: 394
effective length of database: 2,222,360
effective search space: 875609840
effective search space used: 875609840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0083.8


